Abstract
The A32L gene from a Goatpox virus (GTPV) strain isolated from a goat in Yunlin County (Taiwan) displays several substitutions compared with the sequence of the Kenyan GTPV vaccine strain SGP0240 and the Pellor GTPV strain. Samples from the skin lesions on 6 goats with GTPV infection or from goats with Orf virus (ORFV) infection were tested in a multiplex polymerase chain reaction (PCR) system that used primers GPF, GPR1, and GPR2 as well as previously published primers specific for ORFV. These primers were able to amplify either GTPV or ORFV without cross-reactivity. A high-resolution melt analysis (HRMA) was carried out on amplified DNA from the skin lesions of 6 goats with GTPV infection and with the GTPV SGP0240 strain. The results indicated that the melting temperature profiles amplified from samples with Yunlin GTPV infection can be differentiated from the GTPV SGP0240 strain. The findings showed that a successful differential assay for these GTPVs had been developed. Accordingly, both methods can be used to detect and differentiate GTPV isolated from animals that may have either been vaccinated or been infected with a wild strain. The multiplex PCR and HRMA could be used on skin samples of suspected cases to serve as the front-line and confirmative assays, respectively, which will be beneficial to the eradication of GTPV.
Introduction
Goatpox virus (GTPV), Sheeppox virus (SPPV), and Lumpy skin disease virus (LSDV) all belong to the Capripoxvirus (CaPV) genus, a genus associated with goat and sheep diseases. Capripoxviruses have a linear double-stranded DNA genome, are large (approximately 154 kilobase pairs), and are classified in the family Poxviridae. 11 Capripoxvirus virions are brick-shaped enveloped viruses with rounded ends and display a complex symmetry. 12 Generally, SPPV and GTPV exhibit particular host specificities, but some strains of these viruses are able to infect both goats and sheep.11,12
Members of the CaPV genus cause significant economic losses because of high morbidity, a decrease in milk yield, reduced weight gain, increase in abortion, damage to hides, damage to wool, and a higher risk of pneumonia. 12 Up to 75% of animals can be infected in a susceptible flock and the mortality may be as high as 50%. 11 In naïve and young animals, the morbidity and mortality can approach 100%. 2
The clinical manifestations of SPPV, GTPV, and LSDV are similar. 12 Animals infected with CaPVs initially present with concurrent fever and skin macules. Consequently, animals develop excessive salivation, nasal discharge, lacrimation, coughing, conjunctivitis, and an arching of the back. The macules on the skin further enlarge and progress to become papules, vesicules, pustules, and then scabs.1,2 These pox lesions are also evident on the tongue, on the gums, in the lungs, and within the gastrointestinal tract. 12
Capripoxviruses are transboundary diseases and are World Organization for Animal Health (OIE)-listed diseases. 13 For the past half century, the geographic range of goat pox and sheep pox was limited to Africa and Asia. However, during the past 50 years, goat pox and sheep pox have been found in the Middle East, Turkey, the Indian subcontinent, Greece, Mongolia, and Israel.1,11
Capripoxviruses can be readily diagnosed in acute cases. However, the diagnosis of CaPV-associated disease can be complicated by other viruses that cause exanthema or by the fact that infection is caused by a low virulent strain. 3 For example, CaPV-associated disease can be confused with the parapoxvirus that causes contagious pustular dermatitis (contagious ecthema, scabby mouth, or orf disease) and with the herpesvirus that causes pseudolumpy skin disease.7,9 Therefore, it is important that a laboratory test is established for the definitive diagnosis of GTPV. 3
Taiwan reported a goat pox outbreak that lasted from 2009 to 2010. The strategy used to control the spread of goat pox initially after the outbreak began was the immediate culling of infected animals. 8 However, this outbreak spread rapidly throughout the country and the policy was adjusted to one where the number of susceptible animals was reduced via a comprehensive vaccination program, as described elsewhere. 14 A live attenuated vaccine, which contains the Kenyan vaccine strain of GTPV, SGP0240, was chosen and used to immunize all goats in Taiwan based on the recommendation of the World Organization for Animal Health (OIE).8,14
A simple and fast method for goat pox diagnosis would be useful to the field veterinarian. A quick response is vitally important when there is a suspected outbreak of poxvirus in goats. An easy differential diagnostic method for poxviridae infection, including GTPV, would be a convenient tool to control this disease in goats. The aims of the current study were to develop a multiplex polymerase chain reaction (PCR) assay and high-resolution melt analysis (HRMA) in order to detect GTPV infection and then to determine whether a suspected goat has been vaccinated or is infected by a wild-type goat pox strain. Live vaccines may in some circumstances cause lesions and therefore the development of a differential diagnosis system will allow the safety of the GTPV vaccine strain to be monitored.
