Abstract
Despite active research into methicillin-resistant Staphylococcus aureus (MRSA) in pigs since 2004, the ecology of the susceptible ancestral organism has been neglected. A longitudinal study of pigs in 2 intensive production systems was conducted to investigate the effects of age and anatomical site on detection of S. aureus. Sampling was replicated in 2 cohorts per farm, with swabs collected from the nares, tonsils, skin (axilla), and rectum in lactating sows, suckling, weaned, and market-age pigs, plus the vagina of sows. No MRSA were isolated, but S. aureus was detected in a least 1 site in 175 (91.1%) out of 192 pigs. Pig-level prevalence did not differ among the age groups, but the proportion of positive samples (all sites) was higher in market-age pigs (75.2%) and nursery-age pigs (63.2%) than in sows (40.7%) and suckling piglets (38%). Prevalence did not differ among nasal (67.9%), skin (62.3%), and tonsil (61.7%) swabs, but was lower in rectal (42%) and vaginal swabs (39.6%). Multiple multilocus sequence typing (MLST) and spa types were found in both production systems, but all isolates were of ST398, ST9, or ST5. These MLST lineages have been variably predominant among reports of MRSA in pigs on 3 continents, and the presence of methicillin-sensitive variants in several countries raises the likelihood that MRSA in pigs has likely resulted from independent acquisition of the mecA gene by multiple S. aureus lineages that have been adapted to swine over the long term, rather than recent introduction of novel clones into swine populations.
Keywords
Introduction
Several species of staphylococci are recognized as common commensals or opportunistic pathogens of pigs. 9 From the perspective of swine health, only Staphylococcus hyicus is considered a primary pathogen as the cause of exudative epidermitis (greasy pig disease). 9 There are numerous reports of the isolation of Staphylococcus aureus from pigs, and it is generally assumed that, as in human beings, S. aureus is part of the “normal” bacterial flora of swine. 9 Anatomical sites from which S. aureus has been isolated from pigs include skin, 23 tonsils,28,34 feces,5,11 nose, 10 internal organs,28,31 and arthritic joints. 30 Although commonly detected in a range of lesions in pigs, S. aureus is viewed as an opportunistic pathogen, as cases tend to be sporadic and outbreaks of disease are rare. 9 There are reports of S. aureus being isolated from all age groups of pigs from newborn to adult.4,14 However, owing to the relative unimportance of S. aureus as a swine pathogen, the ecology and epidemiology of this organism in swine populations has not been extensively studied.
The revelation in The Netherlands in 2004 that livestock may represent a substantial reservoir of methicillin-resistant S. aureus (MRSA) unleashed a flurry of research of MRSA in pigs and other animals over the past decade.8,13,32 However, with few exceptions, researchers have employed selective culture methods to focus on MRSA alone, and the biology and ecology of the ancestral organism (S. aureus) remains neglected. This focus was initially appropriate due to the need to understand the extent of the livestock MRSA reservoir in the early phase of great uncertainty. However, as discussions begin to consider options to control MRSA in livestock, 24 a more comprehensive understanding of the ecology of S. aureus in pigs is required to assess the feasibility of potential interventions. The objectives of the current study were to compare the effect of anatomical site and age of pigs on detection of S. aureus and to describe the diversity of S. aureus spa types within herds.
Materials and methods
Study design and source population
A longitudinal study was conducted during 2011 and 2012 in 2 unrelated multiple-site pig production systems located in Minnesota. The farms were selected by convenience based on willingness of the producers to participate. In both systems, piglets were weaned at approximately 3 weeks of age and transferred to off-site growing facilities. Sampling was conducted in the farrowing rooms at the breeding herd sites, and during both the nursery and finishing phases at growing pig sites. Two cohorts were sampled in each system, with an interval of at least 6 months between cohorts. At each breeding herd site, samples were collected from lactating sows (n = 12) in rooms housing the youngest litters, and samples were also collected from 1 piglet per litter of the sows sampled (n = 12). Subsequent sampling of the same birth cohorts of pigs (but not the same individual pigs) occurred 4 and 20 weeks later at “nursery” (n = 12) and “market” (n = 12) age phases of pig production, respectively. Individual pigs (1 per pen) were purposively selected from different pens throughout the barns. Swab samples were collected from the nose, tonsil, rectum, and skin (axilla) of each pig. Additionally, vaginal samples were collected from sows. In one system, the respective cohorts were moved to different wean-to-finish facilities. The market-age pigs of the second cohort were reared at relatively low density in a naturally ventilated hoop barn with straw bedding. All other groups were housed in total confinement on perforated floors in power-ventilated buildings.
