Abstract
Brucella species infect a wide range of hosts with a broad spectrum of clinical manifestations. In mammals, one of the most significant consequences of Brucella infection is reproductive failure. There is evidence of Brucella exposure in many species of marine mammals, but the outcome of infection is often challenging to determine. The eastern Pacific stock of northern fur seals (NFSs, Callorhinus ursinus) has declined significantly, spawning research into potential causes for this trend, including investigation into reproductive health. The objective of the current study was to determine if NFSs on St. Paul Island, Alaska have evidence of Brucella exposure or infection. Archived DNA extracted from placentas (n = 119) and serum (n = 40) samples were available for testing by insertion sequence (IS) 711 polymerase chain reaction (PCR) and the Brucella microagglutination test (BMAT), respectively. As well, placental tissue was available for histologic examination. Six (5%) placentas were positive by PCR, and a single animal had severe placentitis. Multilocus variable number tandem repeat analysis profiles were highly clustered and closely related to other Brucella pinnipedialis isolates. A single animal was positive on BMAT, and 12 animals had titers within the borderline range; 1 borderline animal was positive by PCR on serum. The findings suggest that NFSs on the Pribilof Islands are exposed to Brucella and that the organism has the ability to cause severe placental disease. Given the population trend of the NFS, and the zoonotic nature of this pathogen, further investigation into the epidemiology of this disease is recommended.
Introduction
Brucella spp. are facultative aerobic intracellular, Gram-negative bacteria that are most commonly associated with reproductive disease in a wide range of animals. 33 Since the 1990s, the organism has been increasingly documented in marine mammals, the range and distribution of which are well reviewed elsewhere. 20 Species within the Brucella genus are identified by their preferred host and, in 2007, 2 new species, Brucella pinnipedialis and Brucella ceti, were recognized. 12 The significance of Brucella spp. in marine mammals is largely unknown, as the organism has been identified in animals both with and without associated pathology. Clinical disease is most commonly reported in dolphins where B. ceti infection can result in severe meningoencephalitis, a condition often referred to as neurobrucellosis,17–19 and is also reported to cause abortion with placentitis.11,27 In contrast, pathology associated with B. pinnipedialis infection has not been reported despite evidence of exposure across a wide range of pinnipeds. 20 Interestingly, the organism has also been identified in pulmonary nematodes in a pacific harbor seal (Phoca vitulina richardii), 14 a harp seal (Phoca groenlandica), 25 and a harbor porpoise (Phocoena phocoena). 7
The northern fur seal (NFS, Callorhinus ursinus) is found throughout the North Pacific Ocean, with large breeding colonies on a small number of islands. 15 The largest segment of the population resides on the Pribilof Islands of Alaska where it has experienced significant population declines since the late 1950s. 36 In 1988, the eastern Pacific stock of NFS was listed as “depleted” under the Marine Mammal Protection Act (National Marine Fisheries Service: 2007, Conservation plan for the Eastern Pacific stock of northern fur seal (Callorhinus ursinus). U.S. Department of Commerce. Available at: http://www.alaskafisheries.noaa.gov/protectedresources/seals/fur/cplan/final1207.pdf ). Investigation into potential causes for this decline has led to the identification of multiple previously unreported infectious agents in NFS placentas, including Coxiella burnetii 9 and a novel polyomavirus. 8 Although these pathogens were identified in detached placentas with no information regarding the survival of the associated pup, there is evidence that placental infection with Coxiella can induce a functional change in the NFS placenta, 28 which could have an impact on the development and survival of offspring. Given that the greatest historical loss of individual NFS, other than commercial harvest, is during the first year of life 24 and that pup birth weight is correlated with survival, 4 there is concern that subtle developmental impairments leading to poor recruitment may be associated with the declining population trend. Little is known about other infectious agents that may influence reproductive success in the NFS, but considering the recognized role of Brucella infection in other marine mammals, the objective of the current study was to determine if NFSs have evidence of Brucella exposure or infection.
