Abstract
Analysis of rumen contents is helpful in solving poisoning cases when ingestion of a toxic substance by cattle or other ruminant animals is suspected. The most common technique employs extraction of the sample with organic solvent followed by clean-up method(s) before analysis with gas chromatography–mass spectrometry equipped with a library of mass spectra to help identify unknowns. A rapid method using magnesium sulfate, primary secondary amine, and C18 sorbents following principles of QuEChERS to clean up rumen contents samples is reported herein. The method was validated to analyze fortified bovine rumen contents to detect commonly found organophosphorus pesticides, carbamates, and several other compounds such as atropine, 4-aminopyridine, caffeine, scopolamine, 3-chloro-4-methylaniline, strychnine, metaldehyde, and metronidazole. For each compound, the ratio of 2 ions from the mass spectrum was monitored in fortified rumen contents. The ion ratio of fortified sample was compared with the ion ratio of standard sample spectrum and was found to be within 20%, with the exception of aldicarb and 4-aminopyridine with ion ratio of 26% and 29%, respectively. Usefulness of the method was demonstrated by not only analyzing bovine rumen contents but also canine and avian gastrointestinal contents submitted for organic chemical screening.
Keywords
Introduction
Millions of pounds of pesticides are used worldwide annually to safeguard crops and to improve farm productivity (Grube A, Donaldson D, Kiely T, Wu L: 2011, Pesticides industry sales and usage: 2006 and 2007 market estimates. U.S. Environmental Protection Agency, Washington, DC. Available at: http://www.epa.gov/opp00001/pestsales/07pestsales/table_of_contents2007.htm#pdfreport). Such products are evaluated for their safety before they can be sold for use and are considered safe to human beings and animals, if label directions are followed. However, by nature these pesticides are designed to kill or otherwise adversely affect living organisms. Each year, livestock (cattle, sheep, swine, and horses) and wildlife are killed by consumption of pesticides accidentally spilled on the ground or left on trucks and wagons. Animals may die after eating the pesticide alone, when it is accidentally mixed with feeds, or by coming into contact with contaminated surfaces and treated areas. These compounds have also been implicated in intentional poisonings.
Proper identification of toxicant(s) when making a diagnosis of livestock poisoning is essential in the prevention of future losses and to stop contaminated food products from reaching the market. A veterinary diagnostic toxicology laboratory is routinely presented with samples collected from dead animals for the purpose of identifying the cause of an unexpected death. Liver and/or ingesta are most commonly submitted for analysis. Although liver is the main site of metabolism for many drugs and toxicant residues, the analysis of recovered feed, bait, and either rumen or stomach contents can often better support a diagnosis of toxicant exposure. This can be expected as most xenobiotics are only partially absorbed from the gastrointestinal (GI) tract. The absorbed amount produces its deleterious effects at the target site(s) with sometimes only a fraction of the parent chemical residue remaining in the liver postexposure.
In cases where a limited history makes it difficult to test for either a particular toxicant or even a single class of chemical compounds, the best approach is to utilize a general screening method that is able to detect a wide variety of toxicants in a single analysis. With the increased availability and high sensitivity of liquid chromatography–mass spectrometry (LC-MS), a number of publications have described methods to screen samples for a variety of toxicants. However, the lack of well-developed spectral libraries for LC-MS systems limits its use in quickly solving cases involving unknown toxicants exposures. The routine analysis of such samples is carried out with gas chromatography–mass spectrometry (GC-MS) equipped with a library containing mass spectral data for thousands of compounds that facilitates detection by providing a spectral match. A potential match can be confirmed by injection of analytical standards and demonstrating matching spectra and retention time.
