Abstract
Formalin-fixed, paraffin-embedded (FFPE) tissues represent a unique source of archived biological material, but obtaining suitable DNA and RNA for retrospective “-omic” investigations is still challenging. In the current study, canine tumor FFPE blocks were used to 1) compare common commercial DNA and RNA extraction kits; 2) compare target gene expression measured in FFPE blocks and biopsies stored in a commercial storage reagent; 3) assess the impact of fixation time; and 4) perform biomolecular investigations on archival samples chosen according to formalin fixation times. Nucleic acids yield and quality were determined by spectrophotometer and capillary electrophoresis, respectively. Quantitative real-time polymerase chain reaction assays for the following genes: BCL-2–associated X protein, B-cell lymphoma extra large, antigen identified by monoclonal antibody Ki-67, proto-oncogene c-KIT (c-kit). Two internal control genes (Golgin A1 and canine transmembrane BAX inhibitor motif containing 4), together with direct sequencing of c-kit exons 8, 9, 11, and 17, were used as end points. Differences in DNA/RNA yield and purity were noticed among the commercial kits. Nucleic acids (particularly RNA) extracted from paraffin blocks were degraded, even at lower fixation times. Compared to samples held in the commercial storage reagent, archived tissues showed a poorer amplification. Therefore, a gold standard protocol for DNA/RNA isolation from canine tumor FFPE blocks for molecular investigations is still troublesome. More standardized storage conditions, including time between sample acquisition and fixation, fixation time, and sample thickness, are needed to guarantee the preservation of nucleic acids and, then, their possible use in retrospective transcriptomic analysis.
Formalin fixation and paraffin embedding is the method of choice in combined diagnostic and research settings. Formalin preserves the morphological tissue architecture and allows protein analysis (e.g., by immunohistochemical staining). 6 The analysis of archived formalin-fixed, paraffin-embedded (FFPE) specimens with known diagnosis, prognosis, response to therapy, and outcome represents a valuable resource for retrospective studies aiming to identify biomarkers for risk prediction of disease or prognosis.6,17
Since 2000, rapid advances in profiling DNA, RNA, and proteins by using high-throughput platforms have been made. Nucleic acids recovered from archived FFPE tissues are suitable for a variety of downstream genetic and epigenetic analyses (genomics, transcriptomics, epigenomics). 6 Genomic DNA can be assayed by polymerase chain reaction (PCR), array comparative genomic hybridization, massive parallel sequencing, and methylation assays. Likewise, messenger RNA can be used for quantitative real-time reverse transcription polymerase chain reaction (qPCR), high-throughput gene expression, and even massive parallel sequencing.16,17 Overall, these methodologies hold the promise of rapid biomarker discovery and screening. Therefore, the application of these techniques to FFPE, to improve prognostic and predictive power, has been the subject of intense research, particularly in the field of translational cancer research.6,16
Unfortunately, the isolation of adequate amounts of intact DNA/RNA from FFPE blocks is still challenging. Apart from the known toxic properties for human health, the formalin fixation process results in the formation of chemical cross-links between DNA/RNA and/or proteins and monomethylol groups that are added to nucleotide base pairs. 11 These processes increase the susceptibility of nucleic acids to shearing and fragmentation; consequently, the obtained nucleic acids are often degraded and present in short fragmented segments. This phenomenon can be caused by a number of other factors, such as the time period between biopsy punch and effective fixation, fixation time, the pH of the fixative agent, sample thickness, tissue autolysis, incubation at higher temperature during the paraffin embedding process, and prolonged storage of FFPE blocks.2,16,31
Since 2000, a number of methods for optimal nucleic acid isolation and subsequent assay starting from human FFPE specimens of different origin and fixation times have been developed.2,17,31 This was of particular interest for gene expression profiling,5,18,29 as RNA extraction from FFPE tissues posed some challenges in this regard.6,16 Nevertheless, despite the increasing interest in the use of “-omic” techniques with archived FFPE tissues, little information has been published so far in veterinary medicine and particularly on the dog.15,22,28,34 This gap in knowledge is significant, as the dog represents, for several reasons, an excellent comparative model of complex human diseases, such as naturally occurring cancer. 26 Therefore, the aim of the present study was the optimization of a protocol for DNA/RNA isolation from canine tumor FFPE blocks.
Four different but consecutive steps were evaluated (Fig. 1): 1) the evaluation of different nucleic acid extraction kits; 2) the comparison of target gene expression profiles derived from tumor biopsies stored in a commercial storage solution a and FFPE blocks; 3) the effects of formalin fixation time on extracted nucleic acid in terms of both qualitative and quantitative traits; and 4) the execution of biomolecular investigations on mast cell tumor (MCT) archival samples chosen according to formalin fixation times.