Materials and methods
Samples
Material was obtained during an outbreak of GTPV infection in Taiwan. Most samples were obtained from goats with suspected GTPV infection that were farmed in Yunlin County, Taiwan, during 2009. In addition to a pathological assessment of any lesions in the breasts and tails, skin lesions from 6 goats were biopsied and subsequently subjected to histological examination, PCR analysis, and electron microscopy. The biopsied samples for histological analysis were sequentially fixed in 10% buffered formalin, trimmed, dehydrated, embedded in paraffin, and stained by hematoxylin and eosin before examination by light microscopy. Skin samples were also negatively stained with 2% phosphotungstic acid and then examined by electron microscopy.
DNA extraction
DNA samples were extracted from the skin lesions of 6 goats with suspected GTPV infection. In addition, DNA samples from 2 skin lesions of each goat infected with the Orf virus (ORFV) strain Hoping, Nantou, or Taiping (3 goats per virus strain) were available from a previous study. 4 Furthermore, DNA samples were also extracted from the vaccine strain SGP0240 that had a titer of 102.5 50% tissue culture infectious dose per milliliter (TCID50/ml) and from the field strains of GTPV that had been propagated in lamb kidney cells (minimum titer of 105 TCID50/ml). In all cases, the DNA was extracted using a commercial kit a by following the manufacturer’s instructions.
Cloning and sequencing of PCR products
The full-length of open reading frame of the A32L gene was amplified by PCR from the 6 GTPV Yunlin isolates and from the SGP0240 GTPV. The primers used were GPA32L-F and GPA32L-R (Table 1) and were based on the genome sequence of the Pellor GTPVstrain, which was obtained from GenBank (accession no. NC_004003.1). The Pellor GTPV strain in this study was used as a reference strain against which all other strains were compared. The PCR was performed in a 50-μl reaction mixture containing 0.2 μM of each oligonucleotide primer, 10 μl of 5× Taq master mix, b and 100 ng of each DNA sample. The PCR had an initial denaturation at 94°C for 5 min, 35 cycles at 94°C for 35 sec, 48.7°C for 30 sec, and 72°C for 1 min, which was followed by a final extension at 72°C for 7 min. All PCR products were analyzed by electrophoresis using 2% agarose gels and then the bands were extracted and purified from the gel using a commercial kit c according to the manufacturer’s instructions. The purified fragments were cloned into a commercial vector d and transformed into Escherichia coli strain DH5α. e The plasmids were examined by restriction endonuclease digestion and agarose gel electrophoresis to confirm the presence of inserts. Three positive clones were sent for automated sequencing to obtain the nucleotide sequence of the insert. Both the nucleotide and deduced amino acid sequences of the GTPV Yunlin, SGP0240, and Pellor strains and the Hoping, Nantou, and Taping strains of ORFV were aligned based on the Clustal W algorithm. The phylogenetic relationship was analyzed by the neighbor-joining program. f
Primers used in the current study.
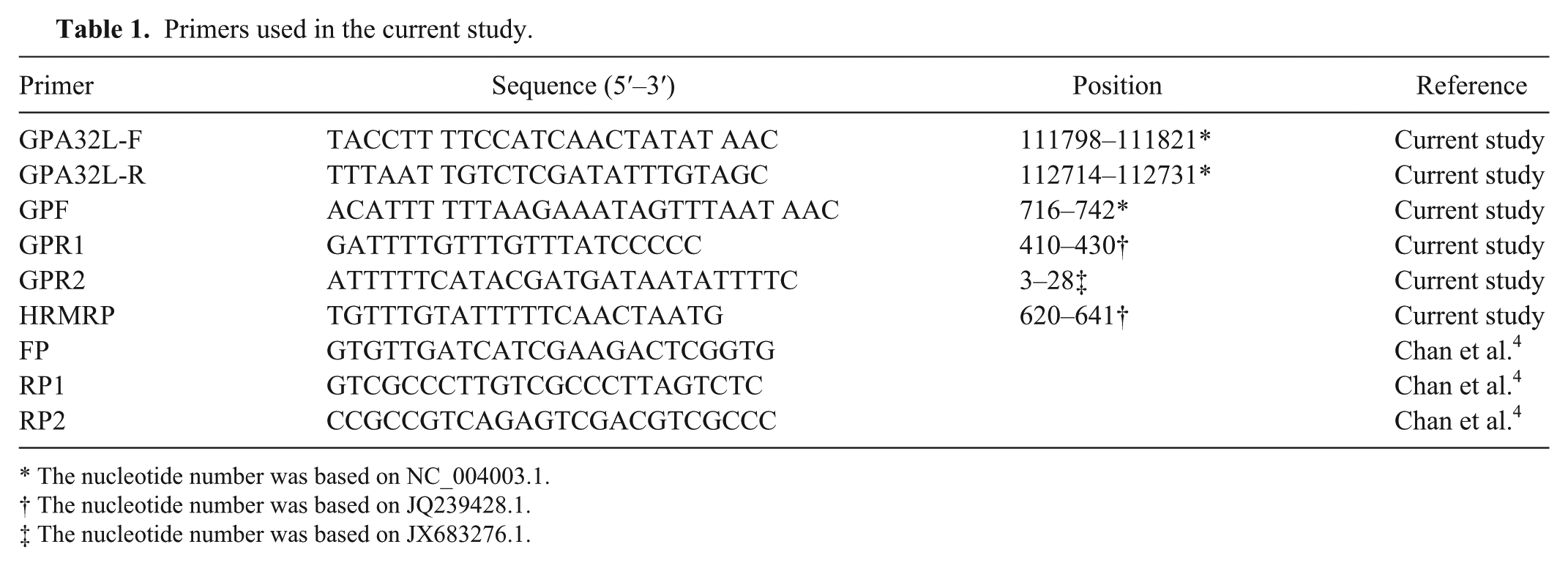
The nucleotide number was based on NC_004003.1.
