Abstract
In the current study, molecular, biological, and antigenic analyses were performed to characterize Border disease virus (BDV) strain FNK2012-1 isolated from a pig in 2012 in Japan. The complete genome comprises 12,327 nucleotides (nt), including a large open reading frame of 11,685 nt. Phylogenetic analysis revealed that FNK2012-1 was clustered into BDV genotype 1 with ovine strains. FNK2012-1 grew in porcine, bovine, and ovine primary cells and cell lines, but grew better in bovine and ovine cells than in porcine cells. Specific pathogen–free pigs inoculated with FNK2012-1 did not show any clinical signs. Noninoculated contact control pigs also did not show clinical signs and did not seroconvert. The results suggest that FNK2012-1 may be of ruminant origin and is poorly adapted to pigs. Such observations can provide important insights into evidence for infection and transmission of BDV, which may be of ruminant origin, among pigs.
Keywords
Border disease (BD) is a viral disease in sheep and goats that is characterized by transient depression, pyrexia, and leukopenia; however, clinical signs are generally mild or not apparent. 28 Border disease virus (BDV) infections are similar in many aspects to Bovine viral diarrhea virus (BVDV) infections, as vertical transmission plays an important role in the epidemiology of BD. 28 Fetal infection in early pregnancy, before the onset of immune competence, can result in the birth of persistently infected (PI) animals. 28 These PI animals are viremic, antibody negative, constantly excrete virus, and are more susceptible to secondary disease; however, they may be clinically normal. 28 Thus, PI animals are the most potent source of BD infection.
Border disease virus is classified into the genus Pestivirus of the family Flaviviridae and is closely related to BVDV genotypes 1 and 2 and Classical swine fever virus (CSFV). 11 As all pestiviruses are antigenically closely related to one another, polyclonal antisera against pestiviruses generally fail to distinguish species when techniques such as fluorescent antibody testing or immunodiffusion are used. 28 While CSFV infection is predominantly restricted to pigs, it is not uncommon for BDV and BVDV to be transmitted between species. Pigs can be infected by both BDV1,12,18 and BVDV.12,19
In swine, CSFV may be associated with acute or chronic clinical disease resulting in high morbidity and mortality, causing massive economic losses. 11 In contrast, infection by BDV and BVDV in pigs usually results in mild clinical or subclinical signs, allowing the viruses to spread undetected through immune-naïve animals.10,18 The direct economic losses associated with BDV or BVDV infection of pigs are minimal; however, such infections may interfere with serologic surveillance and diagnosis of CSFV infection due to cross-reactions between pestiviruses. 10
In Japan, although an epidemiological survey of BDV among sheep documented serological evidence of virus circulation, no clinical cases of BD or PI lambs with BDV were reported. 2 In February 2012, a pestivirus was isolated from a healthy fattening pig kept at a sow–farrow-to-finish farm in Ibaraki Prefecture. No ruminants were allocated in the farm. Virus was detected in the course of a serological surveillance of CSFV, and was diagnosed as BDV. 7 In the current study, molecular, biological, and antigenic analyses were performed in order to characterize the BDV strain FNK2012-1.