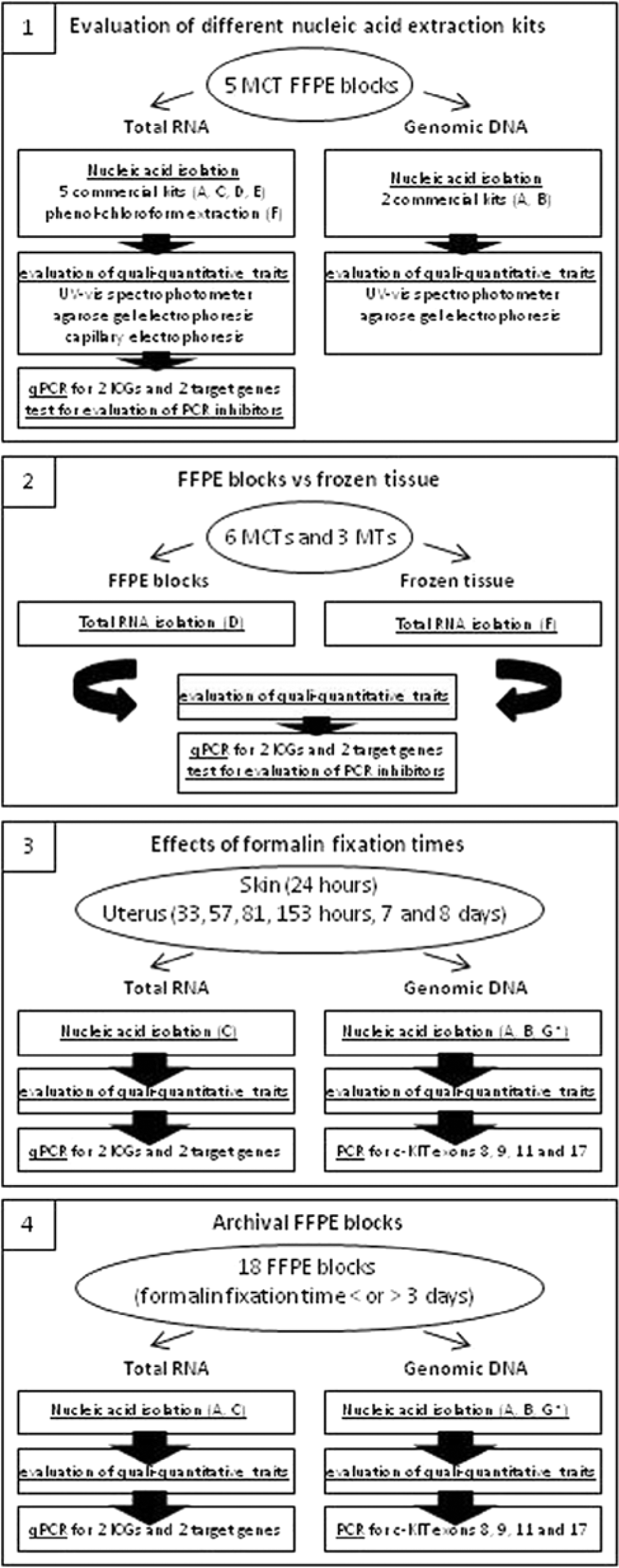
Flowchart showing, for each experimental step (1–4), the sample characteristics, methods, and polymerase chain reaction assays that have been used. MCT = mast cell tumor; FFPE = formalin-fixed, paraffin embedded; qPCR = quantitative real-time reverse transcription polymerase chain reaction; ICGs = internal control genes; MT = mammary tumor; c-kit = proto-oncogene c-KIT.
With the exception of step 3, molecular analyses were carried out on 28 MCT and 3 mammary tumor (MT) biopsies collected either at surgery or necropsy. The complete list of tumors enrolled in the study is reported in Tables S1–S3 of the supplemental material online (Supplementary File 1). Tumors were histologically classified according to the guidelines provided by the World Health Organization,12,20 while grading was assigned in accordance with published guidelines. 25 Immunohistochemical staining for the detection of Ki-67 and c-kit proteins was performed as previously described.9,32
For step 1, in which a comparative evaluation of different nucleic acid extraction kits was undertaken, FFPE sections (n = 3, 10 µm each) derived from 5 distinct MCTs were used. The MCTs were graded as either G1 (4 tumors) or G2 (1 tumor: see Table S1). Five commercial kits were used: kit A, b kit B, c kit C, d kit D, e and kit E. f The first 2 kitsb,c extracted both RNA and DNA. Nucleic acids extraction protocols were similar, with some differences in time of execution, mostly due to the length of proteinase K digestion step. Besides the aforementioned kits, the classical phenol–chloroform g extraction protocol was also tested. All kits required a deparaffinization step, brought about either by treatment with xylene and ethanol (kits B, C, D, and E) or by using a specific solution supplied with the kit itself (kit A). h The extracted RNA yield and purity were measured by a spectrophotometer, i while the RNA integrity was checked by 1% agarose gel electrophoresis and by capillary electrophoresis. j Total RNA was reverse transcribed using a commercial kit. k A qPCR approach was used to detect differences in the threshold cycle (Ct) value of the extracted RNA. The complementary DNA was tested by using qPCR assays for 2 internal control genes (ICGs) and 2 target genes, canine transmembrane BAX inhibitor motif containing 4 (CGI-119) and Golgin A1 (GOLGA1), and BCL-2–associated X protein (Bax) and B-cell lymphoma-extra large (Bcl-xL), respectively. Reactions were run on a PCR machine, l by using commercial probes m and a commercial master mix. n The primers for Bax and Bcl-xL and the corresponding probe as well as the amplicon sizes are reported in Table S4. The qPCR reaction is subject to inhibition by substances (e.g., solvents used for extraction) that copurify with nucleic acids during their isolation. 7 Thus, a qPCR-based test 23 was used to check for the presence of possible qPCR reaction inhibitors.
In step 2, a comparison between target gene expression profiles derived from biopsies stored in a commercial storage reagent a (at −20°C) and corresponding FFPE tissues was made. Six MCTs and 3 MTs, for which both stored material and FFPE blocks were available, were considered. The 6 MCTs were graded as G1 (2 tumors) or G2 (4 tumors) while the MTs were a simple tubular, a complex tubular, and a squamous carcinoma (see Table S2). Total RNA was extracted from frozen specimens and FFPE sections (n = 3, 10 µm each) by using a commercial lysis buffer g for the classical phenol–chloroform nucleic acids extraction and a commercial kit, d respectively. Methods to measure RNA concentration, purity, and integrity as well as to obtain complementary DNA (cDNA) were the same mentioned for step 1. Equal amounts of cDNA were used to compare the commercial storage reagent a and the FFPE sourced specimens. Target genes were the same as mentioned above.
