At present, despite extensive laboratory investigations, most cases of porcine abortion remain without an etiological diagnosis. Due to a lack of recent data on the abortigenic effect of order Chlamydiales, 286 fetuses and their placentae of 113 abortion cases (1–5 fetuses per abortion case) were investigated by polymerase chain reaction (PCR) methods for family Chlamydiaceae and selected Chlamydia-like organisms such as Parachlamydia acanthamoebae and Waddlia chondrophila. In 0.35% of the cases (1/286 fetuses), the Chlamydiaceae real-time PCR was positive. In the Chlamydiaceae-positive fetus, Chlamydia abortus was detected by a commercial microarray and 16S ribosomal RNA PCR followed by sequencing. The positive fetus had a Porcine circovirus-2 coinfection. By the Parachlamydia real-time PCR, 3.5% (10/286 fetuses of 9 abortion cases) were questionable positive (threshold cycle values: 35.0–45.0). In 2 of these 10 cases, a confirmation by Chlamydiales-specific real-time PCR was possible. All samples tested negative by the Waddlia real-time PCR. It seems unlikely that Chlamydiaceae, Parachlamydia, and Waddlia play an important role as abortigenic agents in Swiss sows.
Introduction
Chlamydiae are Gram-negative, obligate intracellular bacteria that occur worldwide and affect a broad host range. The bacteria show a biphasic life cycle: metabolically inactive elementary bodies (EBs) get internalized into host cells and reorganize into reticulate bodies (RBs, metabolically active forms). After replication and condensation back into EBs, cell lysis is initialized, and new cells can be infected. In the host, the primary replication is localized in mucosal epithelial cells of the respiratory, gastrointestinal, and urogenital tract, or conjunctival epithelium, in trophoblastic epithelium of the placenta, as well as monocytes and macrophages.
26
Recent nomenclature16,22 describes the Chlamydiaceae family as including genus Chlamydia, with 9 species.
Chlamydia was first detected in pigs in 1955, from animals with arthritis and pericarditis.
34
Since then, many studies have demonstrated a broad range of clinical manifestations due to chlamydial infections in pigs. To date, the following Chlamydia species have been detected in pigs, with mixed infections being a common finding: C. suis, C. abortus, C. psittaci, C. pecorum, and C. trachomatis.20,21,25 Infections of the female genital tract that induce miscellaneous reproductive failures in sows such as return to estrus, dysgalactia, abortions, neonatal mortality, and mummification have been regarded as being of important economic relevance.
20
Furthermore, C. psittaci, C. suis, C. trachomatis, and C. abortus were isolated from the genital tract (oviduct and uteri) of sows with conception problems.
20
In sows, C. abortus is primarily related to the incidence of abortions and less viable neonates.
31
Chlamydia pecorum in pigs is associated with urogenital tract infections, enteritis, inapparent intestinal infections, pericarditis, conjunctivitis, mastitis, pleuritis, pneumonia, encephalomyelitis, polyarthritis, and also abortions.15,20,34 Chlamydia suis can cause enteritis, conjunctivitis, rhinitis, pneumonia, and reproductive disorders, such as return to estrus and inferior semen quality.15,31 Chlamydia psittaci infects primarily birds, but was isolated from the genital tract of Swiss breeding sows
9
and was used to infect gnotobiotic pigs in an experimental model, indicating that pigs are a possible host for C. psittaci.
18
Chlamydia trachomatis has been detected in a study examining oviducts and uteri of sows with conception failures.
20
Recent research has indicated an abortigenic potential of Chlamydia-like organisms in ruminants, such as Parachlamydia acanthamoebae6,12 and Waddlia chondrophila.13,19 Therefore, the current study aimed to investigate the significance of chlamydial infections due to the Chlamydiaceae family as well as Chlamydia-like organisms such as Waddlia and Parachlamydia in porcine abortions from farms with fertility disorders.
