Abstract
Influenza A virus subtype H1N1 A(H1N1)pdm09 was first confirmed in pigs in the United States in October 2009. In November 2010, lungs and intestines from 2 York piglets from a small, privately owned herd were submitted to the Colorado State University Veterinary Diagnostic Laboratory. The submitting veterinarian reported rapid weight loss and signs of pneumonia in the piglets. Gross lesions included caudoventral pneumonia in both piglets, and histologic lesions in the lungs showed characteristics consistent with influenza virus and bacterial infection. Ribonucleic acid extracted from fresh lung homogenates from both piglets was positive for influenza A(H1N1)pdm09 by a real-time reverse transcription polymerase chain reaction. Virus was isolated from lung homogenates from both piglets in Madin–Darby canine kidney cells, as well as in 10-day-old specific pathogen–free embryonated chicken eggs. Sequence analysis showed 98% homology with 2009 H1N1 human isolates from across the United States and 98% homology against two 2009 and 2010 swine isolates from Nebraska and Minnesota. The current report documents the possible transmission of pandemic influenza A(H1N1)2009 virus [A(H1N1)pdm09] from a human being to a small, privately owned backyard swine herd. The owner was employed as a pharmacist, making occupational exposure to the pandemic influenza A(H1N1)pdm09 a possibility.
Novel Influenza A virus subtype H1N1 (family Orthomyxoviridae, genus Influenzavirus A) emerged in April 2009 and circulated widely among human beings across the globe as a pandemic [A(H1N1)pdm09]. Evaluation of influenza A(H1N1)pdm09 showed that the virus contained human, swine, and avian genetics. 9 Influenza viruses commonly infect swine herds worldwide, and swine workers are recognized as high-risk populations for swine influenza viruses through occupational exposure.6,12–15 Concerns emerged during the H1N1 pandemic that infection of swine with influenza A(H1N1)pdm09 may result in mutation or reassortment of the virus and possible emergence of a more virulent pandemic influenza virus.
In swine, influenza infection usually results in high morbidity but low mortality 10 unless secondary infections are involved. 16 In studies of swine herds infected with influenza A(H1N1)pdm09, affected animals exhibited mild-to-moderate lethargy, anorexia, cough, dyspnea, fever, nasal discharge, and inappetance.5,8,15 The first pigs to be confirmed positive for influenza A(H1N1)pdm09 in the United States were apparently healthy pigs sampled at the Minnesota State Fair in October 2009 (Harless A: 2009, USDA conducting confirmatory testing on possible detection of 2009 pandemic H1N1 influenza in U.S. swine. News Release 0513.09. Available at: http://www.usda.gov/wps/portal/usda/usdahome?contentid=2009/10/0513.xml). An outbreak of influenza in a group of children housed in a dorm at the fair at the same time occurred; however, no direct link between the affected children and apparently health pigs was confirmed. Reports from other countries of influenza A(H1N1)pdm09 infections in pigs detail situations of close human-swine interaction, usually in high-density commercial swine production operations such as in Buenos Aires, Argentina in winter 2009, when a closed herd of 519 sows on a farrow to finish operation was infected with influenza A(H1N1)pdm09. During the outbreak, 30% of nursery pigs and 15% of pigs in growing and fattening barns exhibited clinical signs consistent with influenza A(H1N1)pdm09 infection. Clinical signs of illness lasted for 1 week, with rapid resolution after that time. In this outbreak, it was documented that the farm owner and his wife had reported symptoms of influenza-like illness (ILI) in themselves 10 days earlier. 15
In November 2010, lungs and intestines from 2 York piglets, less than 1 month of age, were submitted for histology and diagnostic testing to the Colorado State University Veterinary Diagnostic Laboratory (Fort Collins, Colorado). The submitting veterinarian performed necropsy on site, and tissues were collected in the field. The submitting veterinarian reported that both piglets had rapidly lost weight and showed signs of pneumonia, and that previous to the submission of these samples, 1 other piglet on the farm had died. All 3 piglets were housed in the same pen of 12 total piglets.