In step 3, the effect of different formalin fixation times was investigated. Uterine and skin surgical samples were fixed in 10% buffered formalin for 24 (skin only), 33, 57, 81, 153 hr, 7 days, and 8 days. Samples were then embedded in paraffin and processed. Nucleic acids were extracted from FFPE sections (n = 3, 10 µm each) by using 2 RNAb,d and 3 DNAb,c,o commercial kits. The assessment of nucleic acids quality, quantity, and integrity as well as reverse transcription were made as reported in step 1. The suitability of the messenger RNA for amplification was measured by using qPCR assays for 2 ICGs (i.e., CGI-119 and GOLGA1) 1 and 2 target genes (Ki-67 and c-kit).9,32 On the other hand, the suitability of the DNA for amplification was evaluated by PCR end-point amplification of c-kit exons 8, 9 11, and 17, whose primers and amplicon sizes are reported in Table S5.
In the last step (4), molecular investigations on archival MCT FFPE tissues, chosen according to formalin fixation time, were performed. Based on step 3 results, 2 sets of FFPE blocks were chosen, and a formalin fixation time of either shorter or longer than 3 days was used as the discriminatory criterion (see Table S3). All the tumors were assessed as G2, and 9 had a fixation time of less than 3 days while 8 had a fixation time of more than 3 days. Nucleic acids were extracted as reported in step 3 except for DNA, for which only 1 kit was used. b Nucleic acid quality, quantity, and integrity was determined as reported in step 1. Likewise, reverse transcription was performed as in step 1. The suitability of the nucleic acids for amplification was evaluated by using the same approach reported in step 3.
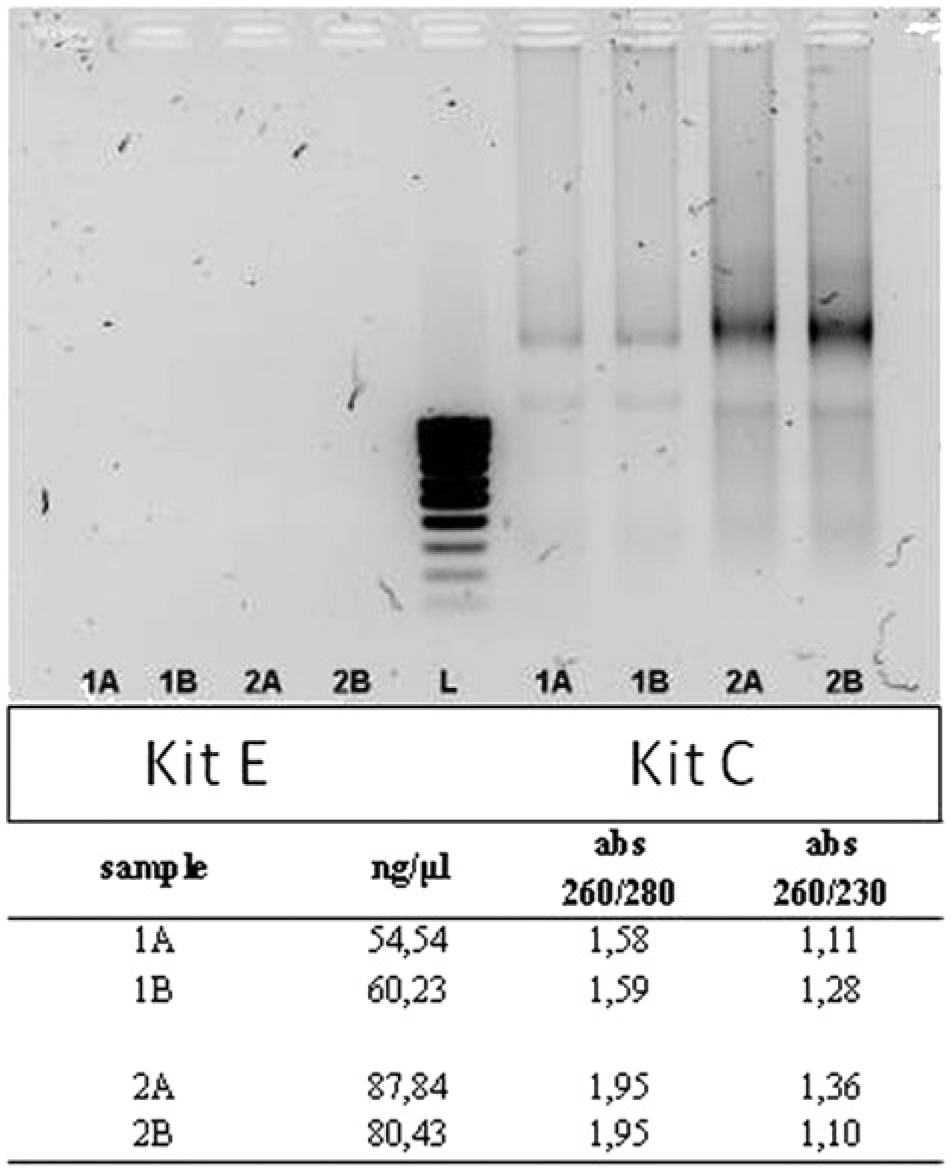
Limited amounts of a highly fragmented RNA were obtained in step 1. The qualitative and quantitative features of the nucleic acid were not comparable to those obtainable by specimens stored in the commercial storage solution, a although in some instances the features were considered acceptable (data not shown). Further experiments were carried out to solve this problem (e.g. increasing either the number of FFPE sections or the duration of proteinase K digestion). Moreover, distributor technical services were sought for advice. Finally, fairly acceptable results were obtained with kit C d only. The agarose gel electrophoresis analysis of 2 samples (A, B), extracted twice with kits C d and E, f as well as data referring to RNA concentration and purity, are shown in Figure 2. Amplification results were anything but encouraging—Ct values were high, and duplicates differed from each other. Similar results were obtained for both ICGs and target genes. Moreover, the qPCR-based test, 23 used to check for the presence of possible qPCR reaction inhibitors in the samples, was negative (data not shown). Based on these results, the only kit giving satisfactory results, kit C, d was used in steps 2–4.