Materials and methods
Sampling
From January until October 2009, fetuses with their placentae were collected. The material originated from 59 pig-breeding farms with fertility disorders (reproductive problem group). The farms were distributed over 12 Swiss cantons, of which 44 farms (74.4%) represented areas with a high swine density. All investigated farms had not yet started the Porcine circovirus-2 (PCV-2) vaccination scheme. From each farm, up to 3 sows with their piglets (normally 3 fetuses, including predominantly mummies and 1 nonautolytic fetus) from the same gestation period were examined. In summary, 286 fetuses from 113 sows on 59 farms with reproductive disorders including abortions, mummies, stillborn, and weak born piglets were collected. In the majority of cases, the placenta was available (n = 99/113). Of the 113 sows, 42 had early abortions before the 110th pregnancy day and 71 sows had at-term delivery but with at least 1 dead piglet (mummified, autolytic, stillborn piglet). As a control group, 70 fetuses out of 7 litters (10 fetuses/litter) were investigated. The fetuses were from 7 healthy slaughter sows in late-stage pregnancy (110th pregnancy day). Immediately after slaughtering of the sows, the pregnant uteri were removed and the first 10 fetuses were taken for further investigations.
Histopathological, viral, and bacteriological analysis
All fetuses from the reproductive problem group (n = 286) and the control group (n = 70) were investigated by necropsy and histological examination of heart, lung, liver, kidney, spleen, mesenterial lymph node, thymus, brain, and placenta. The reproductive problem group was tested by immunohistochemistry for PCV-2 as previously described24,32 and for Porcine parvovirus (PPV) by immunoelectron microscopy and indirect fluorescent antibody test.
35
Preliminary selected fetuses (n = 44) were tested for Porcine enterovirus (PEV) types 8, 9, and 10 and Porcine teschovirus (PTV) by polymerase chain reaction (PCR).
36
Placenta and fetal organs were examined for mesophilic aerobic bacteria by cultivation followed by differentiation using biochemical and molecular biological methods as described previously.
27
Smear preparations of the placentae were investigated for Brucella sp. by Köster and Gimenez staining. Warthin–Starry staining was performed in histological slides indicative for Leptospira spp. infection. The following selected organ tissues of both groups were investigated for Chlamydia, Parachlamydia, and Waddlia species: placenta, liver, heart, mesenterial lymph node, lung, spleen, thymus, kidney, and (when available) umbilicus.
DNA extraction
Thirty micrometer sections of formalin-fixed and paraffin-embedded tissue samples were deparaffinized in xylene and then centrifuged at maximal speed (20,238 × g for 5 min). The supernatant containing the xylene was removed. Xylene was further removed by 2 additional centrifugation steps with ethanol with ensuing removal of ethanol. The resultant pellet was lysed overnight with proteinase K on a thermomixer (55°C). DNA was extracted using a commercial kit
a
following the manufacturer’s instructions. All samples were examined for the presence of DNA of Chlamydiaceae, Parachlamydia, and Waddlia by real-time PCR. The DNA concentration of randomly selected samples was quantified, and DNA content was appropriate for PCR investigation; however, positive controls for each extraction were not performed.
Chlamydiaceae real-time PCR
The extracted DNA was investigated on a real-time PCR system
b
using the 23S-based, Chlamydiaceae family–specific real-time PCR assay described previously.4,14 A 111-bp product, specific for members of the Chlamydiaceae family, was generated as well as a 177-bp product for the internal amplification control. A final volume of 25 µl for each reaction was obtained by adding 2.5 µl of extracted DNA to 12.5 µl of 2× commercial master mix
b
and a final concentration of 500 nM of each primer and probe.
c
The cycling program consisted of an initial denaturation (95°C, 20 sec), followed by 45 cycles of denaturation and amplification (95°C, 3 sec; 60°C, 30 sec). All samples were tested in duplicate. The threshold value was calculated automatically. A sample was considered positive when both values showed a threshold cycle (Ct) of <38, and questionable positive if only 1 value was obtained or the mean Ct value was >38. Samples with questionable positive Ct values were tested again in duplicate. The repeat results were interpreted as positive if the duplicates showed a Ct value of <38. If the mean Ct value was >38 the result was considered questionable positive, and samples without a Ct value were considered as negative. DNA of C. abortus was used as the positive control, whereas a reaction mixture with water was the negative control. Positive or questionable positive samples by real-time PCR for Chlamydiaceae were further examined by the species-specific commercial microarray assay
d
as previously described.