The nucleotide number was based on JQ239428.1.
The nucleotide number was based on JX683276.1.
Differential diagnosis in goats of poxvirus disease by multiplex PCR
The primers used for differential diagnosis of GTPVs were GPF, GPR1, and GPR2, together with the 3 published primers, FP, RP1, and RP2, used previously for diagnosis of ORFV in Taiwan (Table 1). 4 The PCR reaction was performed in a 25-μl reaction mixture. The mixture contained 80 nM of GPF primer, 40 nM of GPR1 primer, and 40 nM of GPR2 primer for GTPV as well as 120 nM of FP primer, 60 nM of RP1 primer, and 60 nM of RP2 primer for ORFV, 2.5 μl of 10× reaction buffer, 5 μl of MgCl2 (50 mM), 7.5 μl of deoxynucleotides (2.5 mM), 1 unit of Taq DNA polymerase, g and 100 ng of extracted DNA. The PCR conditions consisted of an initial denaturation at 94°C for 5 min, 30 cycles of 94°C for 35 sec, 64°C for 30 sec, and 72°C for 1 min, which was followed by a final extension at 72°C for 7 min. All PCR products were analyzed by electrophoresis on 2% agarose gels and were also subjected to sequencing analysis. For the differential diagnosis, DNA from 2 skin lesions of each goat infected with the Hoping, Taping, or Nantou strain of ORFV (3 goats per virus strain), GTPV-infected goats, and the vaccine strain SGP0240 were tested. To determine the detection limit of the multiplex PCR, DNA extracted from 102, 10, 1, 0.1, and 0.01 TCID50/ml of the vaccine strain SGP0240 and Taiwan isolated GTPV were used as templates for tests.
HRMA for the differential diagnosis of GTPV
High-resolution melt analysis was performed on a commercial instrument h to allow the differential diagnosis of the GTPV strains. Two primers, GPF and HRMRP, were used to amplify a short fragment (Table 1). Initially, dan of EMBOSS (http://emboss.sourceforge.net/apps/release/6.1/emboss/apps/dan.html) was applied via suite, the Mobyle portal, to calculate the melting temperatures of the targeted part of the A32L gene of the GTPV Yunlin and SGP0204 strains that had been