In cases utilizing wide-scope screening, it is important that the method used to clean up the sample retains the toxicants to confirm its presence or absence in the submitted sample. A number of articles have been published that describe general screening methods for the analysis of pesticides in feed using GC-MS.1–3,5,7,9,11–13 However, the typical bovine diet of grain, hay, silage, and mineral supplements results in the presence of a complex mixture of digested and undigested feed components and chemical residues in the rumen contents. The methods developed for feed materials for grain cannot be assumed to be also suitable for rumen contents. Rumen contents contain significant amounts of plant material such as hay and silage in addition to partially digested grain and compounds produced by rumen bacteria. Analysis of feed samples by GC-MS (results not shown) exhibits very different spectra as compared with rumen contents (Fig. 1; Table 1). Since the first published application of the QuEChERS (quick, easy, cheap, effective, rugged, and safe) method for the analysis of pesticides in produce, this method has gained usefulness in pesticide analysis in feed,2,5 –7,11 biological fluids, and tissue samples.4,8,10,14 In the current study, a QuEChERS clean-up method followed by GC-MS analysis using a library search for unknowns has been used for the analysis of bovine rumen contents to screen for a variety of toxicants commonly encountered in veterinary toxicology laboratories. Using the principles of QuEChERS, method purification of extracted samples was achieved that could replace standard methods such as liquid-liquid or solid-phase extraction. Initially, 18 control rumen content samples from healthy animals were analyzed to identify compounds normally found in the rumen contents, which then facilitated the identification of xenobiotic compounds in a submitted diagnostic sample. A selected list of toxicants, based on the clinical cases submitted to the authors’ laboratory, was screened using this method. The identity of each compound was confirmed by comparing the retention time of each analyte to a standard and the mass spectrum to that of the compound as found in the spectral library. Control rumen contents were spiked to determine the method detection limit of each toxicant.
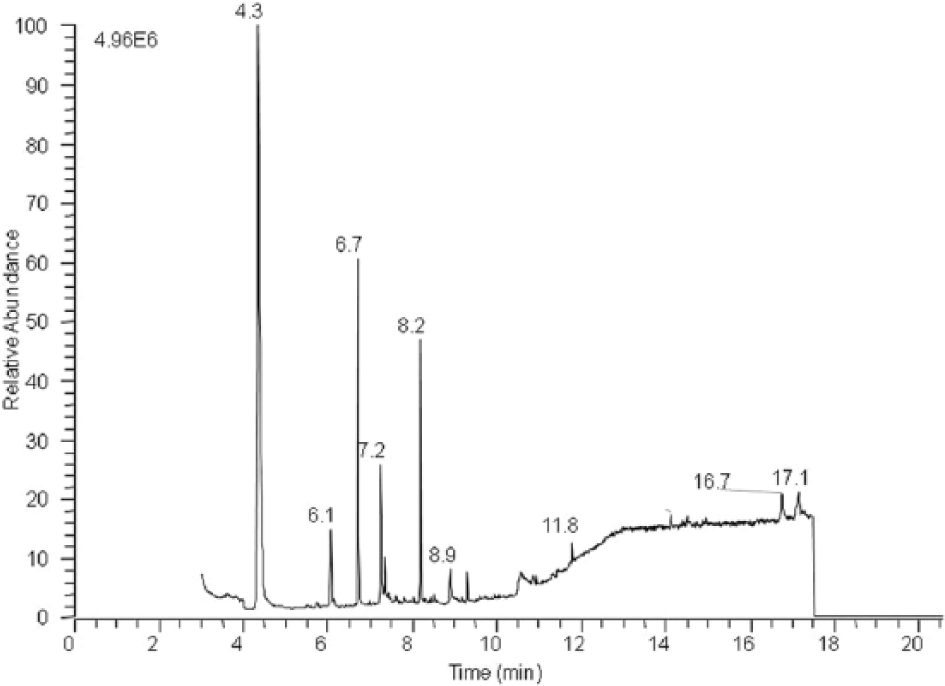
Representative chromatogram of a control rumen contents.