The FNK2012-1 strain was isolated from white blood cells of a clinically healthy pig using a cloned porcine kidney (CPK) cell line and propagated by passage twice.7,9 Seed virus was propagated using the swine kidney cell line L (SK-L) by passaging. Primary swine kidney (SKp), bovine testis (BT), bovine fetal muscles (BFM), and ovine kidney (OK) cells were prepared from pestivirus-free animals and were used for virus growth within 10 passages. Primary cells were grown as a monolayer in Eagle minimum essential medium (MEM)
a
containing 0.075 mg of sodium bicarbonate per ml, 0.292 mg of L-glutamine per ml, inactivated 10% fetal bovine serum, and antibiotics. The porcine kidney cell lines (CPK and a purified porcine kidney 3F [PPK3F]) and the bovine kidney cell lines (Madin–Darby bovine kidney [MDBK] and MDBK-SY) were confirmed to be free of pestiviruses and were also used for virus growth.13,17,21 Cell lines were grown as a confluence in MEM containing 0.295% tryptose phosphate broth,
b
1.125 mg of NaHCO3 per ml, 0.292 mg of
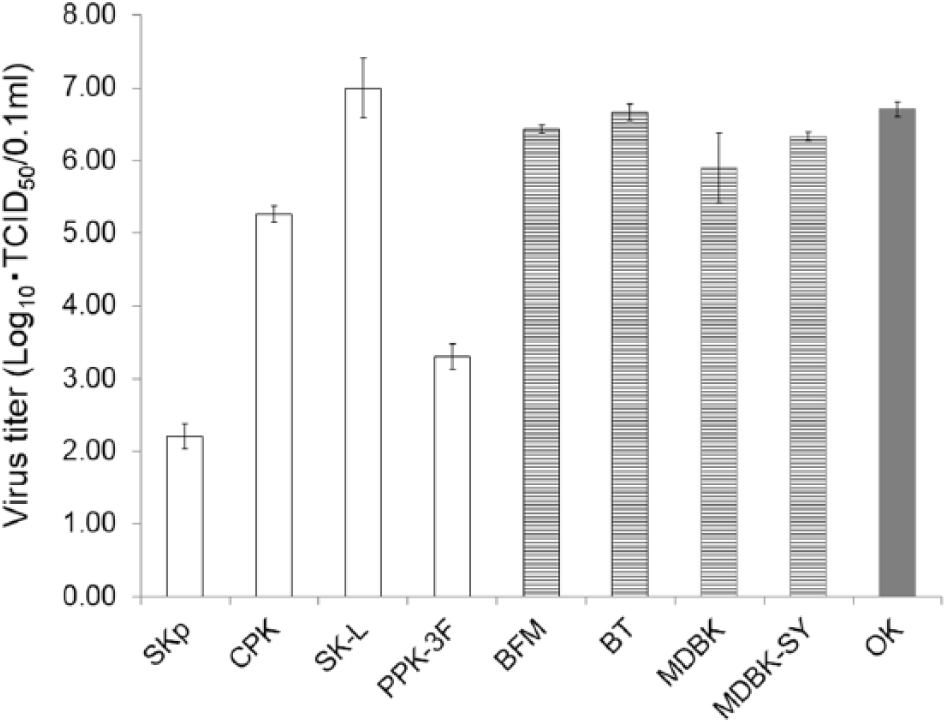
Virus titers of Border disease virus strain FNK2012-1 in primary cells and cell lines of swine (open box), bovine (striped box), and ovine (gray box) at 5 days postinoculation. Cells were infected with FNK2012-1 at a multiplicity of infection of 0.1. Mean values from 3 independent experiments are shown.
Viral RNA was extracted from virus-infected SK-L cells using a commercially available kit. c Next-generation sequencing and data analysis were performed as previously described. 14 The nucleotide sequences of the 5′- and 3′-termini were determined using the rapid amplification of complementary (c)DNA ends method.3,6 Nucleotide sequences were aligned with other pestivirus strains using ClustalW, and phylogenetic analyses were performed by the neighbor-joining method using MEGA5.22.22,25,27
The complete genome of FNK2012-1 is 12,357 nucleotides (nt) in length. The 5′- and 3′-untranslated regions are 374 and 265 nt long, respectively. The full genomic sequence of FNK2012-1 was deposited in the DNA Data Bank of Japan–European Molecular Biology Laboratory–GenBank database under the accession number AB897785. The phylogenetic trees constructed for the entire Npro and E2 genes of pestiviruses are shown in Figure 2A and 2B, respectively. FNK2012-1 was clustered into BDV-1 genotype with ovine strains BD-31 and X818, supported by high bootstrap values. The entire Npro gene sequence of FNK2012-1 shares 88.8%, 88.0%, and 79.4% identity with BD-31, X818, and 87/6, respectively. For the E2 genes, sequence identity between FNK2012-1 and BD-31, X818, and 87/6 was 85.4%, 85.4%, and 75.3%, respectively.