5
16S ribosomal DNA PCR and sequencing
Positive or questionable positive samples by Chlamydiaceae real-time PCR but negative by microarray were further investigated by a 16S ribosomal DNA (rDNA)-based PCR
16
followed by direct sequencing of the obtained PCR product.
37
Sample sequences were compared with those available in GenBank using the BLAST server from the National Centre for Biotechnology (http://www.ncbi.nlm.nih.gov/blast/).
Waddlia real-time PCR
The real-time PCR was performed as previously described
17
using the same PCR machine as for the Chlamydiaceae detection. A 101-bp product within the 16S rRNA gene specific for Waddlia spp. was generated. The cycling program consisted of 3 min at 95°C and 45 cycles of 15 sec at 95°C and 1 min at 60°C. Samples were tested at least in duplicate and were considered as negative if no amplification was observed during all 45 cycles. A Ct value of ≤35.0 was considered as positive, Ct values between 35.0 and 45.0 were considered as questionable positive, and samples without a Ct value as negative. As the positive control, a plasmid containing a partial sequence of the 16S rRNA gene of Waddlia spp. was used. As the negative control, a reaction mixture with water instead of the template DNA was used.
Parachlamydia real-time PCR
The real-time PCR, generating a 103-bp product within the 16S rRNA gene specific for Parachlamydia spp., was performed as previously described
11
using the same conditions as for the Waddlia real-time PCR, with a cycling program of 3 min at 95°C and 45 cycles of 15 sec at 95°C and 1 min at 60°C. The samples were tested as described above. Samples were tested at least in duplicate and were considered as negative if no amplification was observed during all 45 cycles. A Ct value of ≤35.0 was considered as positive, and Ct values between 35.0 and 45.0 were considered as questionable positive. As the positive control, a plasmid containing partial sequence of the 16S rRNA gene of Parachlamydia spp. was applied. As the negative control, a reaction mixture with water instead of the template DNA was used.
Chlamydiales real-time PCR
All Parachlamydia real-time PCR questionable positive samples were further investigated by a 16S rDNA-based Chlamydiales-specific real-time PCR described previously.
23
A product of approximately 207–215 bp (variability in length depends on the species) was generated. The real-time PCR was run in a final volume of 20 µl, obtained by adding to a commercial mastermix,
e
0.1 µM concentrations of each primer,
f
0.1 µM concentration of probe,
f
molecular biology–grade water,
g
and 5 µl of DNA sample. The PCR program started with an initial denaturation (95°C, 3 min), followed by 50 cycles of denaturation, annealing, and extension (95°C, 15 sec, 67°C, 15 sec; 72°C, 15 sec). All samples were tested in duplicate. As the positive control, a recombinant plasmid from Parachlamydia acanthamoebae strain Hall’s coccus was used. As the negative control, a reaction mixture with water instead of the template DNA was used.
Immunohistochemistry
All samples that were positive by PCR for Chlamydiaceae were further investigated by immunohistochemistry (IHC) for the presence of chlamydial antigen in formalin-fixed and paraffin-embedded specimens. A Chlamydiaceae family–specific mouse monoclonal antibody detecting the chlamydial lipopolysaccharide
h
was used. A detection kit
i
was used according to the manufacturer’s instructions. After deparaffinization in xylene and rehydration passing an ethanol-to-water gradient, the antigen retrieval was performed by a 10-min enzyme digestion using proteinase K.
i
The endogenous peroxidase activity was inhibited with peroxidase-blocking solution for 5 min at room temperature (RT), and an incubation for 30 min with the primary antibody (diluted 1:200 in antibody diluent
i
) followed. After incubation with the link-antibody and the horseradish peroxidase–conjugated streptavidin performed at RT for 10 min each, the slides were developed in 3-amino-9-ethyl-carbazole substrate solution for 10 min and counterstained with hematoxylin. A negative control of each section was performed replacing the primary antibody by antibody diluent. The positive controls were from experimentally infected intestinal tissues of gnotobiotic piglets with porcine C. suis strain S45.