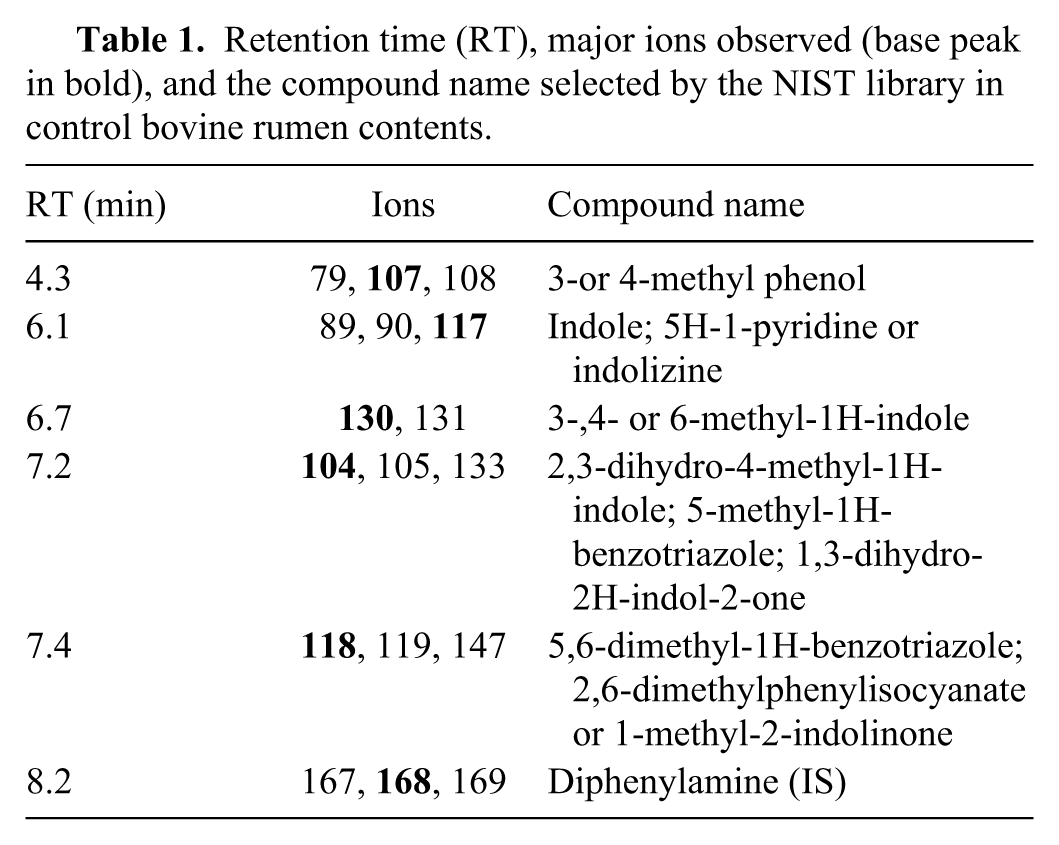
Retention time (RT), major ions observed (base peak in bold), and the compound name selected by the NIST library in control bovine rumen contents.
Materials and methods
Standards and reagents
Standard solutions used for the current study were prepared using high-performance liquid chromatography–grade methanol, a and solutions of 10 mg/ml of each carbamate (aldicarb, b 3-hydroxycarbofuran, b propoxur, b bendiocarb, b carbaryl, b carbofuran, b carbosulfan, b and methiocarb b ) were prepared by dissolving 10 mg of the neat powder in 1 ml of methanol. A solution containing all 8 carbamates was prepared by mixing 100 µl of each 10 mg/ml solution with 200 µl of methanol. The working standard solutions of carbamates used for checking instrument response and for fortifying rumen contents were prepared at 1, 10, 50, 100, and 500 µg/ml by serial dilution of 1 mg/ml mixed standard.
Ten mg/ml standard solutions of individual organophosphorus pesticides (chlorpyrifos, b diazinon, b disulfoton, b famophos, b fenthion, b malathion, b methyl parathion, b tebupirimphos, b and terbufos b ) including amitraz, b were prepared by dissolving 10 mg of neat powder of each pesticide into 1 ml of methanol. Subsequently, 100 µl of each standard was mixed to give 1 mg/ml of mixed standard containing amitraz and organophosphorus pesticides. A serial dilution of 1 mg/ml was performed using methanol to provide 1, 10, 20, and 100 µg/ml solutions to check instrument response and to fortify the rumen contents.
Standard solutions of atropine, c metronidazole, c naphthalene, c scopolamine, c 3-chloro-4-methylaniline, c strychnine, c 4-aminopyridine, a caffeine, a and metaldehyde c were also prepared by dissolving 10 mg of each of the individual neat powder in 10 ml of methanol. A solution containing 1 mg/ml each of atropine, caffeine, scopolamine, 3-chloro-4-methylaniline, and naphthalene was prepared by mixing 100 µl of each 10 mg/ml individual standard with 500 µl of methanol and then diluted further with methanol to provide 5, 50, 100, and 200 µg/ml standard solutions used for this study. The solutions containing 1 mg/ml each of 4-aminopyridine, aldicarb, metaldehyde, metronidazole, and strychnine were prepared by mixing 10 mg/ml individual standard solutions with 500 µl of methanol. The standards containing 200 and 400 µg/ml of 4-aminopyridine, aldicarb, metaldehyde, and strychnine were prepared by serial dilution of 1 mg/ml standard, containing all 5 compounds, with methanol.