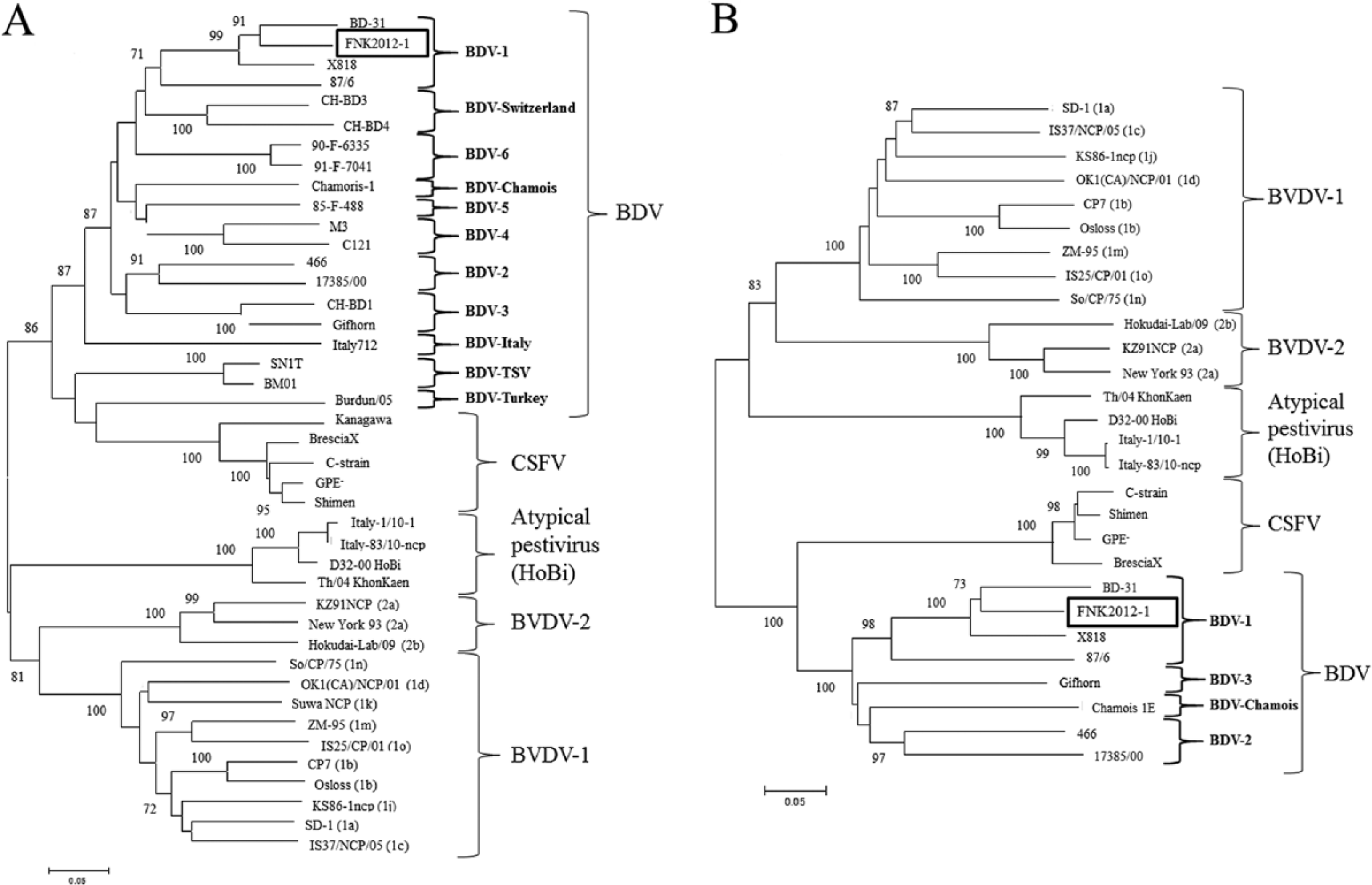
Phylogenetic analysis based on nucleotide sequences of Border disease virus strain FNK2012-1. Phylogenetic trees were constructed for the entire Npro (
In order to assess the pathogenicity of FNK2012-1 in pigs, four 6-week-old crossbred (Landrace × Duroc × Yorkshire) female specific pathogen–free (SPF) pigs (nos. 1–4) were inoculated intranasally by nasal spray with 2 ml of 106.0 TCID50/ml virus from the cell culture supernatant. Furthermore, 2 SPF pigs (nos. 5 and 6) were placed in contact with inoculated pigs at 1 day postinoculation (dpi). Two inoculated pigs (nos. 3 and 4) were euthanized at 5 dpi, and tissues from brains, tonsils, spleens, adrenal glands, kidneys, mesenteric lymph nodes, and colons were collected aseptically. Nasal swabs and whole blood were collected at 0, 3, and 5 dpi. The remaining 2 infected pigs and 2 contacted pigs were monitored for rectal temperature and clinical signs over a period of 37 days. Swabs and blood were collected at 0, 3, 5, 7, 10, 15, 22, 29, and 37 dpi. Sera were collected for detection of antibodies against FNK2012-1 using a commercial CSFV antibody enzyme-linked immunosorbent assay (ELISA) kit d and a neutralization test using FNK2012-1 at 0, 5, 7, 15, 22, 29, 37, and 49 dpi. Total white blood cells and platelets were counted using a commercially available kit. e All animal experiments were performed in a biosafety level 2 facility at the Graduate School of Veterinary Medicine, Hokkaido University, Sapporo, Japan. The Institutional Animal Care and Use Committee of the Graduate School of Veterinary Medicine authorized this animal experiment (approval numbers: 13-0154), and all experiments were performed according to the guidelines of this committee.