amplified by the GPF and HRMRP primers. Taking into consideration the thermodynamic behavior of nearest neighbor bases, the melting curve profiles are presented as an average within a sliding window. The reference value was set as a 20 nucleotide window. The PCR mixture consisted of 80 nM of GPF, 80 nM of HRMRP, 5 μl of 5× Taq master mix, b and 100 ng of extracted DNA in a total volume of 25 μl. The samples examined came from the skin lesions of 6 goats infected with the GTPV Yunlin strain or given the vaccine strain SGP0240. Samples negative for GTPV were included. The PCR reaction conditions consisted of 94°C for 5 min and followed with 35 cycles of 94°C for 35 sec, 48°C for 30 sec, and 72°C for 1 min, followed by a final extension at 72°C for 7 min. The HRMA was performed in a total reaction of 10 μl, which was comprised of 9 μl of amplified PCR product from the skin samples and 1 μl of dye. i The DNA fragments were melted from 60°C to 95°C at a rate of 0.3°C/sec. The melting profiles were analyzed using the commercial software. j
Results
Gross lesions, light microscopy analysis, and electron microscopy analysis
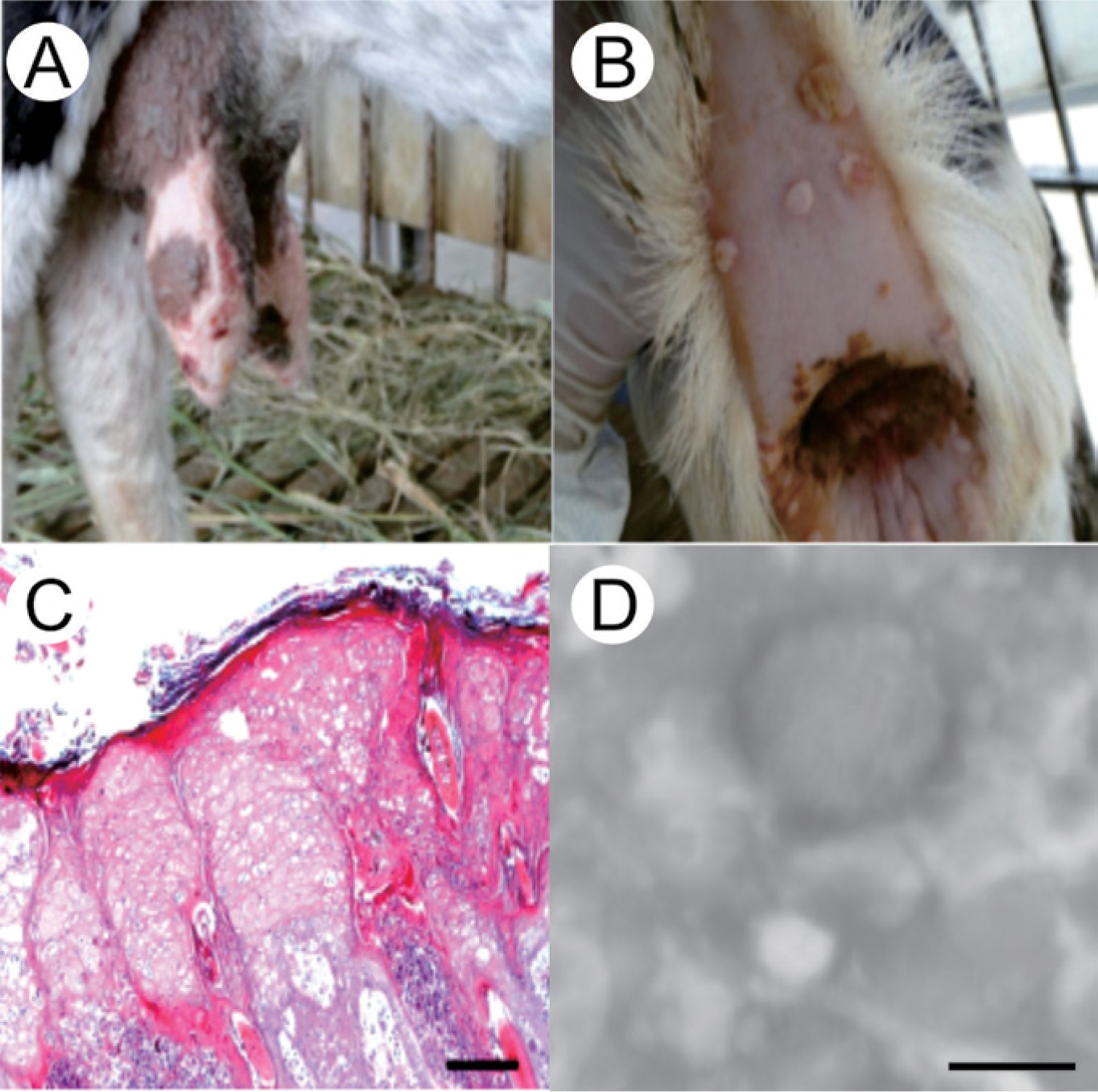
Among the 10 goats examined, 6 were characterized as having multifocal, verrucose, and ulcerated proliferative nodules over the body. The most obvious lesions were present on the skin of the breast and tail (Fig. 1A, 1B). Light microscopy analysis showed the presence of marked hyperplasia and hyperkeratosis in the epidermis of the lesions. Degenerative changes, such as pustule formation and vesiculation, were present in the superficial epidermis. Ballooning degeneration, vacuolation, and intracytoplasmic eosinophilic inclusion bodies were found in keratinocytes of the stratum spinosum (Fig. 1C). Using electron microscopy, the presence of brick-shaped virions with tiny globular structures on the surface could be seen against an electron dense background core. The size of virion was approximately 250 nm × 300 nm in diameter (Fig. 1D).