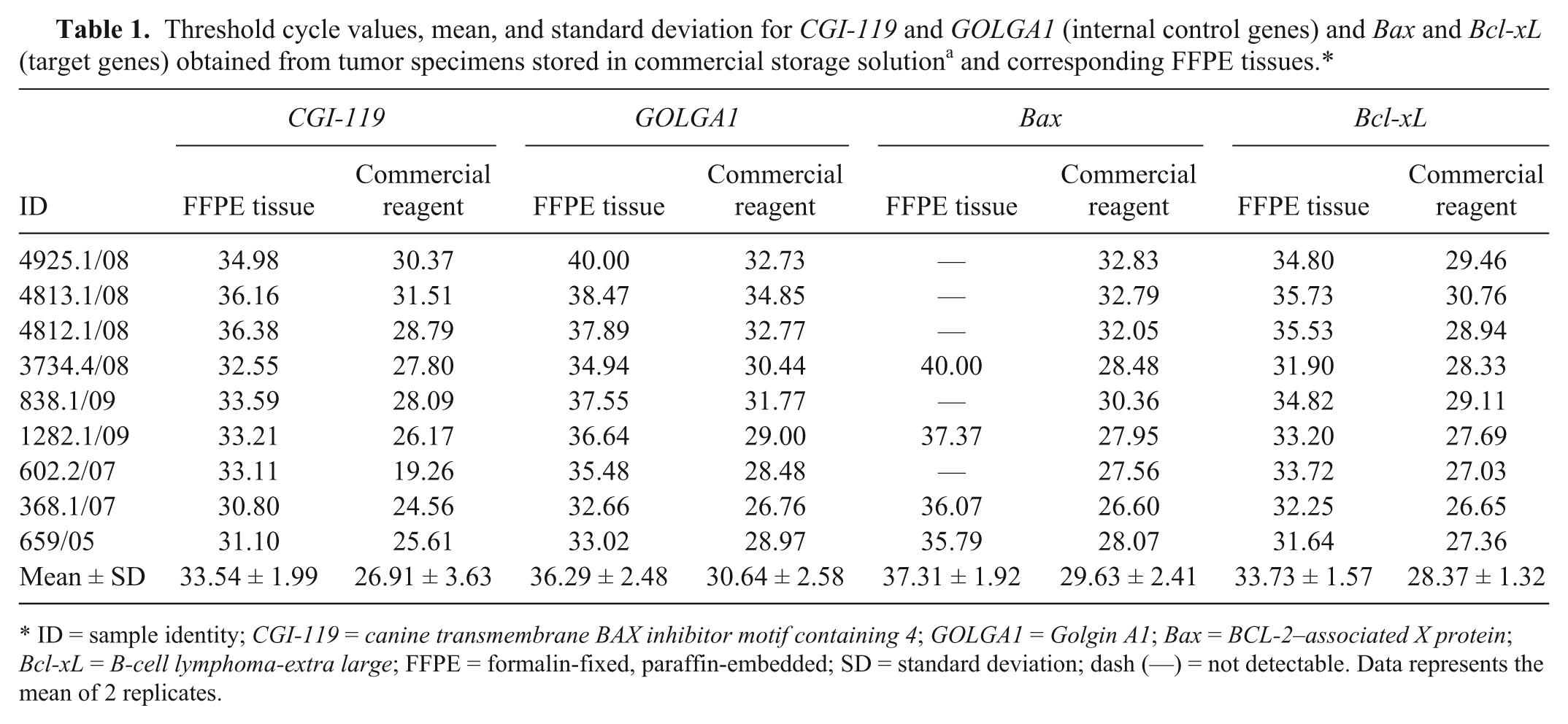
In step 2, a comparison of target gene expression profiles derived from biopsies stored in a commercial storage solution a (−20°C) and corresponding FFPE blocks was made. Higher Ct values (up to 10) were obtained in FFPE tissues (see Table 1). These results are clear evidence of a low qPCR reaction efficiency, not attributable to the presence of inhibitors (the inhibitory qPCR test was negative: data not shown). Contradictory results of Bax gene (no amplification in 5 samples, Ct value higher than 40 in another one) might be due to the minor constitutive expression of this gene. The qPCR analyses were conducted using the same amount of cDNA (in accordance with the validation of each qPCR assay). Despite this precaution, qualitative data (presence or absence of amplification products and corresponding Ct value) indicate that RNA extracted from FFPE tissues was comparatively of poor quality (data not shown). In Figure 3, ultraviolet (UV)–visible spectra and RNA integrity number (RIN, acceptable when higher than 7.0) of 3 tumor specimens are shown.
ID = sample identity; CGI-119 = canine transmembrane BAX inhibitor motif containing 4; GOLGA1 = Golgin A1; Bax = BCL-2–associated X protein; Bcl-xL = B-cell lymphoma-extra large; FFPE = formalin-fixed, paraffin-embedded; SD = standard deviation; dash (—) = not detectable. Data represents the mean of 2 replicates.