Rumen samples
The blank rumen content samples were obtained from diagnostic laboratory necropsy specimens submitted to the Pennsylvania Animal Diagnostic Laboratory System (Kennett Square, Pennsylvania) or from a local slaughterhouse and stored frozen at −20°C until analysis. A blank rumen content sample mixture was prepared by mixing 5 individual blank rumen content samples and was used for the fortification study. This mixture was tested prior to fortification to ensure the absence of any analyte in this study.
Rumen contents (1 g) were spiked at various concentrations with 1 of the following 4 mixes: 1) carbamates mix (aldicarb, 3-hydroxycarbofuran, propoxur, bendiocarb, carbaryl, carbofuran, carbosulfan, and methiocarb) at 1, 2, 5, 10, and 20 µg/g using 100 µl of 10, 20, 50, 100, and 200 µg/ml carbamate standard solution; 2) organophosphosphorus pesticides mix (chlorpyrifos, diazinon, disulfoton, famophos, fenthion, malathion, methyl parathion, tebupirimphos, and terbufos) with amitraz, at 1, 2, and 10 µg/g using 100 µl of 10, 20, and 100 µg/ml organophosphosphorus pesticides standard solution containing amitraz; 3) miscellaneous low mix: caffeine, naphthalene; 3-chloro-4-methylaniline, atropine, and scopolamine mix at 5, 10, and 20 µg/g using 100 µl of 50, 100, and 200 µg/ml standard solution containing the 5 miscellaneous compounds; and 4) miscellaneous high mix: 4-aminopyridine, aldicarb, metaldehyde, metronidazole, and strychnine at 20, 40, and 100 µg/g using 100 µl of mixed standard solution containing 200, 400, and 1000 µg/ml of 4-aminopyridine, aldicarb, metaldehyde, metronidazole, and strychnine.
Equipment and analytical conditions
The analysis was performed using GC-MS d with a 30 m × 0.250 mm column e and National Institute of Standards and Technology (NIST) library f to identify compounds. The GC oven temperature was held at 70°C for 1 min and then ramped to 290°C at the rate of 20°C/min. The oven temperature was maintained at 290°C for the next 8.5 min. Helium was used as carrier gas at a flow rate of 1 ml/min, and 1 µl of sample was injected into the system for analysis. The inlet temperature was maintained at 200°C, and the ion source temperature was set at 230°C. The MS was scanned in 3 segments: full scan mode of 3–5 min in the 40–450 atomic mass unit (amu) range, 5–18 min in the 50–650 amu range, and tandem mass spectrometry (MS/MS) mode from 18 min. The MS/MS conducted on ion m/z 334 was for analysis of strychnine with 30% energy at a voltage of 1 and excitation energy of 0.225, with product ion scan of 200–340 amu; the ions extracted were 306, 277, and 220 amu.
Sample treatment
A 5-g portion of rumen contents was mixed with 10 ml of acetonitrile a and 0.2 g of sodium chloride a by vortexing for 15 sec. The mixture was sonicated for 15 min at 23 ± 2°C to extract analytes in organic solvent. The sample was centrifuged for 8 min at 450 rcf, and 0.8 ml of the supernatant was transferred to a centrifuge tube containing MgS04 a (175 mg), primary secondary amine c (PSA; 50 mg), and C18 sorbent c (50 mg). The sample was vortexed for 15 sec, allowed to sit for 5 min, and revortexed for 15 sec. The sample was centrifuged for 8 min at 16,060 rcf, and the supernatant was filtered using a 0.22-µm nylon filter. The clear filtrate was transferred to a vial to be analyzed by GC-MS.
Results
To obtain the best conditions for the determination of various toxins, sorbents such as PSA, C18, and charcoal were tested by analyzing control rumen contents fortified with 1–40 µg/g of carbamates, organophosphorus pesticides, or miscellaneous compounds. The chosen amount of 5 g of rumen contents was found to be adequate to give a proper representation of the sample, and 10 ml of acetonitrile was sufficient for the extraction of the desired pesticides. Each toxicant was tested at a minimum of 3 concentrations to determine the method detection limit for the toxicant using the method described herein.