None of the inoculated pigs or directly contacted pigs showed fever, leukocytopenia, thrombocytopenia, or any other clinical symptoms (data not shown). The ELISA was used to detect antibodies against FNK2012-1 in serum from experimentally infected pigs, as antibodies against BVDV and BDV may cross-react with this ELISA. A neutralization test used for detection of anti-BDV antibodies in the experimental infection and cross-neutralization test was conducted by the microtitration methods described in the OIE Manual. 28 Briefly, serial 2-fold dilutions of sera were prepared in a volume of 0.025 ml in a microplate with cell culture media, mixed with an equal volume of virus suspension containing 200 TCID50 and incubated at 37°C for 1 hr. Next, 0.1 ml of SK-L cells (for BDV and CSFV) and MDBK cells (for BVDV) suspension (approximately 2 × 105 cells/ml) were added into all the wells of the microplate, and were incubated at 37°C under 5% CO2 for 4 days. Plates were stained using an immunoperoxidase staining system, as described by the OIE Manual. 28 Neutralizing antibody titer was expressed as the reciprocal of the highest serum dilution that showed complete viral neutralization.
Inoculated virus was recovered from the mesenteric lymph nodes of inoculated pigs (no. 3) at 5 dpi. All inoculated pigs seroconverted to FNK2012-1, but contacted pigs did not at 49 dpi. Seroconversion against FNK2012-1 was detected in all inoculated pigs at 22 dpi and 29 dpi by neutralization and ELISA, respectively (Fig. 3).