The piglets were from a privately owned backyard herd of 12 sows raised and sold by the owner as club (show) pigs. The owner was employed as a pharmacist and was not vaccinated for influenza A(H1N1)pdm09, making occupational exposure a possibility. The owner’s other family members were all vaccinated. Neither the owner nor any family members reported any history of ILI in the month prior to the piglets becoming ill.
Histologic lesions in the lungs showed characteristics consistent with influenza virus and bacterial infection, including multifocal mucosal necrosis and effacement with neutrophils spilling into the adjacent parenchyma; consolidation of parenchyma by filling of alveolar spaces with neutrophils and histiocytes; loss of alveolar walls in the absence of distinct necrosis; prominent perivascular lymphocytic cuffs; and congestion of alveolar walls (Fig. 1A, 1B).
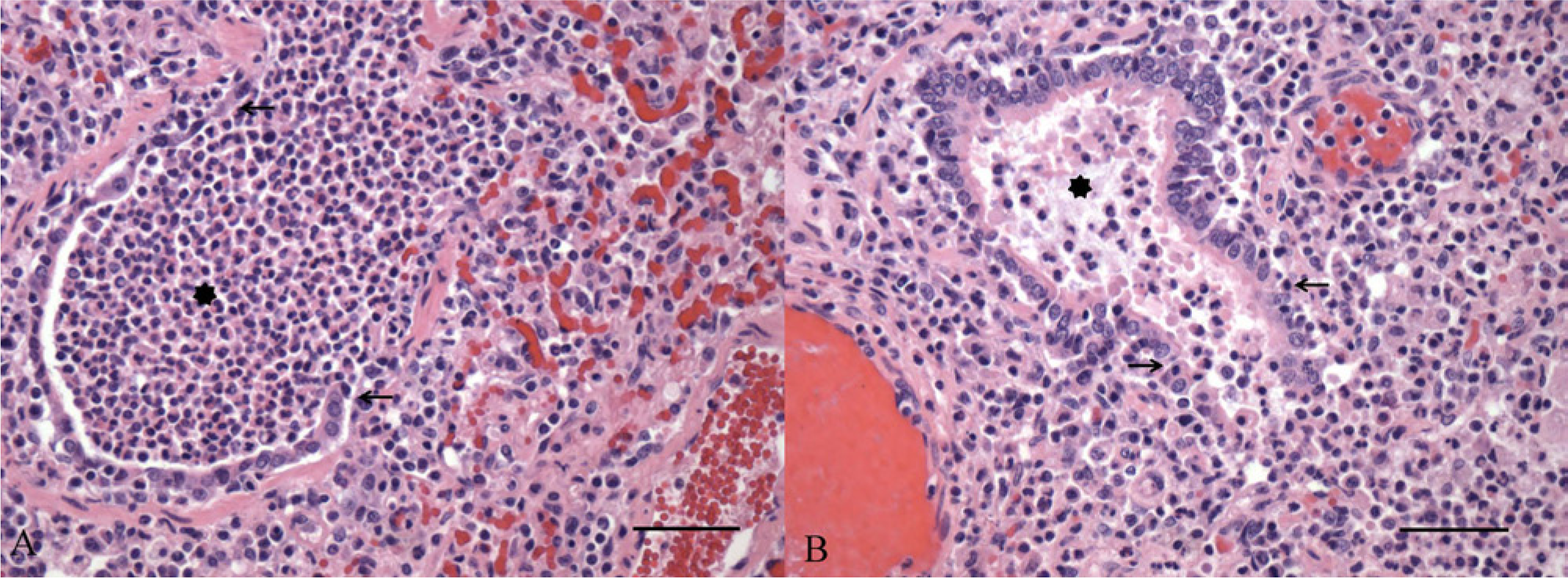
Lungs from piglet 1 (
Fresh lung and small intestinal contents from piglet 2 were submitted by the pathologist for differential diagnostic testing. Fresh lung was negative for Porcine reproductive and respiratory syndrome virus by polymerase chain reaction (PCR). A lung swab submitted for aerobic culture produced heavy growth of secondary pathogens common in swine including Pasteurella multocida and Klebsiella pneumoniae. Small intestine submitted for bacterial culture was negative for Salmonella spp. and Clostridium spp.