Materials and methods
Archived serum and placentas collected on St. Paul Island, Alaska, as part of routine live animal, rookery, and postmortem sampling were available for use in this study. Serum from 40 animals had been collected by standard venipuncture technique and stored at –80°C until testing. Twenty-five samples were from subadult males sampled immediately postmortem as part of the St. Paul subsistence harvest in 1994 (n = 15) or 2011 (n = 10). Fifteen serum samples from female NFSs represented 11 adults and 4 pups handled in 2009 and 2011. Serum samples were tested by Brucella microagglutination test (BMAT) with minor modifications. 2 A titer of ≥160 was considered positive, 20–80 for borderline, and <20 was considered negative. A high, low, and negative reference control was included with each run. a
In 2011, 119 placentas were collected from a single NFS rookery on St. Paul Island, Alaska as part of an ongoing health and disease investigation program. Each placenta was subsampled, and both fresh and formalin-fixed tissues were archived. DNA was extracted from each placenta as previously described 10 and available for use in the present study. Additionally, DNA was extracted from serum samples using a commercial kit. b Brucella spp. DNA was detected by real-time PCR (qPCR) using a fluorescently labeled hydrolysis probe targeting the insertion sequence (IS) 711 gene as previously described. 21 The PCR conditions were modified for detection in clinical specimens. The qPCR amplification was performed using 5 µl of DNA extract, 200 nM of each primer and probe, and a commercially available PCR master mix c in a 25-µl reaction volume. An exogenous internal positive control c was added to each reaction to test for inhibition. Amplification and fluorescence detection was performed on a commercially available PCR platform c with an initial denaturation for 10 min at 95°C, 45 cycles of denaturation for 15 sec at 95°C, and annealing for 1 min at 60°C. A sample was considered positive if the fluorescent growth curves crossed the threshold line within 40 cycles.
Multilocus variable number tandem repeat (VNTR) analysis was performed on IS711 PCR–positive placentas. The PCR amplification of the 15 VNTR loci was performed as described previously23,35 with modifications for amplification in low template samples. Multiplex PCR assays (AI, BI, AII, and BII) were performed to amplify 15 VNTR loci in a final volume of 10 µl containing 1× PCR buffer, 2 mM MgCl2, 2 mM deoxynucleoside triphosphate mix, 0.04 U DNA polymerase, d and 5 µl of DNA template. All reactions were performed on a commercial thermocycler using the primer concentrations reported previously. Thermal cycling amplification cycles were increased from 35 to 45. Multiplex PCR assays (AI + BI and AII + BII) were combined 1:1. Amplicons underwent fragment analysis on an automated fluorescent capillary DNA sequencer. c Allele designations were assigned by internal binning capabilities in the commercially available software, c and a distance tree was generated by clustering analysis using the unweighted pair group method with arithmetic averages (UPGMA).
All placentas positive on PCR were examined microscopically. Formalin-fixed tissues were paraffin embedded, sectioned at 5 µm, and stained with hematoxylin and eosin for histologic review. Sections with histologic lesions were further reviewed using immunohistochemical staining (IHC) for C. burnetii 9 and Brucella 14 as previously described.
Results
Six (5%) of the 2011 placentas were IS711 positive by PCR; all of these were also positive for C. burnetii as previously reported. 10 Six sections from each of the PCR-positive placentas were available for histologic examination. In a single section of 1 placenta there was a regionally extensive area of necrosis and suppurative placentitis resulting in substantial tissue loss (Fig. 1A). At the base of the affected region, the wall of a chorionic artery was expanded by lymphocytes, plasma cells, rare neutrophils, and edema; perivascular cuffing was also prominent (Fig. 1B). Within rare, viable trophoblasts at the periphery of the lesion, aggregates of intracytoplasmic bacteria expanded the cytoplasm (Fig. 1C). There was strong Brucella immunostaining within the cytoplasm of trophoblasts at the periphery of the lesion as well as within the adjacent necrotic cellular debris (Fig. 1D); no C. burnetii immunostaining was identified. No organism was identified in the other placentas that were positive by IS711 PCR; however, 4 of the 5 had small foci of necrosis or suppurative inflammation in at least 1 of the examined sections, and a single placenta had foci of mineralization that appeared slightly increased relative to previously reviewed placentas from this species.