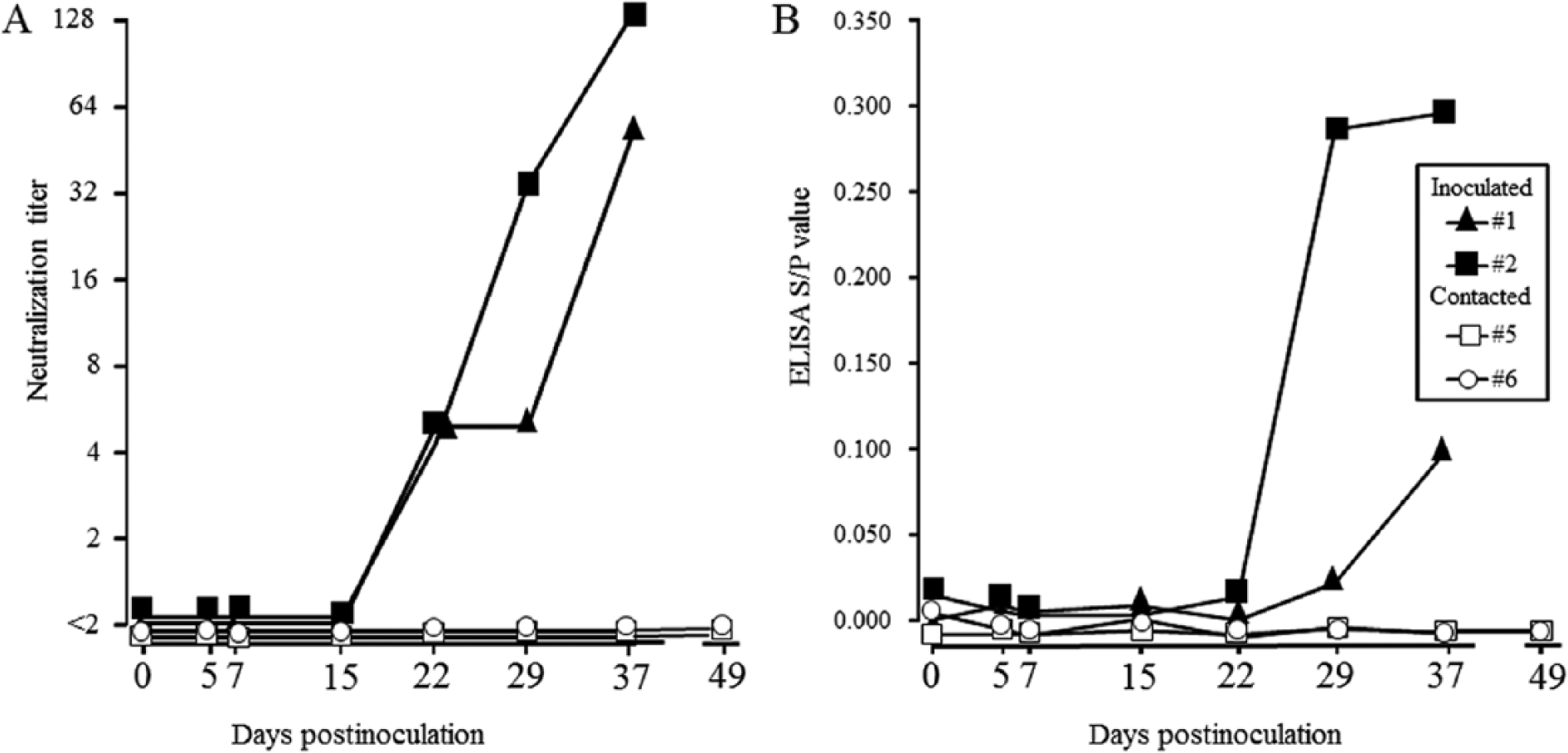
Profiles of virus neutralization titers against Border disease virus strain FNK2012-1 (
Six strains (CSFV GPE–, CSFV Kanagawa/74, BVDV-1 Nose, BVDV-2 KZ-91-NCP, atypical bovine pestivirus D32/00_HoBi [HoBi], and BDV 87/6) were propagated by SK-L cells (GPE–, Kanagawa/74, and 87/6) and MDBK cells (Nose, KZ-91-NCP, and Hobi), and used for neutralization test as representative pestivirus strains.5,8,16,18,23,24 Antisera against CSFV GPE–, CSFV Kanagawa, BVDV-1 Nose, BVDV-2 KZ-91-NCP, BDV FNK2012-1, and BDV 87/6 were prepared with pigs by intravenous injection of virus suspensions, as described previously. 23 Antiserum against HoBi was prepared with rabbits by repeated intramuscular injection of virus suspensions with adjuvant component. Sera were stored at −20°C, and were heat-inactivated at 56°C for 30 min before use.