Animal sampling
Using sterile swabs, a nasal, tonsil, rectal, and skin (axilla) samples were collected from 192 pigs, and vaginal swabs were also collected from the 48 sows. To collect nasal samples, a single swab was inserted 2–7 cm (according to pig size) into both nostrils and gently rotated against the mucosal epithelium. To enable collection of tonsillar samples, a speculum was used to open the mouth, and the swab was gently rotated against the tonsillar epithelium. Skin samples were obtained by rubbing a swab onto the axillary folds of pigs. Rectal swabs were obtained by introducing a swab approximately 5 cm into the rectum of pigs and gently rotating the swab in the rectal contents. For vaginal sampling, using circular rotations, a swab was gently rubbed against the vaginal epithelium approximately 5 cm into the vagina. Following sample collection, the swabs were individually identified and stored on ice until processed in the laboratory within 24 hr of collection. All procedures with animals were approved by the University of Minnesota Institutional Animal Care and Use Committee under protocol number 1105B99155.
Isolation and identification of Staphylococcus aureus bacteria
All samples were cultured in parallel by 2 procedures, described as “nonselective” (for S. aureus) and “selective” (for MRSA). All swabs were placed in 15-ml tubes b with 5 ml of Mueller–Hinton broth c containing 6.5% NaCl d and incubated at 37°C for 24 hr. The procedure to isolate S. aureus consisted of transferring a 1-ml aliquot from the Muller–Hinton broth into 9 ml of phenol–red mannitol broth. e After 24 hr of incubation at 37°C, if the broth changed color, one 10-µl inoculating loop of broth was streaked onto a Columbia CNA (colistin and nalidixic acid) agar plate with 5% sheep blood. e The plates were incubated at 37°C for 24 hr. Two presumptive S. aureus colonies (round, golden-yellow, colonies with hemolysis) were then each restreaked onto Columbia CNA plates and incubated at 37°C for 24 hr. Once pure cultures were obtained, all suspected isolates were confirmed as S. aureus using the tube-coagulase test f and a commercial S. aureus latex agglutination assay. g
To detect MRSA, a 1-ml aliquot from the Mueller–Hinton broth was transferred into 9 ml of phenol–red mannitol broth e containing 4 mg/l of oxacillin. d Samples showing color change indicating mannitol fermentation after 24-hr incubation at 37°C were streaked (a 10-µl inoculating loop) onto a chromogenic agar plate h and incubated at 37°C for 24 hr. Isolates confirmed to be S. aureus were stored in lysogeny broth f containing 17.5% glycerol at −80°C prior to genotypic characterization.
Genotypic characterization
From the glycerol solution, each S. aureus isolate (n = 463) was streaked to a Columbia CNA agar plate with 5% sheep blood e and incubated at 37°C for 24 hr. One fresh colony was added into a polymerase chain reaction (PCR) microplate well i with 19.5 µl of 10 mM Tris–HCl j and 0.5 µl of 1 mg/ml lysostaphin. d Then, the plates were heated at 37°C for 15 min and stored at −20°C for further procedures.
All S. aureus isolates were typed by DNA sequence analysis of the X region of the staphylococcal protein A (spa) gene as previously described with some modifications. 27 The PCR amplification of the simple sequence repeat region of the spa gene was performed as follows. The reaction mix consisted of the following: 1.0 μl of the DNA of the samples, 9.5 μl of nuclease-free water, 1.0 μl of 1095F forward PCR primer at 10 μM, 1.0 μl of 1517R reverse PCR primer at 10 μM, and 12.5 μl of a commercial PCR master mix (2×) j were added to 0.2-ml PCR tubes. A negative control (nuclease-free water) and a positive control were included. The PCR tubes were placed in a thermocycler. The thermal cycling parameters were 2 min at 95°C, then 30 cycles of 30 sec at 94°C, 30 sec at 55°C; 1 min at 72°C, and a final extension at 72°C for 5 min. Completed reaction mixtures were stored at −20°C. The amplicons were sequenced using regular Sanger sequencing at the Biomedical Genomics Center (BMGC), University of Minnesota. The sequences of the spa genes from the BMGC were submitted to the eGenomics program (www.egenomics.com) and Ridom SpaServer (www.spaserver.ridom.de), which use computerized algorithms to analyze the variability of the spa gene X region and assign spa types. The eGenomics spa types are reported using a numerical system (e.g., spa type 539) while Ridom spa types are reported using a numerical system preceded by a “t” (e.g., spa t034). Multilocus sequence typing (MLST) of selected isolates of each spa type was performed as previously described. 6 The sequences of the relevant 7 housekeeping genes were submitted to the MLST site (http://www.mlst.net) in order to obtain a sequence type.