Because the goal was to develop a simple method, the samples were vortexed after the addition of acetonitrile and sodium chloride and sonicated to extract out the toxicants. Rumen samples do not require homogenizing or grinding, which avoids cross contamination. Different QuEChERS sorbents such as MgSO4, PSA, C18, basic alumina, Florisil (magnesium silicate), a and/or charcoal were tested to clean up the extract, and the qualitative detection of the compounds was monitored. Inclusion of charcoal c in the sorbent mix gave cleaner extracts but even a small amount of it in the sorbent mix resulted in partial or complete loss of analytes. The best recoveries were obtained by using a mixture of 0.2 g of MgSO4 and 0.05 g each of PSA and C18.
The analysis of rumen content samples obtained from 20 different healthy cattle exhibited very similar profiles by GC-MS. The identity of common matrix compounds in the rumen contents was assigned based on identification made by NIST library–equipped GC-MS, and the compounds are listed in Table 1. The most commonly found compounds are methyl phenol (retention time [RT] = 4.3 min) and unsubstituted (RT = 6.1 min) and substituted indoles (RT = 6.7 and 7.2 min) with diphenylamine used as a surrogate compound. Diphenylamine was added to the samples along with acetonitrile used for extraction. Optimum recovery of diphenylamine was monitored to ensure extraction of analytes of interest from the sample. A representative chromatogram of the rumen contents is shown in Figure 1.
The GC-MS analysis was carried out in 3 segments, the first being a short segment of 3–5 min with scan range of 40–450 amu to capture early-eluting 4-aminopyridine and metaldehyde. The first segment scan range was lowered to 40 amu to capture metaldehyde, which has its major ion at m/z = 45. The second segment of 5–18 min with mass range of 50–650 amu was set for the analysis of the rest of the compounds with the exception of strychnine. To reduce the background noise that could impede detection, the lowest mass was set at m/z of 50 in the second segment, where most of the analytes eluted. To overcome poor sensitivity of strychnine, a third segment was set, and the detection was switched from full scan mode to MS/MS at 18 min. The collision energy of 30% was used to fragment the 334 amu parent ion of strychnine from which product ions of 200–340 amu were scanned and ions 306, 277, and 220 were extracted.
The detection of carbamates, organophosphorus pesticides, and miscellaneous compounds listed in Table 2 was monitored by spiking control rumen contents at a minimum of 3 concentrations. As a group, organophosphorus pesticides and amitraz were found to be the most sensitive and could be detected at the lowest concentration of 1 ppm; most of the carbamates and miscellaneous compounds could be detected at 5 ppm with a few exceptions. For metaldehyde, 3-chloro-4-methylaniline, and scopolamine, the lowest detection limit was established at 20 ppm, whereas strychnine and aldicarb could not be detected below 40 ppm. For each compound, 3 ions were used for spectral identification and the ratio of the 2 ions, shown in Table 2, was used to confirm the presence of each toxicant. The first ion chosen was the most abundant ion and the second ion was either the next most abundant ion or an ion unique to the compound. The parent ion was absent for many of these compounds but when detected, it was used in monitoring the ion ratio. A representative chromatogram for the analysis of fortified rumen contents, with organophosphorus pesticides and amitraz, is shown in Figure 2. The expected ion ratio was calculated using the analysis of standards for each compound. That ratio was compared with that of fortified rumen and was within 20% for all compounds with the exception of aldicarb and 4-aminopyridine where the ion ratio difference was 26% and 29%, respectively.