Antibody titer against each strain is shown in Table 1. Each antiserum showed the highest neutralization titer against homologous strains. Significant antigenic differences between BDV, BVDV-1, BVDV-2, atypical bovine pestivirus (HoBi), and CSFV were observed. Interestingly, the neutralization titers of antisera against FNK2012-1 to 87/6, and 87/6 to FKN2012-1 were significantly lower than those against homologous virus, despite being classified in the same genotype, BDV-1.
Cross-neutralization titers of antisera raised different strains of pestivirus.*
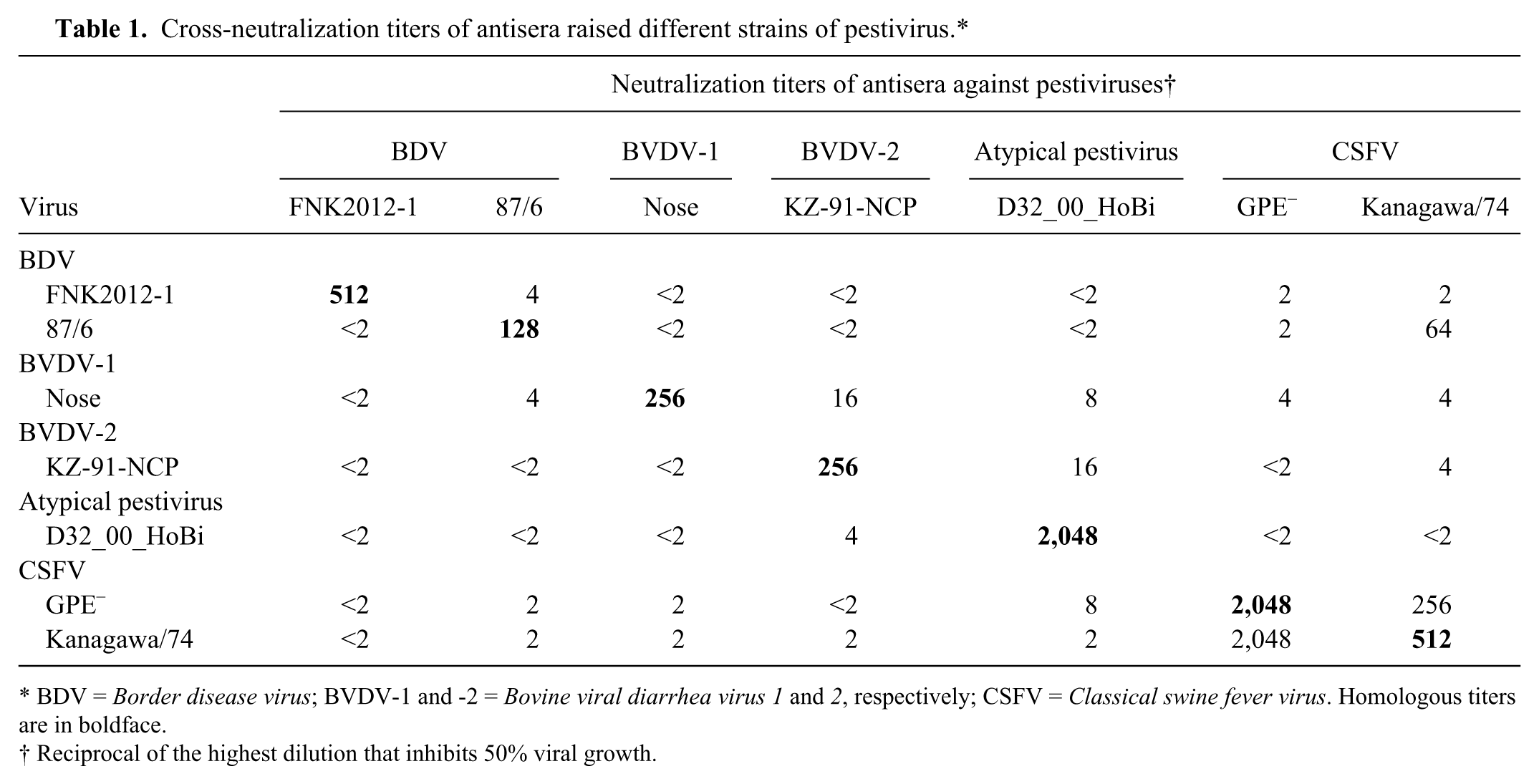
BDV = Border disease virus; BVDV-1 and -2 = Bovine viral diarrhea virus 1 and 2, respectively; CSFV = Classical swine fever virus. Homologous titers are in boldface.