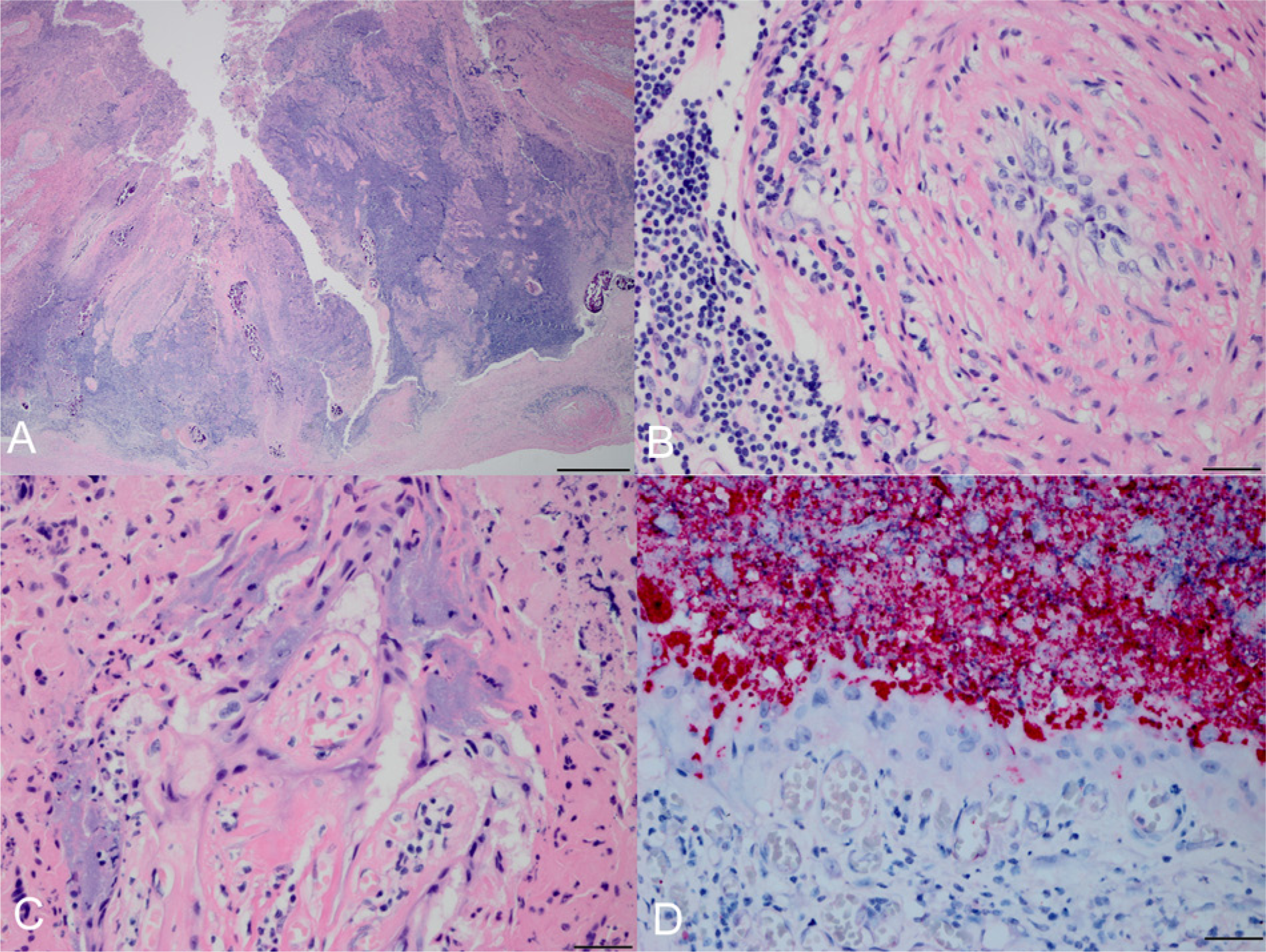
Northern fur seal (Callorhinus ursinus) placenta with histologic lesions associated with Brucella infection.
Complete VNTR profiles (15/15 loci) were obtained from 2 of the IS711 PCR–positive placentas (P57 and P53) while nearly complete profiles were obtained from samples P55 (12/15) and P58 (13/15). The UPGMA clustering analysis of the VNTR profiles indicated a great deal of similarity between these NFS samples (Fig. 2). Samples P57 and P53 have identical VNTR profiles. Despite the lack of amplification of certain loci in samples P55 and P58, these 2 samples clustered most closely with the other 2 NFS placenta samples.