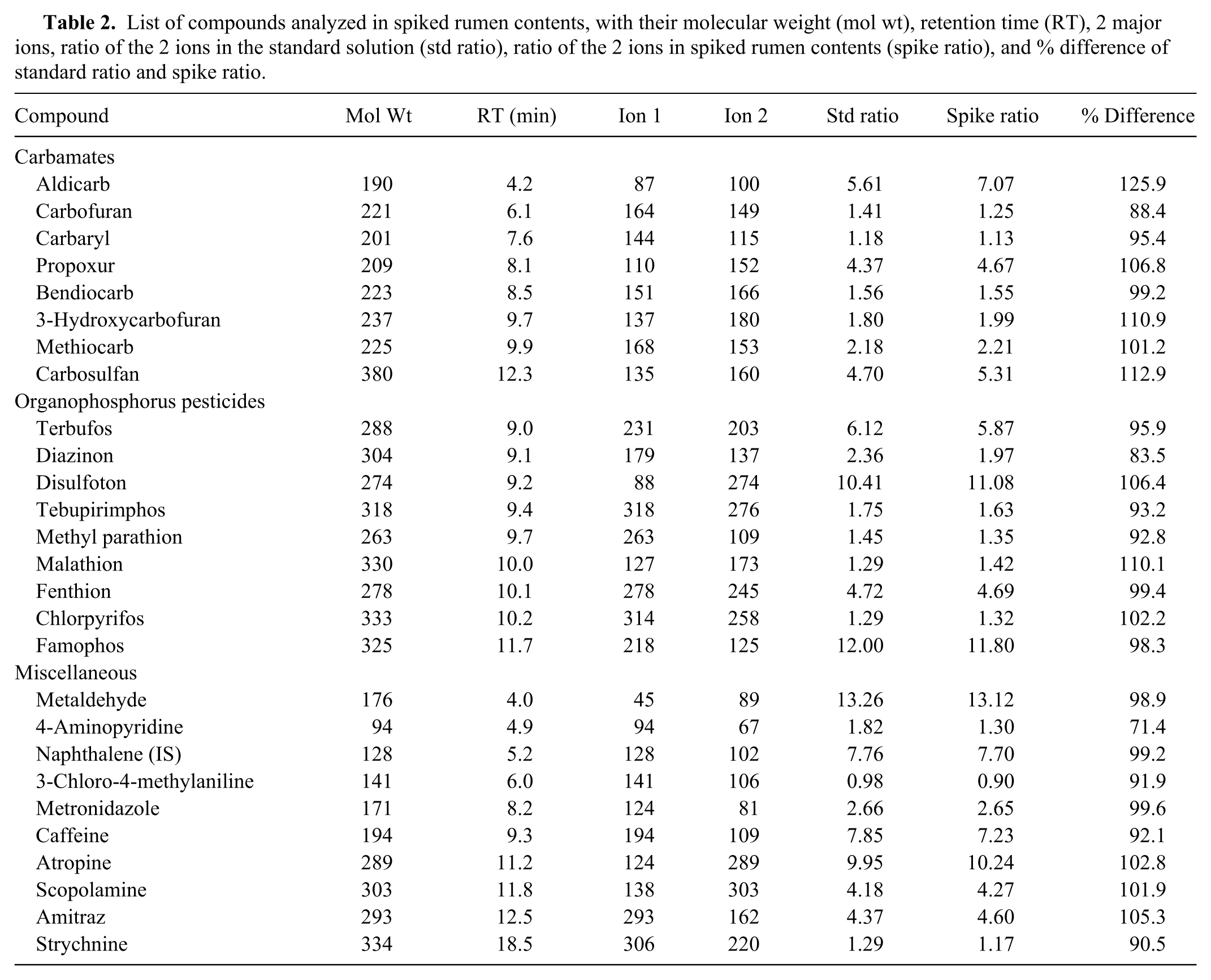
List of compounds analyzed in spiked rumen contents, with their molecular weight (mol wt), retention time (RT), 2 major ions, ratio of the 2 ions in the standard solution (std ratio), ratio of the 2 ions in spiked rumen contents (spike ratio), and % difference of standard ratio and spike ratio.