18
Results
A total of 115 of the 286 (37.2%, 42/113 litters) investigated fetuses of the reproductive problem group were abortions; the other 171 (62.8%, 71/113 litters) were at-term delivered piglets. Eight of 113 litters (7.1%) were positive for PCV-2 (n = 5) or PPV (n = 3). Five out of 44 fetuses (11%) were positive for PEV (n = 4) and PTV (n = 1), respectively. In 16 of 113 litters (14.2%) with inflammatory lesions, a bacterial etiology was determined: Escherichia coli (n = 6), Streptococcus sp. (n = 3), Actinomyces pyogenes (n = 2), Klebsiella sp. (n = 2), Enterococcus sp. (n = 1), Mycobacterium sp. other than M. tuberculosis complex (n = 1), and Leptospira sp. (n = 1). In a further 14 litters, inflammatory lesions were present (12.4%), but no etiological agent could be demonstrated. All placental smears were negative for Brucella sp.
Malformations and dystocia were present in 4% of litters, and 63% (n = 71) had no indication for an infectious abortion cause. In the control group, no inflammatory lesions could be detected in all 7 litters.
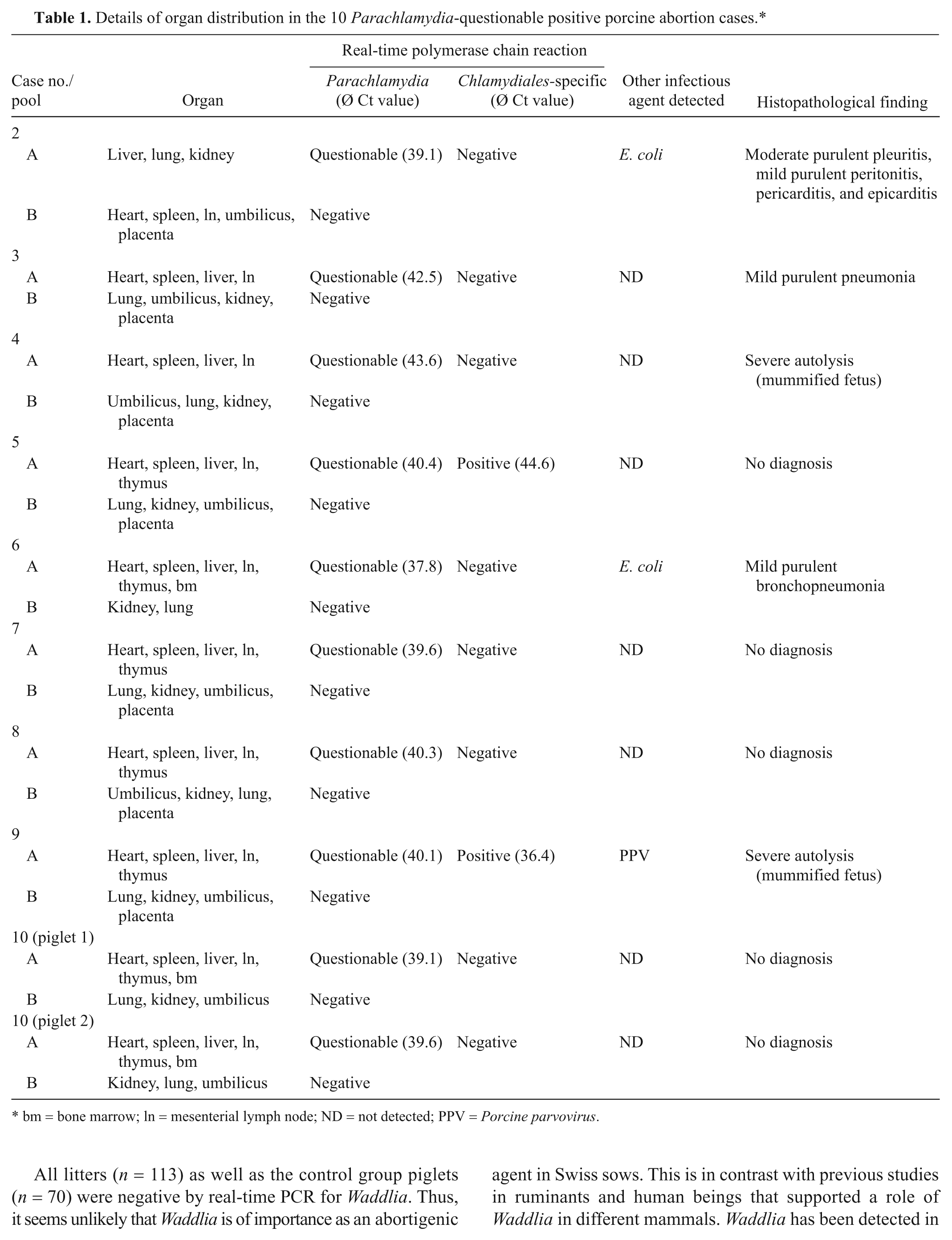
Of 286 fetuses out of 113 swine abortions, 1 fetus of 1 litter originating from central Switzerland was positive (460 g, crown-rump length 21 cm) by real-time PCR for Chlamydiaceae. From this fetus, both paraffin blocks consisting of lung and heart (pool A) and liver, kidney, and brain (pool B) were positive by real-time PCR with a mean Ct value of 36.1 and 37.7, respectively. These samples were further investigated by microarray. In lung and heart (pool A), C. abortus was identified. Liver, kidney, and brain (pool B) were negative by microarray and thus further analyzed by 16S rDNA PCR. The PCR product was positive for C. abortus by sequencing. The IHC test for Chlamydiaceae was negative in all organ samples of this fetus. PCV-2 antigen was detected in heart and liver of this fetus by IHC. Advanced autolysis of the fetus limited the possibility of thorough histopathological assessment. However, meconium was present in the lung alveoli indicative of intrauterine asphyxia. All fetuses of the control group (n = 70) were negative by the investigation for Chlamydiaceae. By the Waddlia real-time PCR, none of the fetuses of the reproductive problem group (n = 286), as well as all of the fetuses of the control group (n = 70), were positive. By the Parachlamydia real-time PCR, 10 out of 286 fetuses of the reproductive problem group were questionable positive. Details of Parachlamydia questionable positive cases are shown in Table 1. In all 10 cases, the Ct values were higher than 37.5, therefore the