RNA was extracted from fresh lung homogenates from both piglets using a commercial viral isolation kit a according to the manufacturer’s recommendations. Samples were screened for the influenza matrix gene, according to U.S. Department of Agriculture (USDA) real-time reverse transcription PCR (real-time RT-PCR) protocols, as previously described. 19 Both samples were PCR positive for the matrix gene and also subsequently tested positive for influenza A(H1N1)pdm09, according to a USDA real-time RT-PCR protocol specific for the 2009 N1 influenza subtype. Lung homogenates from both piglets 1 and 2 were inoculated onto Madin–Darby canine kidney cells,b,20 as well as into 10-day-old specific pathogen–free embryonated chicken eggs. c Influenza A(H1N1)pdm09 was isolated by both methods.
The hemagglutinin (HA) gene segment was amplified using conventional PCR and purified from an agarose gel using a commercial gel extraction kit according to the manufacturer’s recommendations, d then sequenced using a commercial kit. a Sequence analysis showed a 98% homology with influenza A(H1N1)pdm09 human isolates from across the United States (including Nevada, North Carolina, Ohio, Oklahoma, South Carolina, Texas, California, New York, and Boston) and throughout the world (including Hong Kong, Canada, Cambodia, Thailand, England, Argentina, Athens, and Taipei). Sequence analysis also showed a 98% homology with 2 influenza A(H1N1)pdm09 swine isolates from 2009–2010: A/swine/Nebraska/A01049048/2010(H1N1) (GenBank accession no. JF812280.1) and A/swine/Minnesota/165A/2009(H1N1) (HQ840330).
Transmission of swine influenza from infected pigs to contact workers (i.e., veterinarians, swine farmers, and processing plant workers) has been widely documented throughout the United States since 1974, when swine influenza was first reported to have been isolated from an immunocompromised individual. 18 Human-to-pig transmission of influenza A(H1N1)pdm09 occurred during the 2009 pandemic period, 5 and typically involved contact with human beings exhibiting ILI.7,8
The current case represents possible human-to-swine transmission of influenza A(H1N1)pdm09 in a small, closed backyard herd. No workers other than the owner and immediate family were in contact with the pigs in this closed herd, and as a pharmacist, the owner was potentially occupationally exposed to influenza A(H1N1)pdm09. Although pharmacists have been shown to have higher than average vaccination rates for influenza, 17 the owner was not vaccinated. The owner and owner’s family did not have a history of ILI, and so may have been subclinically infected or served as fomites for transmission of the virus to the pig herd. Studies of influenza transmission dynamics in human beings have shown that the virus is shed maximally 1 day before clinical signs occur. 1 Subclinical infection with shedding is possible, 1 and during infection, particles containing influenza virus are exhaled in normal breath in the absence of coughing or sneezing. 4 These infection characteristics may have allowed this and many other human-to-swine transmission events of influenza A(H1N1)pdm09 to occur, and has implications for all swine workers, including owners of small herds, for recommendations of personal protective equipment to prevent transmission of influenza viruses to pigs. In 2011, several studies documented the occurrence of novel reassortment of influenza viruses in pigs,3,11 including cases involving human illness. 2 The current case highlights the potential ease with which even a small backyard swine herd—a group in which transmission of influenza virus from human beings to swine is not well understood—may be unknowingly infected, risking further virus mutation and potential reassortment with other influenza viruses.
Footnotes
Acknowledgements
The authors would like to thank referring veterinarians Drs. Michael J. Mathis and Lawrence R. Mackey for their work on this case.
a.
Ambion MagMAX-96 AI/ND Viral Isolation Kit, BigDye Terminator Kit; Life Technologies, Carlsbad, CA.
b.
American Type Culture Collection (ATCC), Manassas, VA.
c.
Sunrise Farms, Catskill, NY.
d.
QIAquick Gel Extraction Kit, Qiagen Inc., Valencia, CA.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Diagnostic testing was conducted under the USDA SIV surveillance program as a participating National Animal Health Laboratory Network laboratory.