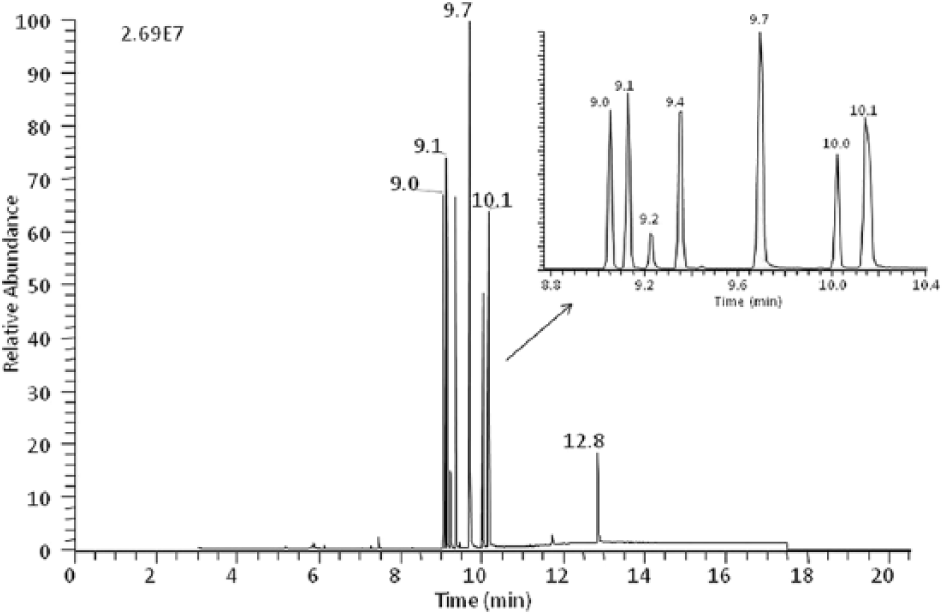
Chromatogram of a control rumen content spiked with organophosphorus pesticides and amitraz mix at 10 µg/g. Inset: region expanded from 8.8 to 10.4 min.
Application of the method
The analyses of several bovine rumen content samples as well as stomach contents of different animal species (birds and dogs) were done using the proposed method. The pH of stomach contents was adjusted to 6–8, if needed. Actual case samples were taken from specimens submitted as suspected pesticide poisonings. Representative chromatograms of the rumen and stomach contents are shown in Figure 3. The analysis of one sample of rumen contents exhibited 2 peaks (Fig. 3A) that were identified by library search as endosulfan. Based on comparison of the sample with pure standards of endosulfan I, II, and endosulfan sulfate, the first peak at 10.9 min was assigned to endosulfan and the second peak at 11.5 min was assigned to endosulfan sulfate. Figure 3B shows the chromatogram of stomach contents from a dog that was found to be positive for both phorate and terbufos. Figure 3C is from ingesta of a pigeon found positive for 4-aminopyridine. The data demonstrates that the developed method has applications beyond rumen content analysis and can be used to detect a variety of toxicants beyond the list of analytes shown in Table 2.
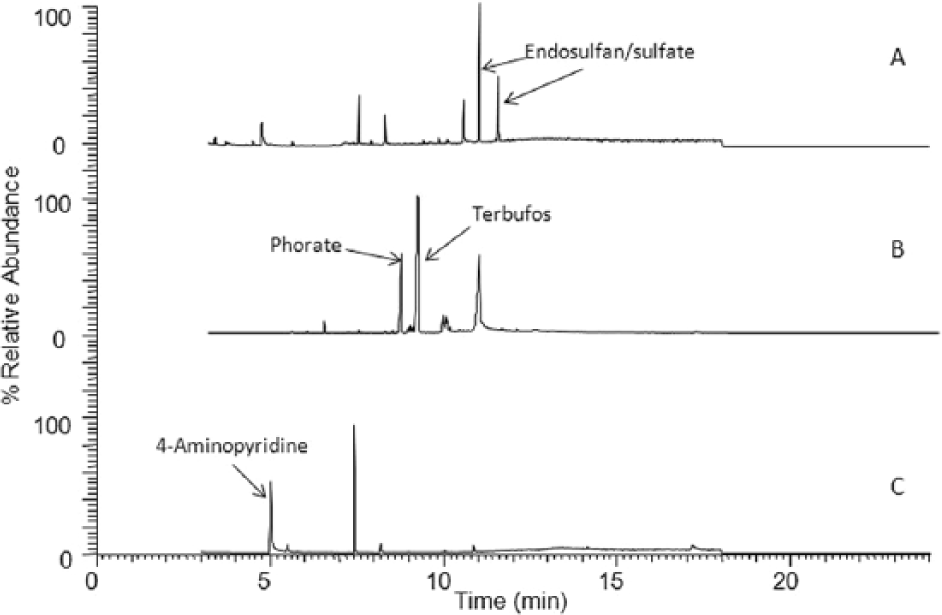
Chromatogram.