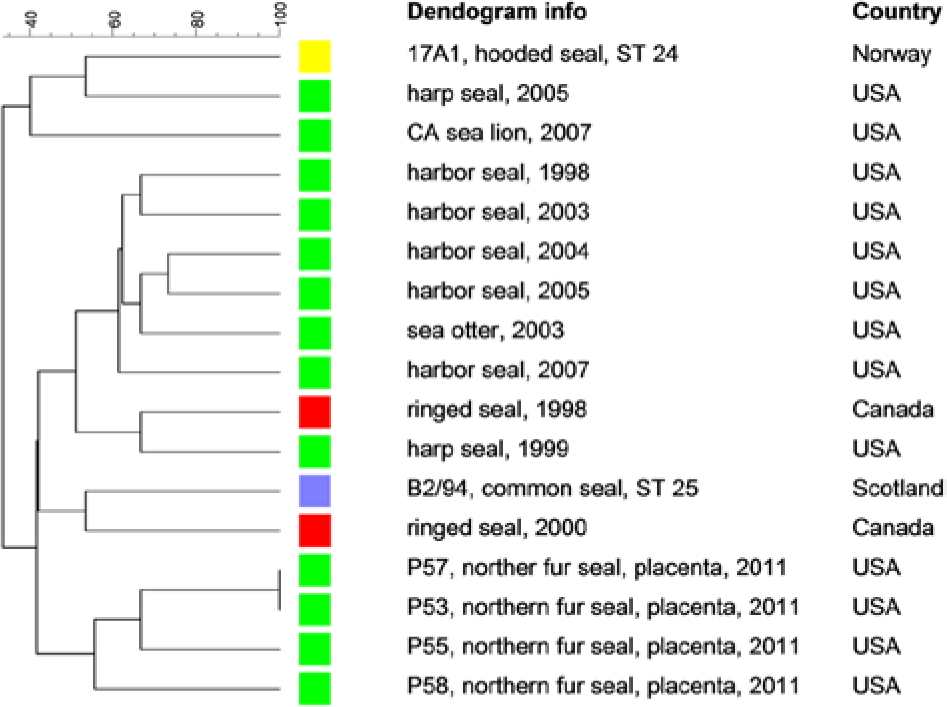
Clustering analysis of multilocus variable number tandem repeat analysis profiles using the unweighted pair group method with arithmetic averages analysis obtained from northern fur seal (Callorhinus ursinus) placenta samples and other Brucella pinnipedialis isolates from the United States and Canada.
The serum of a single adult female, sampled in 2011 with no known reproductive history, was positive on BMAT with a titer of 1:160. Twelve additional animals had titers within the borderline range; these animals were predominantly subadult males (10/12) sampled in both 1994 (5/12) and 2011 (7/12). A single animal, borderline on BMAT, was IS711 positive by PCR on serum. This animal was a subadult male harvested for subsistence purposes in 2011.
Discussion
The histologic lesion identified in the single NFS placenta was striking and unexpected; however, the significance of infection and placental pathology for the pup or the dam is unknown. The consequence of placental infection is dependent on a number of factors with outcomes ranging from normal births to unthrifty offspring or abortions and stillbirths. Bacterial infection during pregnancy can also result in maternal reproductive consequences; in bison, animals aborting due to Brucella infection have decreased subsequent reproductive success relative to seronegative bison.13,31 As this placenta was collected from the rookery, detached from the associated pup, it is assumed that the pup was born alive; however, given the nature of the heavily populated rookery, it is impossible to know the survival of that individual. Although this placenta was also positive for C. burnetii by PCR, no C. burnetii staining was identified by IHC and the lesion is therefore assumed to be due to Brucella infection. It is notable that this placenta specimen (P53) had very low crossing threshold values on the IS711 qPCR indicating high bacterial load in the placenta tissue. Brucella has been identified (culture and PCR) in the placentas of California sea lions (Zalophus californianus) aborting due to domoic acid toxicity, but the placentas were devoid of histologic lesions. 16
Clustering analysis of the NFS placenta samples clearly demonstrates not only the close relatedness of the Brucella strains in the NFS placenta samples but also their overall close association with other B. pinnipedialis strains recovered from other seals and sea lions in the United States and Canada. A number of serosurveys have been conducted in marine mammals of the North Pacific Ocean, and the results are variable. Antibody titers are uncommon (0.005%) in the Steller sea lion (Eumetopias jubatus), 3 but relatively common (46%) in the Alaskan harbor seal (Phoca vitulina). 37 Polar bears (Ursus maritimus) on the Alaskan North Slope have approximately 10% seroprevalence; however, it is unclear if the exposure is via a terrestrial or marine source. 30 Brucella antibodies were rare 29 or not detected 5 in the Pacific walrus (Odobenus rosmarus divergens). Unfortunately, the true level of exposure is often difficult to ascertain as there are large gaps in knowledge regarding the performance of different serologic assays in many marine mammals. For example, an extremely wide range of apparent prevalence (16–74%) was demonstrated in the Alaskan harbor seal depending on the assay used for antibody detection. 22 As such, serologic results should be interpreted with caution.