results were interpreted as questionable positive. Two of the 10 fetuses could be confirmed by a second PCR method targeting the 16S rDNA gene of the order Chlamydiales. One fetus had a mean Ct value of 44.6; the PCR product could not be sequenced. The second fetus had a mean Ct value of 36.4; the PCR product showed closest homology to uncultured Chlamydiales (95.3%) and to uncultured Parachlamydia sp. (94.8%), respectively.
Histological examination showed that 3 of the 10 fetuses showed multiple inflammatory lesions in internal organs, and in 1 case an additional necrotizing placentitis was found. The organs of 2 fetuses could not be interpreted histologically due to severe autolysis and 5 fetuses had no histopathological lesions. Seven of the Parachlamydia questionable positive fetuses were negative for all other investigated abortigenic agents. Another 2 fetuses were also positive for Escherichia coli and, in 1 fetus, a mixed infection with Parachlamydia and PPV was present. In 9 out of 10 fetuses, a Ct value was obtained in pooled organ samples of heart, spleen, liver, mesenterial lymph node, and thymus. In 1 fetus, liver, lung, and kidney (pool B) was questionable positive. All fetuses of the control group (n = 70) were negative by the Parachlamydia real-time PCR.
Discussion
Most cases of porcine abortions and stillbirths remain of unknown etiology.
8
In the present study, 12% of cases had histological lesions such as placentitis and fetal pneumonia indicative for an infectious agent but remained without etiological diagnosis. Common abortigenic agents such as Brucella sp., Leptospira sp., and PPV were excluded or present in only a small number of abortion cases. Noninfectious causes (4%) such as malformation and dystocia were not a frequent finding. However, other noninfectious causes such as high temperature, toxins, and nutritional deficiencies could still be considered but were not investigated in the present study. The need to look for yet uncommon or even unknown abortigenic agents is important. Therefore, the present study investigated uncommon and newly emerging abortigenic agents such as Chlamydiaceae as well as Chlamydia-related bacteria, in particular Waddlia and Parachlamydia in 286 fetuses and their placentae.
A major advantage of the current study was the availability of the placentae in almost all 113 affected litters (n = 99). Under field conditions, corresponding placental tissue cannot be routinely investigated due to maternal behavior (eating the placenta). Placental membranes are the preferred samples to diagnose chlamydial abortion in ruminants.