Discussion
A simple QuEChERS method for the analysis of various carbamates, organophosphorus pesticides, and miscellaneous compounds in rumen contents was developed in the current study. The objective was to develop a qualitative method that could help make accurate diagnoses due to either accidental or intentional exposure to toxicant(s). Initially, a profile of expected compounds present in normal rumen contents was created by analyzing 20 different samples of rumen contents. These compounds are listed in Table 1 with retention time of each compound, major ion observed, and the identity of the compound based solely on the NIST library. This was done to help facilitate identification of unknown toxicants by ruling out the diagnostically insignificant peaks normally present in rumen contents. The method developed is a general organic chemical screen that can not only detect the compounds specifically selected for the current study but also a wide variety of other compounds that are amenable to analysis by GC and spectral match using a NIST library search capability. The method has also been applied successfully to similar matrices often encountered in veterinary diagnostic toxicology.
In veterinary diagnostic toxicology laboratories, GC-MS organic chemical analyses routinely utilize a NIST library search, accessing more than 147,000 spectra to facilitate the identification of unknown substances. Samples, including rumen and/or GI contents, and/or baits, are routinely submitted in cases when an animal has died unexpectedly and ingestion of a toxicant is suspected. In such cases, a sensitive method is desirable, but accurate identification of the toxicant is paramount. Most rumen or GI contents found to be positive by GC-MS analysis in diagnostic veterinary toxicology laboratories tend to contain concentrations of toxicants that are high enough to be easily detected and identified.
When certain drugs are suspected, such as specific pesticides, mycotoxins, ionophores, or anticoagulant rodenticides, LC-MS/MS screening methods with the ability to detect toxicants at low part per billion levels offer enhanced sensitivity. Additionally, general LC-MS/MS screening methods available in the literature are ideal for ensuring that animal foods and feeds are free of potentially harmful levels of contaminants. However, because of the unavailability of a comprehensive library, the usefulness of LC-MS/MS methods is limited to the analytes that are selected as part of in-house developed methods. Furthermore, when an unknown compound is present in a sample that is not part of a developed LC-MS/MS method, it is difficult to identify a potential toxicant based on its chromatogram and spectral data alone.
Until comprehensive libraries with the capability to screen thousands of compounds are available for LC-MS/MS, GC-MS equipped with a NIST library will continue to be a valuable tool in the identification of unknown toxicants. At the same time, the sensitivity attainable by LC-MS/MS is critically needed not only to ensure food and feed safety but also to positively identify toxicants that would otherwise elude detection by GC-MS. Both GC-MS and LC-MS/MS methods are necessary as no one method is ideal for the identification of all toxicants that may potentially harm animals. Since the first introduction of the QuEChERS method for the analysis of pesticides in produce, this simple and quick method has been adopted for a variety of matrices including animal feed. The current study demonstrates the application of the QuEChERS clean-up method for an additional matrix commonly submitted to veterinary diagnostic laboratories, rumen contents. While rumen contents are a fairly uniform sample, GI contents from other animal species may require modified sampling and preparation strategies prior to analysis. Characteristics of GI content samples that may be important for analytical purposes can differ not only between various animal species but also among groups and individuals within a certain species depending on the provided diet and any baits or other abnormal substances that may have been consumed. The decision to homogenize the sample or to preferentially select out suspicious material needs to be made first, followed by adjusting the pH to neutral prior to proceeding with extraction process. The usefulness of this clean-up technique for rumen or GI contents needs to be further explored with more sensitive LC-MS detection methods and additional toxicants at levels of concern in diagnostic veterinary toxicology.
Footnotes
Acknowledgements
The authors thank Ms. Nina Richey for technical assistance in preparing the samples and Penn State University’s Animal Diagnostic Laboratory for providing the control rumen contents.
a.
Fisher Scientific, Fair Lawn, NJ.
b.
AccuStandard Inc., New Haven, CT.
c.
Supelclean ENVI, Sigma-Aldrich, St. Louis, MO.
d.
Thermo Scientific Polaris Q with Trace GC Ultra, Tri-plus autosampler; Thermo Scientific, West Palm Beach, FL.
e.
HP-5MS 30 m × 0.250 mm column, Agilent, Santa Clara, CA.
f.
NIST MS Search 2.0a library, National Institute of Standards and Technology, Gaithersburg, MD.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The study was supported by the U.S. Department of Agriculture (CSREES PENV2008-3048) and Pennsylvania Animal Diagnostic Laboratory System.