The majority of the animals that were borderline on BMAT or seropositive in the present study represent subadult males killed in the subsistence harvest, in good body condition with no reported signs of illness. One of these animals was also PCR positive indicating the presence and circulation of DNA that may be consistent with active infection; the duration of bacteremia in Brucella-infected pinnipeds is unknown. Marine Brucella infection has resulted in human disease,1,26,34 and experimental studies have determined that marine isolates can cause disease in domestic cattle. 32 The NFS is an important cultural and nutritional resource for the Aleut people of St. Paul Island. Since the end of the commercial NFS harvest, the Aleut community has continued to harvest NFS for food with, on average, 388 animals taken per year on St. Paul Island over the last decade (M. Williams, personal communication, 2013). Brucellosis has been described as one of the most important arctic infectious diseases in Alaska (Brubaker M, Berner J, Butler J, Bradley M: 2010, Brucellosis: understanding an important arctic infectious disease. Alaska Native Tribal Health Consortium, Center for Climate and Health bulletin. Available at: http://www.anthc.org/chs/ces/climate/upload/CCH-Bulletin-No-5-Brucellosis-Understanding-an-Important-Arctic-Infectious-Disease-4.pdf), although the public health risk is thought to be due to hunting of terrestrial, and not marine, mammals. As an intracellular organism, known to infect circulating monocytes and tissue macrophages, 6 it is possible that the organism may be present in tissues collected for consumption; however, additional information is needed regarding tissue distribution and persistence during food preparation before these risks can be meaningfully assessed. While the presence of Brucella DNA within the serum of a single male NFS may be consistent with active infection, it could also represent inactive DNA persisting within phagocytic cells that would not pose any threat of transmission.
While the current study represents novel information regarding NFS exposure, infection, and pathology associated with Brucella spp., it also raises new questions regarding the significance of these infections for NFS and sympatric species. To address public health concerns, investigation of the viability and distribution of the organism within different tissues is warranted. Additional testing of biological samples may help elucidate at-risk cohorts and geographic locations that may be more contaminated by bacteria shed at the time of parturition. Finally, concern about the number of reproductive pathogens, and coinfections, identified within this declining population necessitates prospective studies integrating vital rates with microbiology and other health parameters.
Footnotes
Acknowledgements
All animal tissue samples were collected under authorization of U.S. Marine Mammal Protection Act Permit nos. 837, 782-1708, and14327, issued to the National Marine Mammal Laboratory, Alaska Fisheries Science Center, Seattle, Washington. The authors thank Jack Rhyan, E.J. Ehrhart, and Brad Charles for assistance with immunohistochemical staining and The Tribal Government of St. Paul Island for their collaboration in sample collection.
a.
U.S. Department of Agriculture, Animal and Plant Health Inspection Service, Veterinary Services, National Veterinary Services Laboratories, Ames, IA.
b.
DNA blood mini-kit, Qiagen Inc., Valencia, CA.
c.
Life Technologies, Grand Island, NY.
d.
Invitrogen Corp., Carlsbad, CA.
e.
Applied Maths NV, Sint-Martens-Latem, Belgium.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) declared that they received no financial support for their research and/or authorship of this article.