Reciprocal of the highest dilution that inhibits 50% viral growth.
Apart from SK-L cells, the FNK-2012-1 titer in porcine cells was lower than in bovine and ovine cells. Therefore, FNK2012-1 may be of ruminant origin. Further investigation is needed in order to clarify the source of FNK2012-1 in ruminants.
The whole genome of BDV FNK2012-1 was 12,357 nt; the open reading frame was 11,688 nt, coding 3,896 amino acids. No insertions of host or viral genome or deletions were found. Phylogenetic analyses of Npro, E2 (Fig. 2), 5′-untranslated region, and NS5B (data not shown) revealed that FNK2012-1 grouped within BDV-1 with ovine strains. Ovine strain L73/74 from Europe had the most sequence similarity with FNK2012-1 in the GenBank database, and nucleotide sequence identities for the Npro and E2 were 90.0% and 86.0%, respectively. The relatively low homology with other BDV strains limited further inferences about the virus origin.
In the current study, pigs inoculated with FNK2012-1 did not develop clinical disease, fever, leukopenia, or thrombocytopenia. Viremia was not observed but the inoculated virus was recovered from an inoculated pig and subsequent seroconversion was demonstrated in inoculated pigs. However, pigs put in contact with inoculated pigs with FNK2012-1 remained seronegative. Viremia in BDV experimentally infected pigs and in horizontal transmission studies have been reported by some authors.1,18 It is suggested that FNK2012-1 strain was recently introduced into swine populations, and has not yet developed the ability to efficiently spread in swine populations.
The reactivity of pestivirus isolates with defined serological reagents represents a further important criterion for demarcation of the species. 26 Significant antigenic differences were observed between FNK2012-1 and 87/6 within BDV-1. Thus, the similarities of genomic region encoding for the entire E2 of these strains was compared, as the viral structural envelope protein E2 has been demonstrated to be the main target of neutralizing antibodies. 11 The nucleotide and amino acid sequence similarity in E2 between FNK2012-1 and 86/7 were only 75.3% and 78.0%, respectively. As reported, comparison of genomic sequences encoding for E2 is thus useful for predicting antigenic differences among pestivirus. 15
In conclusion, FNK2012-1 grew better in bovine and ovine cells than in porcine cells and induced no clinical disease or viremia in experimental infected pigs, suggesting a ruminant origin. While BDV infection in pigs does not, to date, have a major economic impact, the spread of BDV in swine populations affects serological surveillance of CSFV. The data from the current study provides important evidence on the epidemiological basis of BDV in pig populations.
Footnotes
Acknowledgements
Makoto Nagai, Hiroshi Aoki, and Yoshihiro Sakoda contributed equally to this work. The authors are grateful to Kenhoku Ibaraki Prefectural Livestock Hygiene Service Center for providing BDV FNK2012-1 strain and related information. MDBK-SY was kindly provided by the Tochigi Prefectural Central Livestock Hygiene Service Center. The authors are also grateful to Dr. Horst Schirrmeier (Friedrich-Loeffler-Institut, Germany) and Dr. D. J. Paton (Animal Health and Veterinary Laboratories Agency, UK) for providing atypical bovine pestivirus D32_00 “HoBi” and Border disease virus 87/6.
a.
Nissui, Tokyo, Japan.
b.
BD Biosciences, San Jose, CA.
c.
ISOGEN-LS, Nippongene, Toyama, Japan.
d.
JNC Corp., Tokyo, Japan.
e.
pocH-100iV Diff apparatus, Sysmex, Hyogo, Japan.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