Cloning and sequences analysis
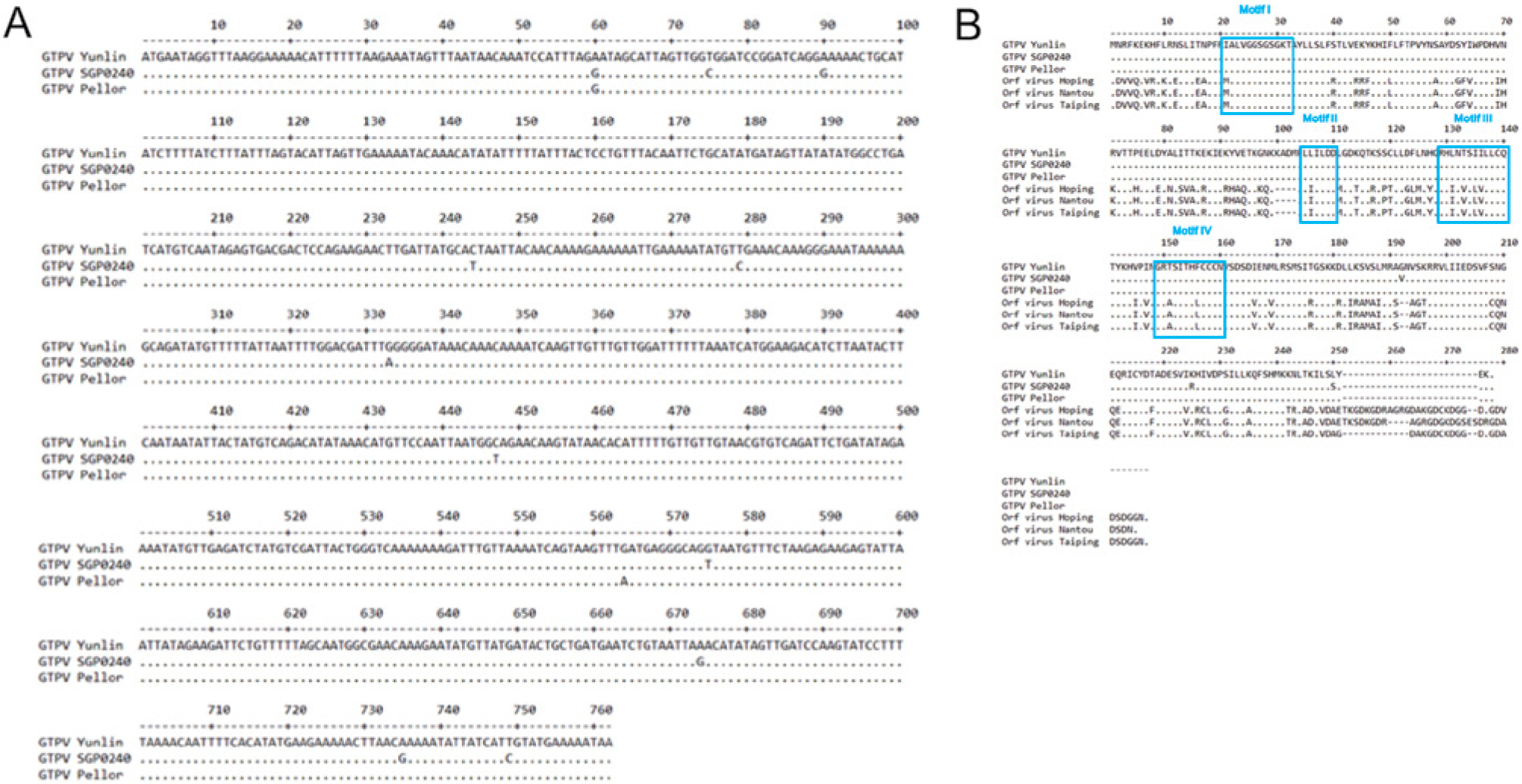
DNA fragments containing the full-length open reading frame of the A32L gene from the 6 Yunlin GTPV isolates and the SGP0240 GTPV strain were amplified by PCR. A prominent band with a size of 940 base pairs (bp) was obtained (data not shown). The various PCR products were purified and cloned into a commercial vector d for DNA sequencing. All sequences from individual clones containing the A32L gene of the 6 GTPV strains isolated from Yunlin were identical, which suggests that, to date, available field isolates from Taiwan are the same strain, named the Yunlin GTPV strain. The GenBank accession numbers of nucleotide sequences of the Yunlin and SGP0240 GTPV strains are JQ239428.1 and JX683276.1, respectively. There are 11 substitutions in the nucleic acid sequence of the A32L gene when the Yunlin and SGP0240 GTPV strains were compared (Fig. 2A). When the amino acid sequence of the A32L protein of GTPV was examined, 4 motifs within the A32L protein, namely IALVGGSGSGKT (motif I), LLILDD (motif II), RHLNTSIILLCQ (motif III), and RTSITHFCCCN (motif IV), were found to be conserved across the Yulin, Pellor, and SGP0240 GTPV strains. Similar to the findings for ORFV, these motifs are representative of the characteristics of a nucleoside triphosphatase (NTPase; Fig. 2B).5,10

Differential diagnosis using multiplex PCR
The optimal annealing temperature for the multiplex PCR was determined by trial and error to be 64°C. The sizes of the PCR products from the Yunlin and GTPV SGP0240 strains were 333 and 740 bp, respectively. No cross-reactivity for GTPV was found with the primers targeting the ORFV. Polymerase chain reaction products with sizes of 180 and 254 bp were amplified from the ORFV Hoping strain. For the ORFV Taiping and Nantou strains, the sizes of PCR products were 271 and 180 bp, respectively. As expected, no product was obtained when ORFV was amplified using the primers designed for GTPV. In addition, no amplified products were observed in the negative control group (Fig. 3). This multiplex PCR system is able to successfully differentiate a range of different GTPV and ORFV strains.