Statistical analysis
The current study investigated the effects of “anatomical sampling site” (nose, tonsil, rectum, and skin or vagina) and “pig age group” (sow, preweaning piglet, nursery age, and market age) on S. aureus prevalence after adjusting for individual pig, cohort, and farm. The comparison of S. aureus prevalence between the different pig age groups was performed at 2 levels: pig level and sample level. At the pig level, a pig was considered S. aureus–positive if at least 1 anatomical site was culture positive for S. aureus. At the sample level, all samples (i.e., from all anatomical sites) contributed to the prevalence calculation. The statistical analyses were performed with a commercial package k using generalized mixed models (Proc Glimmix), with a logit link function for binary outcomes (Proc Glimmix) and random effects to reflect potential correlations at the pig, cohort, and farm levels. The model used Wald-type test statistics and confidence intervals. The pairwise comparisons were made with least-squares means, and statistical significance was set at α = 0.05 for all analyses.
Results
All S. aureus isolates detected in the 2 farms were methicillin-sensitive S. aureus (MSSA). In the pig-level analysis (Table 1), the prevalence of S. aureus–positive pigs (175/192 pigs; 91.1%) did not differ among age groups (P = 0.55). In the sample-level analysis, prevalence of positive samples was highest in market-age pigs, followed by nursery-age pigs and suckling piglets and sows (Table 1). There was no difference in S. aureus prevalence among samples from suckling piglets and sows (P = 0.62). However, the prevalence of S. aureus in nursery pig samples was higher (P < 0.0001) than in samples of sows or preweaning piglets, but lower than that of samples from market-age pigs (P = 0.016). All market-age pigs were positive in at least 1 anatomical site except for 2 pigs in the group kept in less intensive conditions. The proportion of samples positive for S. aureus varied among anatomical sites (p-value < 0.001). Prevalence was higher in nose (68%), skin (62%), and tonsil (62%) samples compared with rectal (42%) and vaginal (40%) samples (Table 2). In the 12 market-age pigs sampled in less intensive housing, tonsil swabs were positive in 10 pigs, 2 of which were also positive on rectal swabs, but the corresponding nasal and skin swabs from these animals did not yield S. aureus.
Staphylococcus aureus prevalence (least-square means) at the pig level (proportion of pigs with at least 1 culture-positive anatomical site) and at the sample level (proportion of culture-positive samples across all anatomical sites) by age group.*
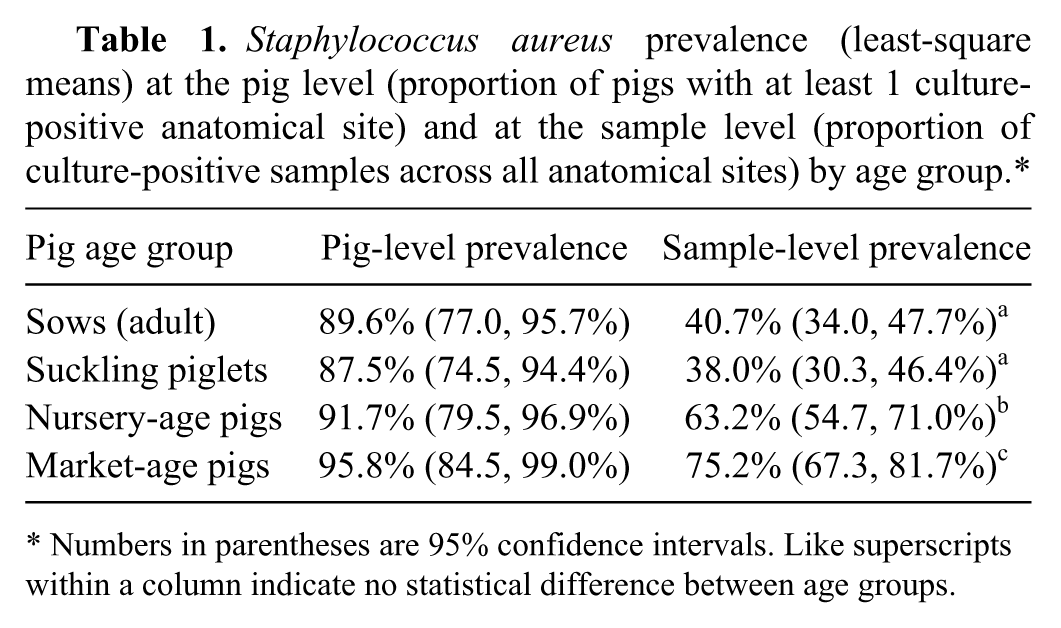
Numbers in parentheses are 95% confidence intervals. Like superscripts within a column indicate no statistical difference between age groups.