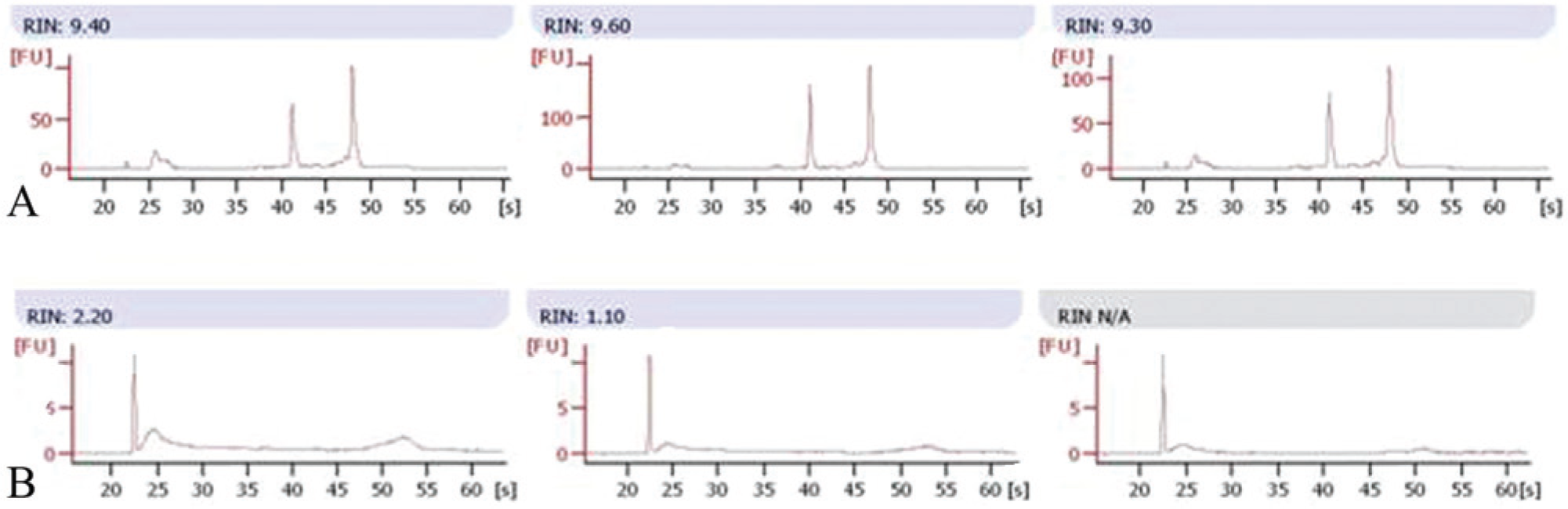
Representative ultraviolet–visible electropherogram curves and RNA integrity number (RIN) of 3 tumor specimens.
In step 3, the effect of different formalin fixation times was investigated. Concerning RNA, the time of formalin fixation did not affect RNA concentration and purity, except for a slight decrease in the amount of extracted nucleic acid and a possible contamination by genomic DNA (260/280 ratio <2.0). Paradoxically, the best results (in terms of presence of 28S and 18S ribosomal bands) were obtained for times higher than 33 hr (see Supplemental Fig. S1). Nonetheless, the results of the UV–visible spectra analysis suggest that RNA was for the most part degraded (data not shown). This result was supported by the results of the qPCR work: the longer the fixation time, the higher the Ct value obtained for candidate genes, with negligible differences between genes (see Fig. 4). Altogether, the data suggests that RNA extraction and the resultant amplification performance are essentially arbitrary. Consequently, a specific causative factor for such a behavior cannot be identified. With regard to DNA, the time of fixation altered the nucleic acid quality (absorbance ratio 260/280 and 260/230, presence of degradation on gel electrophoresis) and quantity (determined by spectrophotometry). Indeed, the best results were obtained after 24 (skin) or 33 and 57 hr (uterus) of storage in buffered formalin (see Supplemental Fig. S2). Exons 8, 9, 11, and 17 of the c-kit gene were used to amplify the extracted nucleic acid, as their different length (400, 450, 650, and 271 bp, respectively) allowed the fine evaluation of amplification efficiency. The data obtained confirmed that prolonged fixation time negatively influences nucleic acid preservation. As shown in Figure 5, the shortest formalin fixation time resulted in better amplification efficiency, except for exon 17, whose pattern of amplification seemed to increase with the increasing time of fixation. As a whole, DNA performances (extraction and subsequent amplification) were better than those noted for RNA. The time of fixation affected the quality of nucleic acids and, consequently, their suitability for amplification (i.e., the shorter time of fixation, the better the PCR performances).