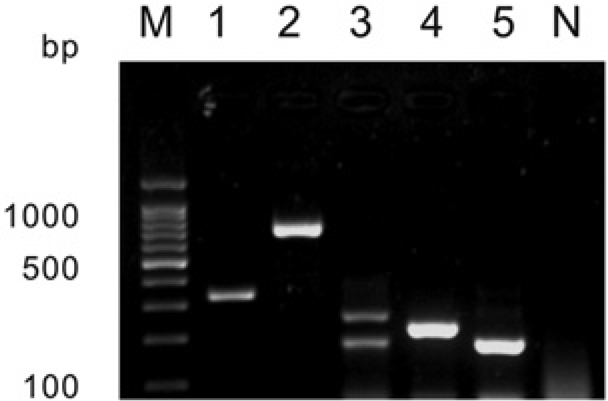
Analysis of the multiplex polymerase chain reaction products amplified from clinical samples for diagnosis of Goatpox virus. Lanes 1 and 2 are the Yunlin and SGP0240 strains of Goatpox virus, respectively, while lanes 3–5 are the Hoping, Taiping, and Nantou strains of Orf virus, respectively. Lane N is the negative control.
Sensitivity of the multiplex PCR
A 333-bp product was obtained by PCR when the amount of the GTPV vaccine strain was as low as 10 TCID50/ml (Supplemental Fig. 1A). Similarly, a 740-bp product was obtained as the amount of the GTPV Taiwan isolate was reduced from 102 to 0.1 TCID50/ml (Supplemental Fig. 1B). This result showed that the multiplex PCR is able to detect GTPV genomic DNA when the amount of virus is as low as 10 TCID50/ml, no matter whether the Yunlin or the SGP0240 strain is present.
Nucleotide and amino acid sequence similarity of the A32L gene across all viruses
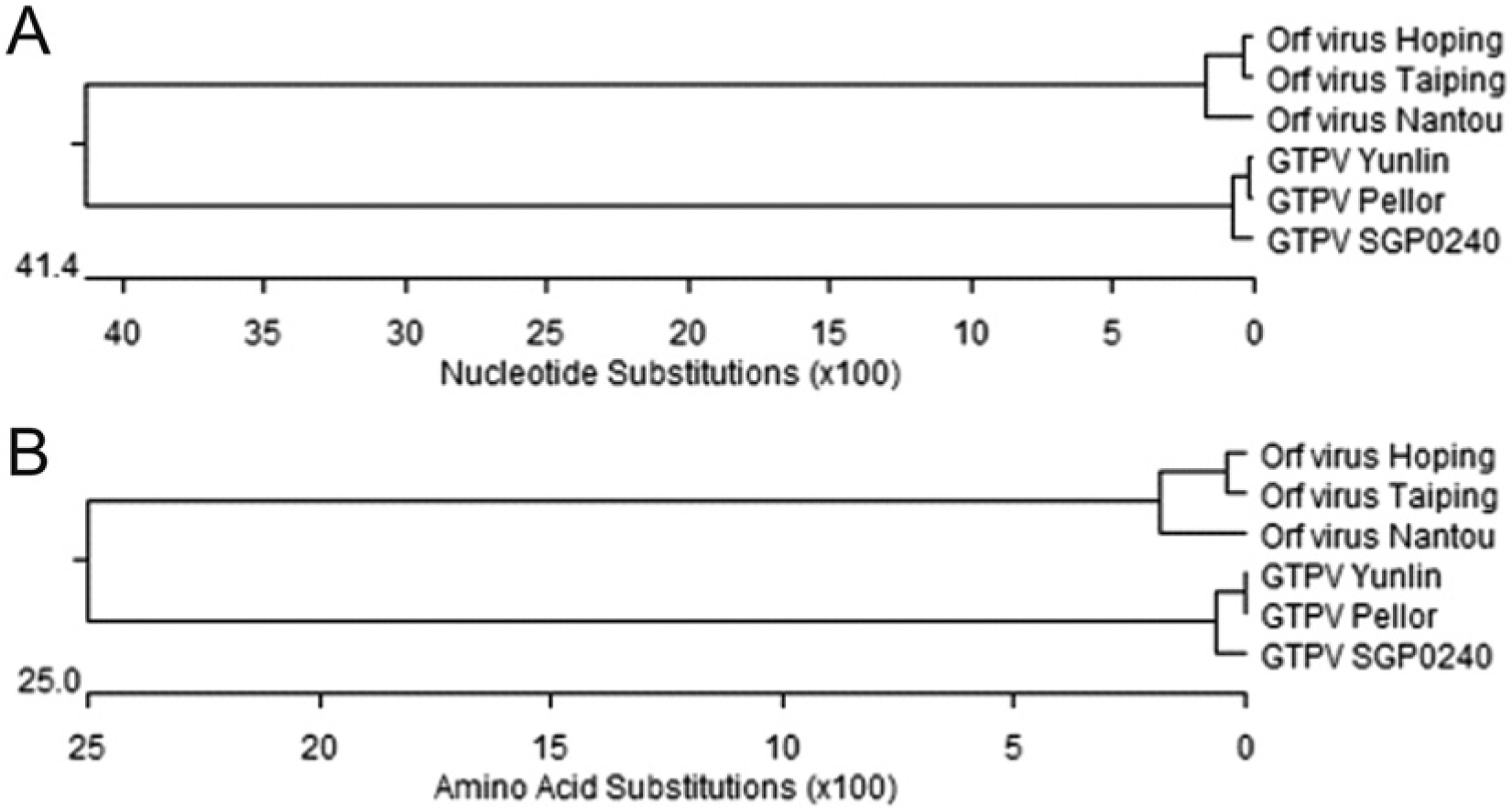
The A32L nucleotide sequence identities between the Yunlin GTPV strain and the SGP0240 and Pellor GTPV strains are 98.6% and 99.7%, respectively. The Yunlin GTPV strain is relatively closer to the Pellor GTPV strain with a divergence of only 0.3%. This divergence is due to the presence of a nucleotide transversion of G → A at position 60 of the Yunlin strain. However, nucleotide substitutions between the Yunlin and SGP0240 GTPV strains were found at positions of 60, 75, 90, 244, 279, 333, 447, 575, 674, 735, and 749, which results in a much higher sequence divergence of 1.5%. Taken together, it seems the endemic GTPV strain in Taiwan is closest to the