Proportion of swab samples positive for Staphylococcus aureus by anatomical site (least-square means).*
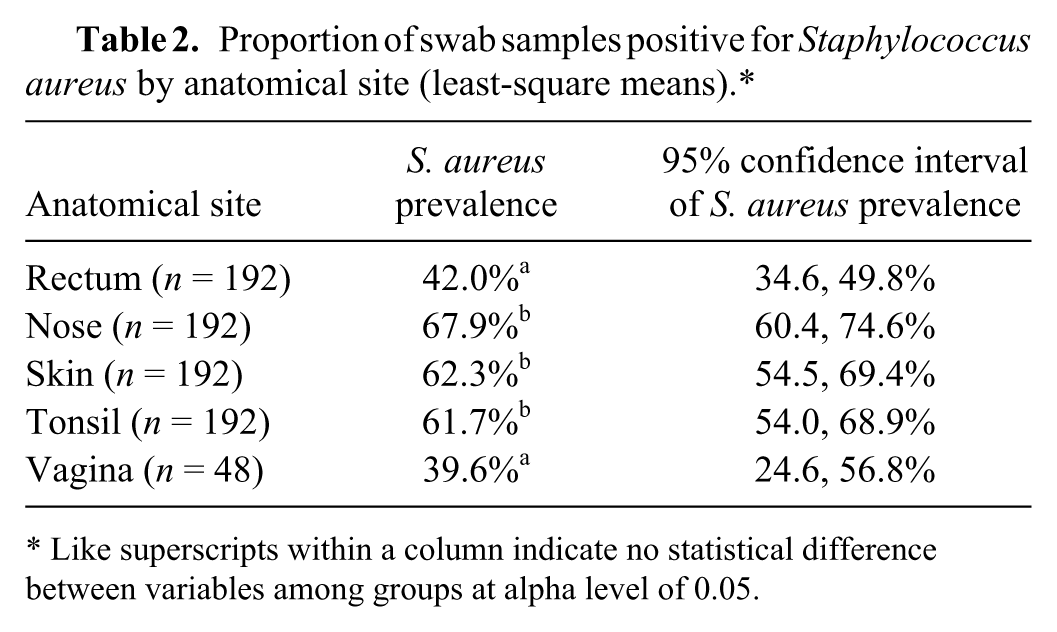
Like superscripts within a column indicate no statistical difference between variables among groups at alpha level of 0.05.
Multiple spa types (6 in farm 1, 5 in farm 2) and multiple sequence types of S. aureus were detected on both farms (Table 3). The predominant spa types were t034 (ST398; n = 163) and t337 (ST9; n = 127), which were found on both farms and comprised 65% of all isolates. All spa types belonged to ST398, ST9, or ST5. Multiple spa types (up to 3) were isolated from individual pigs on 59 occasions. There was no apparent association of spa type with anatomical sites.
Sequence types and spa types of Staphylococcus aureus isolates from pigs in 2 multiple-site systems.