30
Surprisingly, all placentae (n = 99) tested negative in the investigation for Chlamydiaceae and Chlamydia-like organisms by real-time PCR. Using real-time PCR in combination with microarray or 16S rDNA PCR and sequencing, C. abortus was detected only in the internal organs of 1 out of 286 fetuses (0.35%) in the reproductive problem group and in none of the fetuses of the control group (n = 70).
Abortions due to C. abortus in ruminants have the typical histological features of a purulent and/or necrotizing placentitis with vasculitis. Chlamydial abortion is defined if chlamydial antigen and/or DNA is detected within these lesions by PCR and/or IHC.7,30 In the present C. abortus–positive case, the placenta was not available, and due to advanced autolytic processes, a histological examination of the fetus was not practical. Thus, the above mentioned definition of chlamydial abortion could not be completely fulfilled in this case. Interestingly, the Chlamydia-positive fetus was also positive for PCV-2. It remains unclear if the synergistic effect of both pathogens or only 1 of these caused the stillborn fetus. Similar mixed infections were previously diagnosed in an Estonian pig production farm (C. suis, C. abortus, and PCV-2).
31
Although investigations on the seroprevalence of Chlamydiaceae in sows (63%) suggested that Chlamydiaceae are widespread in Switzerland,
10
this could not be confirmed in the present study in the setting of porcine abortion. In previous studies, C. abortus was detected in a porcine abortion case (1/20, 5%), as well as in 7 uteri (7/21, 33%) of pigs with reproductive problems in Switzerland.
10
However, this prior study investigated only a small number of cases, which limited the conclusions on the prevalence estimates. In a German study, only 2 uteri of sows with reproductive disorders (n = 42) were positive for C. abortus.
20
Pregnant sows experimentally infected with C. abortus orally (n = 2) and intravenously (n = 2) seroconverted but their reproductive performance was not impaired. However, Chlamydiae were identified in the afterbirth from 2 sows using stained smears and 2 different antigen detection tests.
33
Thus, the abortigenic effect of C. abortus in pregnant pigs experimentally infected remains to be proved. A zoonotic potential of C. abortus is proven in sheep and goats.
28
The transmission of C. abortus from pigs to human beings has not been reported so far but cannot be excluded.
31
All litters (n = 113) as well as the control group piglets (n = 70) were negative by real-time PCR for Waddlia. Thus, it seems unlikely that Waddlia is of importance as an abortigenic agent in Swiss sows. This is in contrast with previous studies in ruminants and human beings that supported a role of Waddlia in different mammals. Waddlia has been detected in Switzerland in ruminant abortions (3/343 abortions, 0.9%)
4
and has been related to human miscarriage.
3
The current study revealed real-time PCR questionable positive results for Parachlamydia in swine abortions. In total, 10 out of 286 fetuses (3.5%) were questionable positive by real-time PCR. Parachlamydia was already detected in bovine (13.4%, 46/343),
4
ovine, and caprine abortions (0.9%, 2/211).
29
In the present investigation, 2 coinfections with E. coli could be detected, as well as a mixed infection with PPV. In these cases, it remains unclear which pathogen caused the abortion. Previous studies have described mixed infections with Parachlamydia and Waddlia as well as with Parachlamydia and Neospora caninum and even with C. abortus in bovine abortions.
4
Double infections with Parachlamydia and C. abortus in ovine abortions have been reported as well.
29
In conclusion, it remains unclear if Parachlamydia can be considered as a new abortigenic agent in pigs, as very high Ct values were obtained by real-time PCR (ranging from 37.8 to 43.6). Furthermore, only in 2 fetuses could the PCR products be confirmed by the Chlamydiales-specific real-time PCR. False-negative results in the second PCR might be due to advanced autolysis resulting in increased DNA double-strand breaks that make it difficult to detect DNA fragments longer than 200–300 base pairs. Parachlamydia is discussed as an agent of pneumonia in human beings and has been associated with human miscarriages and was found serologically in young men in Switzerland.1,2 However, direct transmission between animal and human being has not yet been demonstrated, and therefore its potential as a zoonotic agent needs to be investigated.