Pellor strain phylogenetically. As expected, when the nucleotide sequences of the GTPVs were compared with those of the 3 ORFV strains from Taiwan, the nucleotide divergences were found to range from 83.1% to 83.9%. When the amino acid sequences of the GTPV A32L protein were explored, the percent identity of the Yulin strain to the Pellor and SGP0240 strains are 100% and 98.8%, respectively. Furthermore, the sequence divergence between the GTPV and ORFV strains are at least 49.3% (data not shown). Such similarities are reflected in phylogenetic relationships at both nucleotide and amino acid levels based on the neighbor-joining algorithm (Fig. 4A, 4B).
HRMA for the differential diagnosis of GTPV using clinical samples
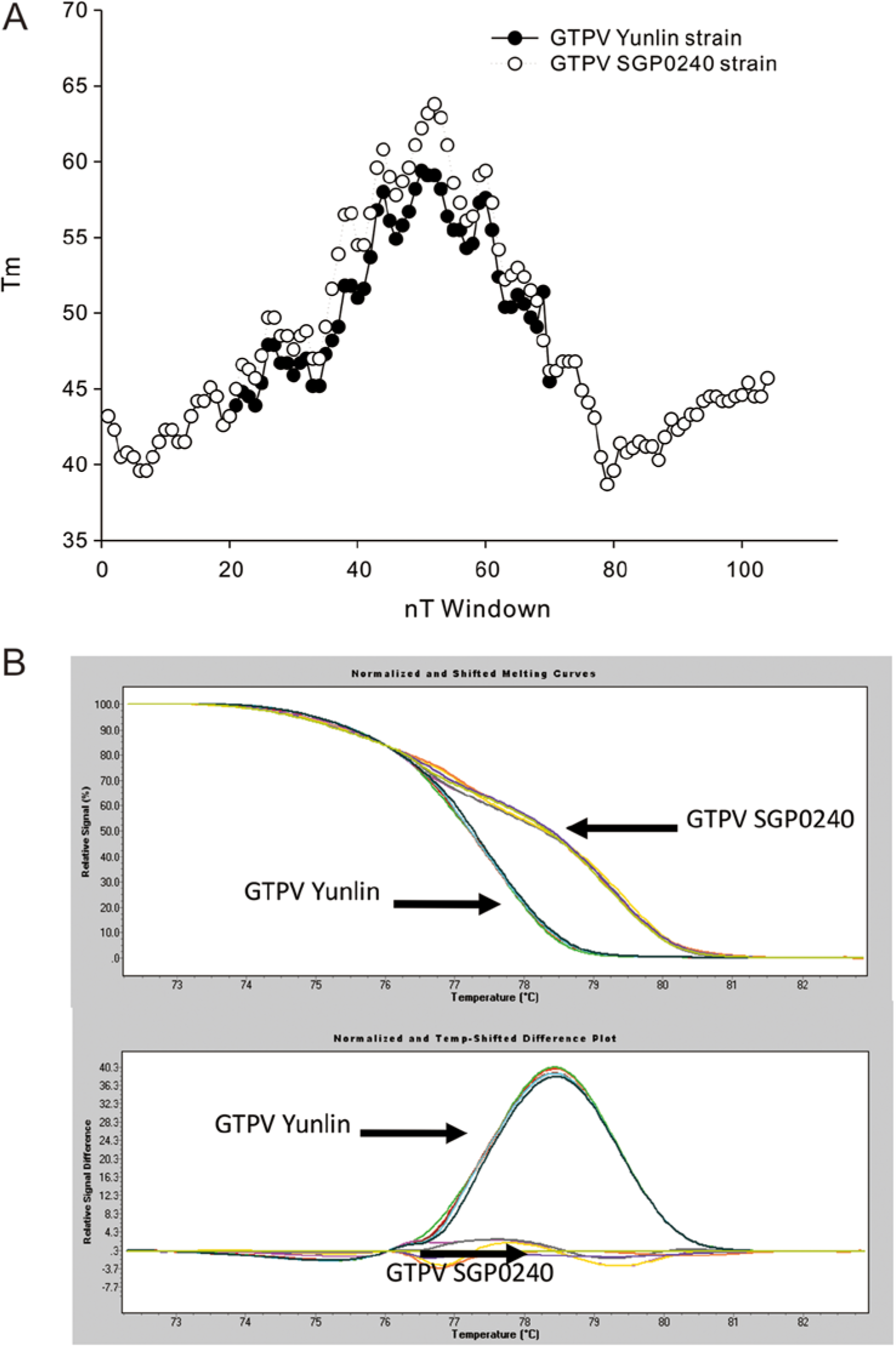
Distinct melting temperature profiles were predicted from the Yunlin and SGP0240 GTPV strains based on the EMBOSS suite algorithm. The difference in melting temperature profiles of the strains affected nucleotide positions 21–70 (Fig. 5A). Results of the HRMA on DNA amplified from the skin lesion DNA of 6 goats with the GTPV infection and that from the vaccine strain showed that the GTPV strains fall into 2 groups on the normalized curve plot (the upper chart of Fig. 5B). A shifted melting curve plot from 2 GTPV groups was constructed by subtracting the melting curve shape of products amplified from the SGP0240 GTPV strain from the curve of products amplified from the skin samples of 6 goats with the Yunlin GTPV infection. This profile, obtained from skin samples, provides a clear-cut differentiation between Yunlin and SGP0240 GTPV strains (the bottom chart of Fig. 5B).