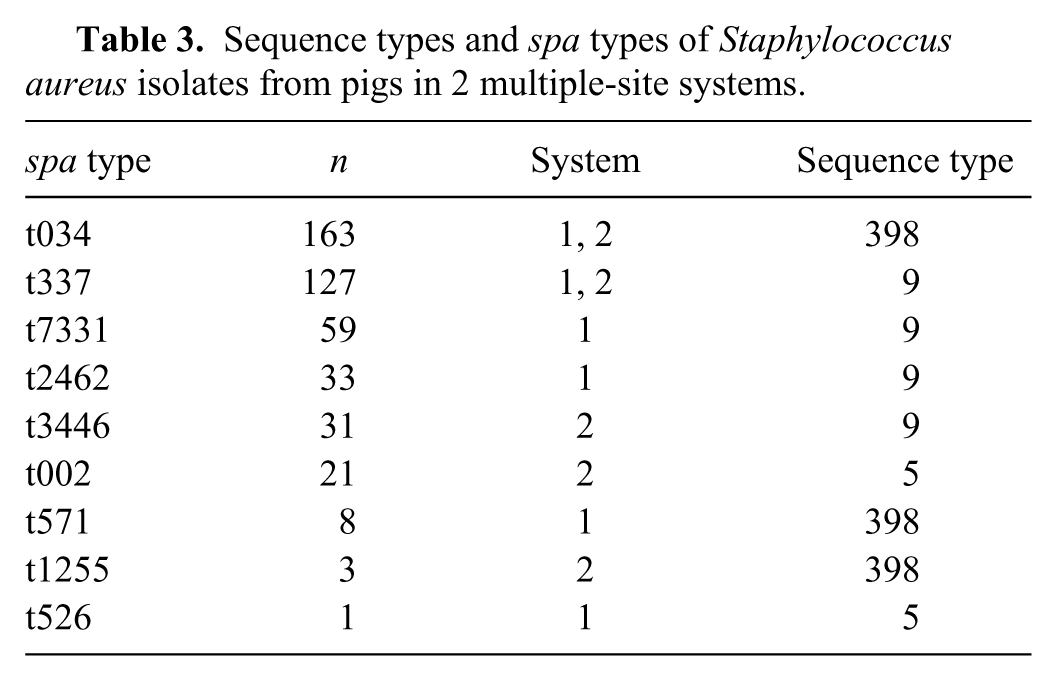
Discussion
Although S. aureus is generally considered to be a member of the normal flora of pigs, there has been minimal research to describe the ecology of S. aureus in swine populations. In a broad review of the ecology of staphylococci in human beings and animals, mammalian skin (including the anterior nares) was stated to be the major habitat of staphylococci, but staphylococci also occur in other regions of the body such as the throat, mouth, mammary glands, and intestinal tract, albeit less frequently or in smaller numbers than on skin. 16 The author of the review also observed that staphylococcal populations of pigs were not well characterized. 16 Research into S. aureus in swine has been stimulated by the discovery that MRSA variants occur commonly in pigs. 32 However, to the authors’ knowledge, systematic investigation of the ecology of S. aureus has not previously been conducted.
Overall, the observations made in the 2 herds in the current study support the contention that S. aureus constitutes part of the normal bacterial flora of pigs. Approximately 90% of pigs sampled in all age groups were culture positive for S. aureus in at least 1 anatomical site. However, a higher proportion of anatomical sites were positive in growing pigs than in sows and suckling pigs. A study comparing nasal and tonsillar flora of pigs found that S. aureus and most lactobacilli became more prevalent after weaning. 2 Previous studies using nasal swabs to detect MRSA in swine herds have also suggested that the prevalence of positive swabs may be higher in growing pigs than suckling pigs.3,19 In contrast to the finding of higher prevalence in samples from older growing pigs in the current study, a prior study reported a decline in prevalence with age in nasal swabs of growing pigs, with prevalence highest in nursery-age pigs. 14 It is important to note that these few studies have been conducted on small numbers of farms, and there is evidence for variation among farms. 3 Furthermore, environmental factors, such as animal density and the markedly different ventilation rates used in confinement facilities in different seasons, have the potential to confound longitudinal studies of prevalence in different age groups of pigs.
The data in the current study indicates that the nose, tonsils, and skin are anatomical sites of pigs that are all likely to be culture positive for S. aureus in pigs reared in confinement. However, as the pigs in these commercial units are housed in groups, it is impossible to determine whether sites are colonized or transiently contaminated. 3 For this reason, the axilla was selected as the location for skin sampling, as it is a relatively protected site that is likely less prone to direct environmental contamination. Theoretically, the skin, nose, or tonsil could be the preferred site for S. aureus colonization of pigs, with other sites being contaminated from that primary reservoir. Perhaps a more likely and nuanced scenario is that different anatomical niches provide conditions that are variably supportive of S. aureus colonization and persistence. 29 The chance observation that the tonsil, but not nasal or skin swabs, was frequently positive in the group of pigs housed less intensively is intriguing and needs to be explored further. One 2013 experimental study reported markedly higher numbers of S. aureus in pharyngeal (including tonsil) tissues of pigs than in nasal samples, 12 and a similar observation was reported for naturally infected pigs. 2 The importance of the oropharynx as a site of colonization of people with S. aureus is well established, with several studies reporting higher prevalence in oropharyngeal than nasal samples. 29 It has been suggested that housing of pigs in isolation could give some insight into which locations are most supportive of S. aureus persistence. 3 A study of wild boar in Spain found S. aureus in 18% of animals, with a higher prevalence in nasal swabs than skin swabs, but tonsil samples were not collected. 25 Furthermore, as in the current study, multiple spa types were also found in individual pigs in that study. Regardless, for pigs housed intensively in groups on commercial farms, any of these sites should be adequate for sampling of herds for epidemiological studies, although sampling of single anatomical sites will underestimate prevalence and diversity of S. aureus at the pig level. For sampling of pigs housed at low density or outdoors, the observations in the current study indicate that it would be prudent to sample the tonsil as well as the nose or skin. For studies exploring mechanisms of colonization, including metagenomic studies, one should not assume the nares are the predominant site of S. aureus colonization in the pig or that pigs harbor only a single spa type.