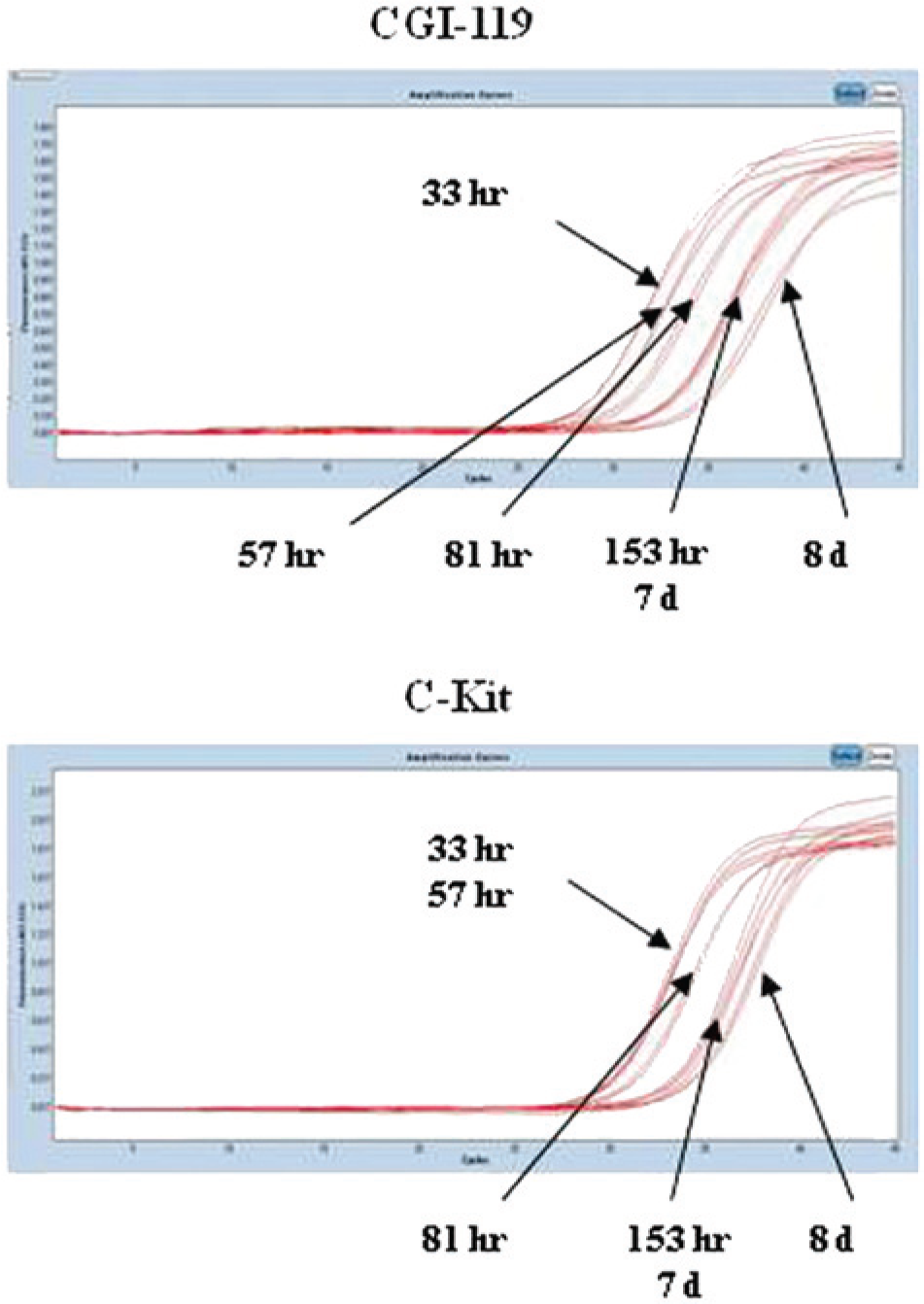
Amplification curves of canine transmembrane BAX inhibitor motif containing 4 (CGI-119; an internal control gene) and proto-oncogene c-KIT (c-kit; a target gene), obtained from uterine specimens subjected to different formalin fixation times before embedding in paraffin. d = days; hr = hours.
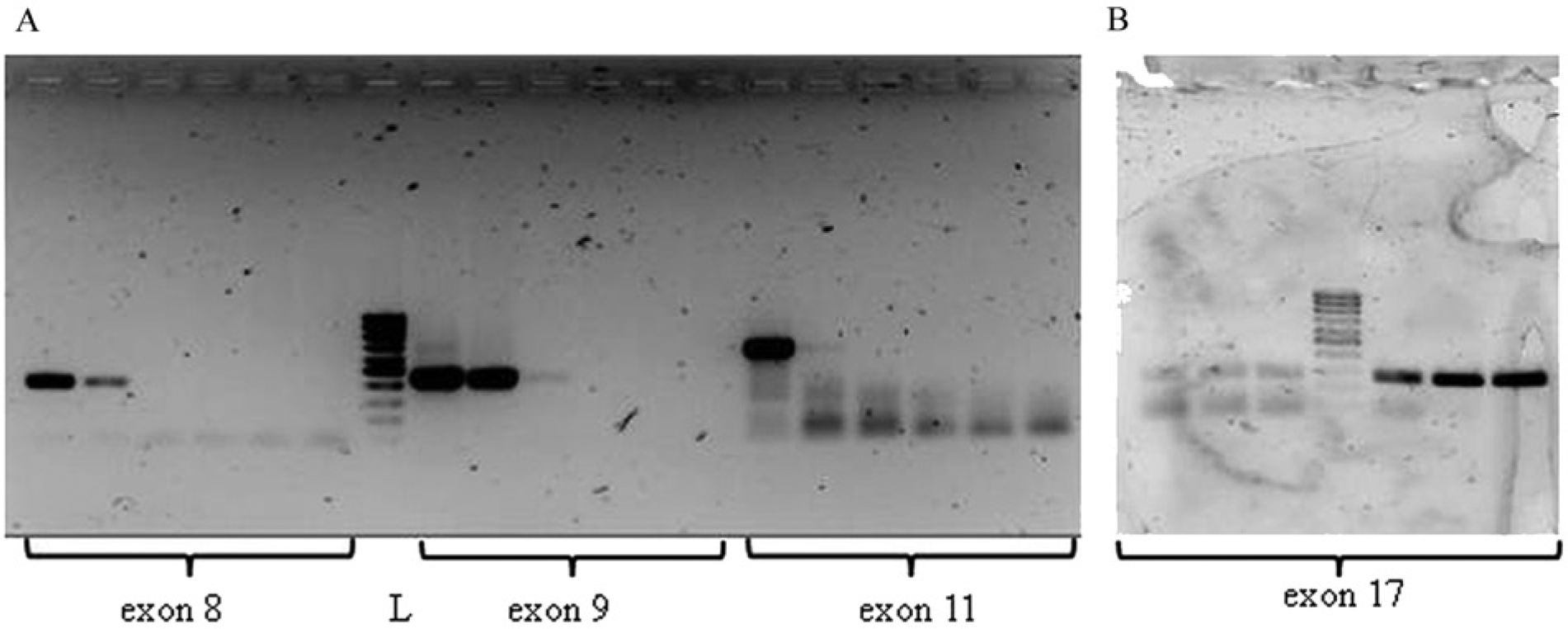
Amplification of proto-oncogene c-KIT exons 8, 9, 11 (~400, 450, 650 bp;
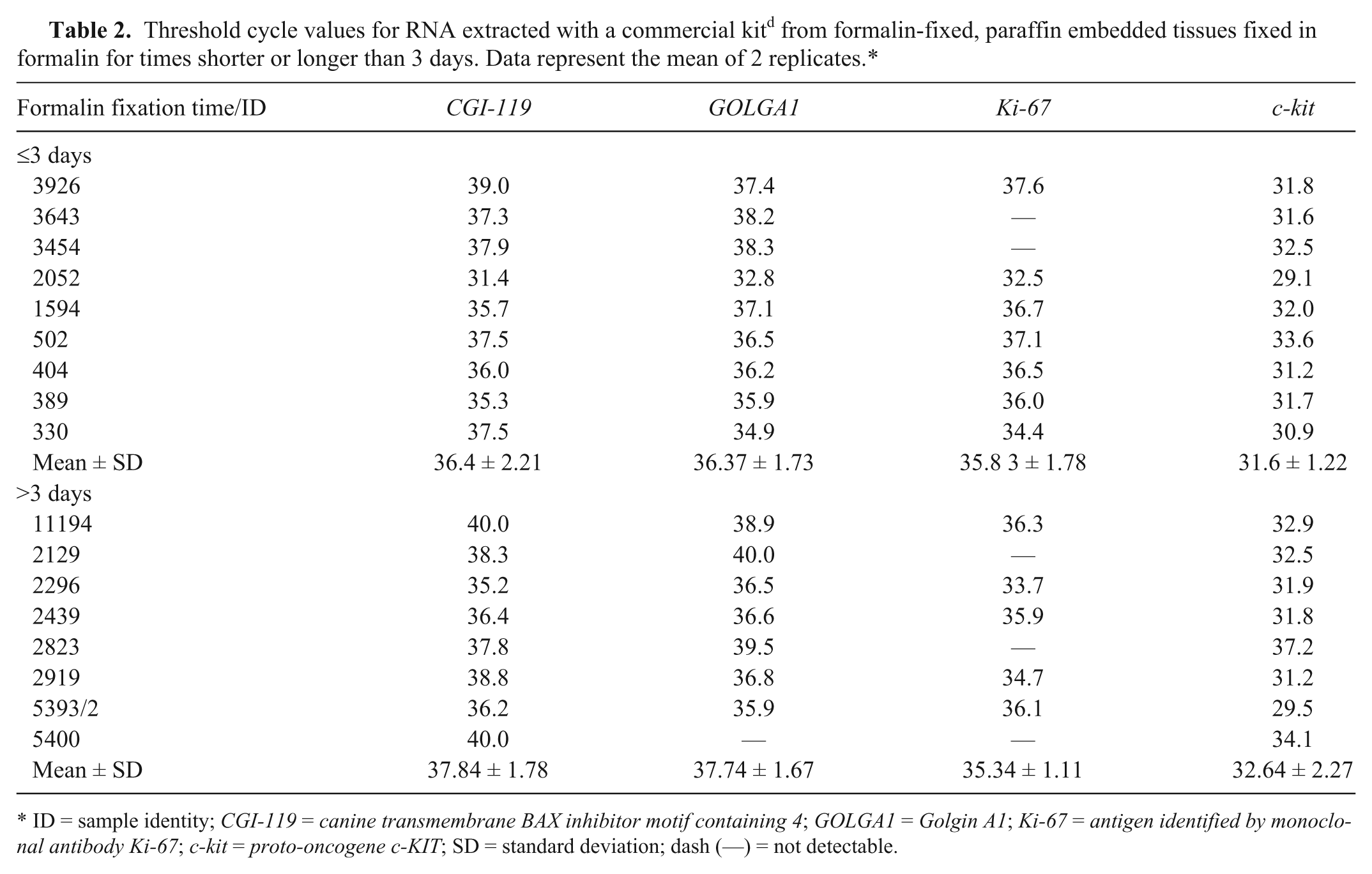
Finally, in step 4, molecular investigations were performed on archival MCT FFPE tissues, chosen according to results obtained in step 3 (i.e., a formalin fixation time of either shorter or longer than 3 days). Two different kits were used for RNA extraction (kits A b and C d ). The best performances were obtained with kit C. d If compared with samples stored in the commercial storage solution, a minor RNA amounts of poor quality were extracted from FFPE tissues (data not shown). At the amplification stage, high Ct values were obtained. An arbitrary pattern of amplification was noticed for both ICGs and target genes, and differences higher than 0.5 Ct were found among replicates. No correlation was ever found between the results of amplification and formalin fixation time. As a whole, these results are more likely the consequence of either a poor RNA quality (e.g., fragmentation) or other interfering factors, such as the sampling procedure, sample dimensions and thickness, and maintenance and/or storage temperature. In Table 2, Ct values obtained for RNA extracted with kit C d are reported. The fixation in formalin affected DNA yield and purity. The agarose gel electrophoresis (presence of smear) and UV–visible spectra results (very low RIN values) supported this finding (data not shown). The amplification of c-kit exons 8, 9, and 11 gave for the most part negative or variable results, independent of the time of fixation. By contrast, exon 17 (the shortest exon 271 bp) was always amplifiable, except for 3 samples (data not shown).
ID = sample identity; CGI-119 = canine transmembrane BAX inhibitor motif containing 4; GOLGA1 = Golgin A1; Ki-67 = antigen identified by monoclonal antibody Ki-67; c-kit = proto-oncogene c-KIT; SD = standard deviation; dash (—) = not detectable.