Discussion
There are at least 2 GTPV strains (the Yunlin and SGP0240 strains) coexisting in Taiwan at the moment. This is complicated by occasional outbreaks of orf infection in Taiwan, which has become an endemic on goat farms since 2009. 8 The clinical lesions of goat pox and orf diseases are similar, being vesiculoproliferative lesions of the skin of the lips and muzzle of goats and sheep. 8 Animals infected with GTPV show high morbidity and mortality, which results in severe economic losses; in contrast, orf infections result in milder symptoms and have a lower mortality.5,6 It is still difficult to distinguish these 2 diseases especially in the early stages of infection. 16 A live attenuated vaccine can provide a more durable and efficacious protection against goat pox. 14 To date, 2 GTPV vaccines, the AV41 and the SGP0240 strains, are available.14,17 Serious complications including skin nodules and deaths have been reported in the AV41 strain–vaccinated goats coinfected with a wild-type GTPV. 17 One possible explanation for these observations is a recovery of virulence by the vaccine strain, while another is that a novel virus strain might have arisen by genetic recombination between the AV41 vaccine strain and a GTPV wild strain.1,17 Based on the fact that goats without clinical signs during the incubation period of goat pox may still be vaccinated, more serious delayed type hypersensitivity may present as skin maculae in some vaccinated goats already infected with GTPV, and it is not possible to exclude recombination events involving the SGP0240 strain. Thus, in these circumstances, an effective tool to differentiate infection from vaccination is necessary.8,14 This will also be useful in a situation where an indigenous vaccine needs to be differentiated from a field challenge in the future. 15
Several conditions for the multiplex PCR were optimized. First, primers were purified by using an oligonucelotide purification cartridge rather than by desalting to efficiently remove the impure oligonucleotide fragments may interfere with the PCR reaction. Second, the concentration of Mg2+ was increased to 10 mM. Third, an approximate 2-fold higher concentration of the upstream primers was used to avoid a competitive interference from both the downstream primers for GTPV and the ORFV. Fourth, because the melting temperatures of various sets of GTPV and ORFV primers are not similar, the ratios of the concentrations of the GTPV primers relative to the ORFV were approximately 2–3-fold high in order to ensure the similar amplification efficiency. Compared with the traditional approach, namely PCR amplification and sequencing of the PCR product, the multiplex PCR was rapid (3 hr vs. 2 days) and inexpensive (US$10 vs. US$18). Furthermore, using the higher saturation capability of DNA binding dyes, HRMA was shown to be able to differentiate the Yunlin and SGP0240 GTPV strains from the skin samples based on the partial conserved A32L genes of both strains. Three nucleotide transversions (A/T ↔ G/C) within a total 123 nucleotides in this region were sufficient to change the melting curve profile behavior between the both strains.
In conclusion, the developed multiplex PCR and HRMA are able to serve as the front-line and confirmative assays, respectively, with good sensitivity and specificity. Both these systems will be especially useful as part of the goat pox eradication program in Taiwan.
Footnotes
Acknowledgements
The authors thank Dr. Ralph Kirby for a critical reading of the article.
a.
Qiagen GmbH, Hilden, Germany.
b.
Protech, Taipei, Taiwan.
c.
Geneaid, New Taipei, Taiwan.
d.
pGEM-T vector, Promega Corp., Madison, WI.
e.
Promega Corp., Madison, WI.
f.
DNASTAR Inc., Madison, WI.
g.
Protech, Taipei, Taiwan.
h.
Transition Technologies, Toronto, Ontario, Canada.
i.
BioFire Diagnostics, Salt Lake City, UT.
j.
Transition Technologies, Toronto, Ontario, Canada.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