The diversity of S. aureus spa types detected on both farms is not unexpected as considerable genetic diversity is typically found among commensal organisms within a host species (e.g., Escherichia coli in most species; Streptococcus suis or Haemophilus parasuis in pigs).17,20,26 The 3 MLST types (ST9, ST398, ST5) detected in the present study correspond closely with those reported in studies of commercial pigs in Europe,1,21 which is significant for 2 reasons. First, this similarity, despite different study designs and considerable temporospatial separation of the study populations, suggests that these lineages may be widely distributed among commercial swine populations. However, data from a study of a single farm in Senegal, 7 and observations on wild boar in Spain, 25 indicate that these MSLT lineages are not universal among swine populations. Second, while ST398 is the predominant lineage of livestock-associated MRSA in Europe, 13 ST9 has been predominant in most Asian countries studied, 33 and ST5 MRSA has been commonly isolated from pigs in North America.10,15,18 A 2014 study of S. aureus in slaughtered pigs in northeastern China found that ST398/t034 was the predominant strain of MSSA, while all MRSA isolates were ST9. 33 It is feasible that the emergence of livestock-associated MRSA has resulted from independent acquisition of the mecA gene by multiple S. aureus lineages that have been adapted to swine over the long term, rather than recent introduction of novel clones into swine populations.
Some limitations of the current study are that the sample size for each sampling event was only 12 pigs, and only 2 herds were involved. There was no credible prior data for expected prevalence, but the core assumption was that prevalence would be high in at least 1 anatomical site based on the general opinion that S. aureus is part of the normal bacterial flora of pigs. Given the high prevalence of S. aureus found in most anatomical sites, the small number of pigs sampled is arguably a minor concern. The large number of isolates evaluated on these farms overall greatly exceeds that in earlier studies, and likely provides reasonably reliable identification of the predominant spa types on the farms overall. By comparison, an early study sampled only 14 swine S. aureus isolates from laboratory submissions to veterinary laboratories across France, 1 while a broader (41 farms) but less intensive study (mean of 8.8 pigs per farm) was undertaken in Switzerland. 21 Similarly, a 2013 study of relatively small farms in Connecticut sampled a mean of 5.1 pigs across 51 farms, reporting a S. aureus prevalence in nasal swabs of 30%. 22 Six spa types were found among the 22 isolates assessed, with t337 (9 isolates) being predominant. 22 More extensive and systematic investigation of S. aureus is necessary to further understand the ecology of this agent in swine production. Design of future studies should consider the likely diversity of S. aureus within farms, and studies of small or extensively raised populations should include tonsillar sampling in addition to the usual approach of nasal swabs.
Footnotes
Acknowledgements
The authors wish to thank the collaborating producers and their veterinarians for facilitating the study on their farms.
a.
BBL culture swab, Liquid Stuart medium single plastic applicator; BD, Sparks, MD.
b.
BD Falcon 15 ml conical centrifuge tubes, Fisher Scientific Inc., Waltham, MA.
c.
Teknova, Hollister, CA.
d.
Sigma-Aldrich, St. Louis, MO.
e.
BBL, BD, Franklin Lakes, NJ.
f.
Difco Laboratories Inc., Detroit, MI.
g.
Pastorex Staph-plus, Bio-Rad Laboratories, Hercules, CA.
h.
MRSAselect, Bio-Rad Laboratories, Hercules, CA.
i.
Genomic, 96-well, Eppendorf, Twin.tec, PCR, yellow, extra-thin polypropylene wells, semiskirted, polycarbonate frame, automation friendly; Fisher Scientific Co., Pittsburgh, PA.
j.
HotStart-IT FideliTaq, Research Products International Corp., Mt. Prospect, IL.
k.
SAS package version 9.3, SAS Institute Inc., Cary, NC.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The research was funded by the National Pork Board (grant 11-016).