As a whole, the results of the current study indicate that DNA/RNA extraction from dog tumor FFPE blocks is still challenging. For RNA extraction, the concentration was generally low, the RIN value less than 2.5 or undetectable, and electrophoresis showed a diffuse smear with 28S and 18S ribosomal bands barely detectable. Although methods used to assess the nucleic acid quality (absorbance ratio 260/280 and 260/230, presence of degradation on gel electrophoresis and capillary electrophoresis) and quantity (determined by UV–visible spectrophotometer) have been disputed, 33 the results of the current study confirm, by and large, that only minor amounts of nucleic acid were obtained from FFPE blocks. Moreover, both nucleic acids (and RNA in particular) were degraded and/or fragmented, likewise to formerly available data.13,16
When compared with specimens stored in solution offering RNA stabilization and protection, thus enabling gene expression profiling investigations, the use of FFPE tissues gave a much reduced qPCR reaction efficiency. Such a result confirms previously published evidence, 27 although some contradictory results have been published. 21
Formalin fixation is a time-dependent process that negatively influences nucleic acid preservation, 10 though no alterations in DNA integrity have also been noticed for specimens fixed for periods of 30 min and up to 20 hr. 8 In the present study, the time of fixation did not interfere with extracted RNA and subsequent use in qPCR assays. In fact, comparable results (for the most part, fragmentation) were obtained in specimens left in formalin from 24 hr and up to 8 days as well as in archival FFPE blocks derived from samples fixed for less or more than 3 days. Thus, the cause of nucleic acids fragmentation could be hypothetically assigned to the work conditions adopted during sample collection and FFPE processing. Notably, nucleic acid integrity increases if the biopsy is kept on ice and then fixed in formalin at low temperatures (+4°C); moreover, this cold-fixation procedure permits the amplification of sequences longer than 600 bp. 4 Although not yet reaching the nucleic acid quality obtained from fresh-frozen material, such an approach might represents an alternative way to enable reliable gene expression profiling of FFPE tissues. The fixation process per se was shown to alter likewise DNA yield, purity, and PCR performances. In particular, negative or inconsistent results were obtained with the amplification of the c-kit exons, except for the shortest exon (exon 17, 271 bp). The amplicon size is another limiting factor in archival FFPE downstream analyses. It is estimated that up to 200–300-bp long DNA fragments are likely to be amplified by using an optimal nucleic acid isolation protocol.3,19,24 In comparison to DNA, the current study has shown that RNA is more susceptible to degradation and that subsequent qPCR reactions were troublesome (high Ct values), even though the amplicon length was always lower than 100 bp. This was not surprising, as RNA is a single-stranded molecule, and a comparatively higher fragmentation of RNA is usually associated with the addition of methylol groups during fixation.6,19,27
In conclusion, the extraction of DNA/RNA from canine FFPE blocks for “-omic” investigations remains problematical. The extracted RNA can be degraded and fragmented, regardless of the extraction kit used, with resultant unsuccessful qPCR performances. Similar problems apply to DNA extraction, even though amplification was obtained with small fragments (i.e., approximately 200 bp). The results of the present study suggest that some other factors are likely to affect nucleic acids yield, integrity, and suitability for subsequent analysis. These factors include the time of paraffin embedding, the age of FFPE material (e.g., months, years), the length of the proteinase K digestion step, and warm tissue ischemia.19,30 The results of the current study, together with the very restricted information actually available in veterinary medicine, strengthen the need for validating the best conditions for collection of surgical tumor biopsies, their fixation, inclusion, storage, and downstream processing. Such an objective might be pursued through the establishment of recommended guidelines, such as those developed already for submission, trimming, margin evaluation, and reporting of tumor biopsies 14 and, then, the execution of multicenter validation studies, as already done in human medicine. 3
Footnotes
Acknowledgements
The authors would like to thank Prof. Paolo Buracco and Drs. Emanuela Morello and Antonella Vercelli for their contribution in providing some of the tumor samples included in the present investigation. Anna Granato and Mery Giantin contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by grants from Istituto Zooprofilattico Sperimentale delle Venezie (RC IZSVe 10/09: Studio retrospettivo da tessuti fissati in formalina ed inclusi in paraffina per la valutazione di biomarcatori prognostici e predittivi nella patologia neoplastica del cane) to Anna Granato, Mauro Dacasto, and Marta Vascellari.
a.
RNAlater, Life Technologies Europe BV, Monza, Milan, Italy.
b.
AllPrep DNA/RNA FFPE kit, Qiagen SpA, Milan, Italy.
c.
NucleoSpin FFPE RNA/DNA, Macherey-Nagel GmbH & Co. KG, Düren, Germany.
d.
High Pure RNA Paraffin kit, Roche SpA, Milan, Italy.
e.
High Pure FFPE RNA Micro kit, Roche SpA, Milan, Italy.
f.
Ambion RecoverAll Total Nucleic Acid Isolation, Life Technologies Europe BV, Monza, Milan, Italy.
g.
TRIzol, Life Technologies Europe BV, Monza, Milan, Italy.
h.
Deparaffinization Solution, Qiagen SpA, Milan, Italy.
i.
Nanodrop ND-1000 spectrophotometer, Thermo Scientific, Erembodegem, Belgium.
j.
Agilent 2100 Bioanalyzer, Agilent Technologies, Cernusco sul Naviglio, Milan, Italy.
k.
High Capacity cDNA Archive kit, Applied Biosystems, Courtaboeuf, France.
l.
LightCycler 480, Roche Diagnostics GmbH, Mannheim, Germany.
m.
Universal Probe Library, Roche Applied Science SpA, Milan, Italy.
n.
LightCycler480 Probes Master, Roche Diagnostics GmbH, Manheim, Germany.
o.
HighPure PCR Template Preparation kit, Roche SpA, Milan, Italy.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.