Abstract
Mink are susceptible to infection with influenza A virus (IAV) of swine and human origin. In 2019, a Utah mink farm had an outbreak of respiratory disease in kits caused by infection with the pandemic influenza A(H1N1)2009 virus [A(H1N1)pdm09]. In 3 wk, ~325, 1–2-wk-old kits died (10% mortality in kits). All deaths occurred in a single barn that housed 640 breeding females. No clinical signs or deaths occurred among adult mink. Five dead kits and 3 euthanized female mink were autopsied. All kits had moderate-to-severe neutrophilic and lymphohistiocytic interstitial pneumonia; adult mink had minimal-to-moderate lymphohistiocytic bronchointerstitial pneumonia. Immunohistochemistry and real-time PCR targeting the matrix gene detected IAV in lung of kits and adults. Virus isolation and genetic analysis identified the A(H1N1)pdm09 virus. The source of the virus was not determined but is thought to be the result of reverse zoonosis. Our case emphasizes the need for close monitoring on mink farms for interspecies transmission of IAV and for safe work practices on farms and in diagnostic laboratories. Additionally, a pandemic virus may continue to circulate at low levels long after the global event is declared over.
In April 2009, a novel strain of influenza A virus (IAV; Orthomyxoviridae, Alphainfluenzavirus) subtype H1N1 was reported in people in Mexico. The virus then spread to the United States, and by the following month had spread to 43 countries. 11 By June, more than 26,000 confirmed cases had been reported in 73 countries, and the World Health Organization (WHO) had classified the disease outbreak as a pandemic. 5 In the following year, the pandemic would result in 18,500 confirmed fatalities. 4 Estimates of total mortality during this time in 2 reports were 123,000–203,000 and 151,700–575,400 deaths.4,8 Unlike typical seasonal influenza, this particular strain of IAV caused most of the confirmed fatalities in individuals under the age of 65. 4 The total number of deaths was considered relatively small for an influenza pandemic, but many of those who died were under the age of 65 and the impact in terms of years of life lost was substantial. 5
The pandemic influenza A(H1N1)2009 virus [A(H1N1)pdm09] is thought to have originated in swine, and unfortunately was referred to widely as the “swine flu” early during the pandemic. In subsequent years, infection with this same virus has been documented in other species. Infection in domestic ferrets, mink, turkeys, cats, dogs, sloth bears, skunk, cheetahs, giant anteaters, a binturong, and a badger has been documented.6,7
Pneumonia in farmed mink caused by A(H1N1)pdm09 virus was first reported in 2011 in Canada. This facility housed approximately 1,200 breeding females, and the losses amounted to ~1,000 dead kits. Diagnosis of A(H1N1)pdm09 virus was confirmed using an IAV-specific reverse-transcription real-time (RT-rtPCR), followed by phylogenetic analysis. 1 An additional outbreak was identified in swine and a single mink sample in 2010 in the province of Quebec in Canada. This outbreak was caused by a novel IAV with genetic elements from the A(H1N1)pdm09 virus and a swine A(H3N2) virus. During that outbreak, affected swine had respiratory signs with no increase in death losses. The swine samples had hemagglutinin (HA) and neuraminidase (NA) genes that were closely related to a swine A(H3N2) virus; the matrix (M), polymerase (PB1, PB2, PA), nucleoprotein (NP), and nonstructural protein (NS) gene segments were closely related to A(H1N1)pdm09 virus. The mink sample from a single animal had an HA segment with 97.5% homology to the swine A(H3N2) virus, and an M gene with 99.8% homology to the A(H1N1)pdm09 virus. 10 Human-to-animal transmission is suspected in most reports of pandemic influenza infection in animals, although this is difficult to prove in farm outbreaks.
Here we report an outbreak of A(H1N1)pdm09 virus on a mink farm in Utah in May 2019. In early May 2019, a Utah mink farm with ~5,000 breeding females housed in 7 barns reported increased mortality in kits in a single barn. Each barn housed ~640 breeding females. The producer reported increased respiratory effort in some kits prior to death; others were found dead with no preceding clinical signs. All kits that died were <14-d-old. Some older kits were dyspneic, and then recovered. No adults in the barn displayed any clinical signs or died. Over the course of 3 wk, ~325 kits died, which resulted in an overall kit mortality of ~10%. After 3 wk, death losses subsided.
Five dead kits and 3 euthanized adults were submitted to the Central Utah Branch Laboratory of the Utah Veterinary Diagnostic Laboratory (Spanish Fork, UT, USA) for autopsy. Gross lesions were not observed in the kits or adults. Aerobic and Salmonella spp. bacterial cultures of lung from adults and kits yielded no growth. Microscopically, all of the kits had diffuse, moderate-to-severe, acute, neutrophilic, lymphocytic, and histiocytic interstitial pneumonia (Fig. 1A, 1B). All 3 adult mink had multifocal, mild-to-moderate, subacute lymphoplasmacytic and eosinophilic bronchointerstitial pneumonia (Fig. 1C). Adult mink had additional lesions that are common background lesions and considered incidental, including mild mononuclear hepatitis (1 of 3), minimal lymphoplasmacytic interstitial nephritis (2 of 3), mild eosinophilic mastitis (2 of 3), mild eosinophilic and suppurative enteritis (2 of 3), and mild lymphoplasmacytic, neutrophilic, and eosinophilic endometritis (2 of 3). Immunohistochemistry for IAV was performed on lung at the University of Minnesota Veterinary Diagnostic Laboratory (St. Paul, MN, USA). A moderate number of mononuclear cells in alveolar spaces and a few pneumocytes in the kit lung demonstrated strong cytoplasmic immunoreactivity for IAV (Fig. 1B). In the lung of adults, rare immunoreactive pneumocytes were observed (Fig. 1D). IAV rtPCR performed at the Utah Veterinary Diagnostic Laboratory detected the IAV M gene in lung samples from kits and adults. The avian IAV subtypes H5 and H7 were not detected by rtPCR. 9 Lung tissue from one of the kits was submitted to the National Veterinary Services Laboratories (NVSL; Ames, IA, USA), which confirmed IAV detection by rtPCR. IAV was isolated using MDCK cells, and whole-genome sequencing was performed for virus characterization. The virus was identified as pandemic lineage H1N1, with analysis of all 8 gene segments showing >99% homology to 2018–2019 available IAV sequences from humans (Suppl. Tables 1, 2).
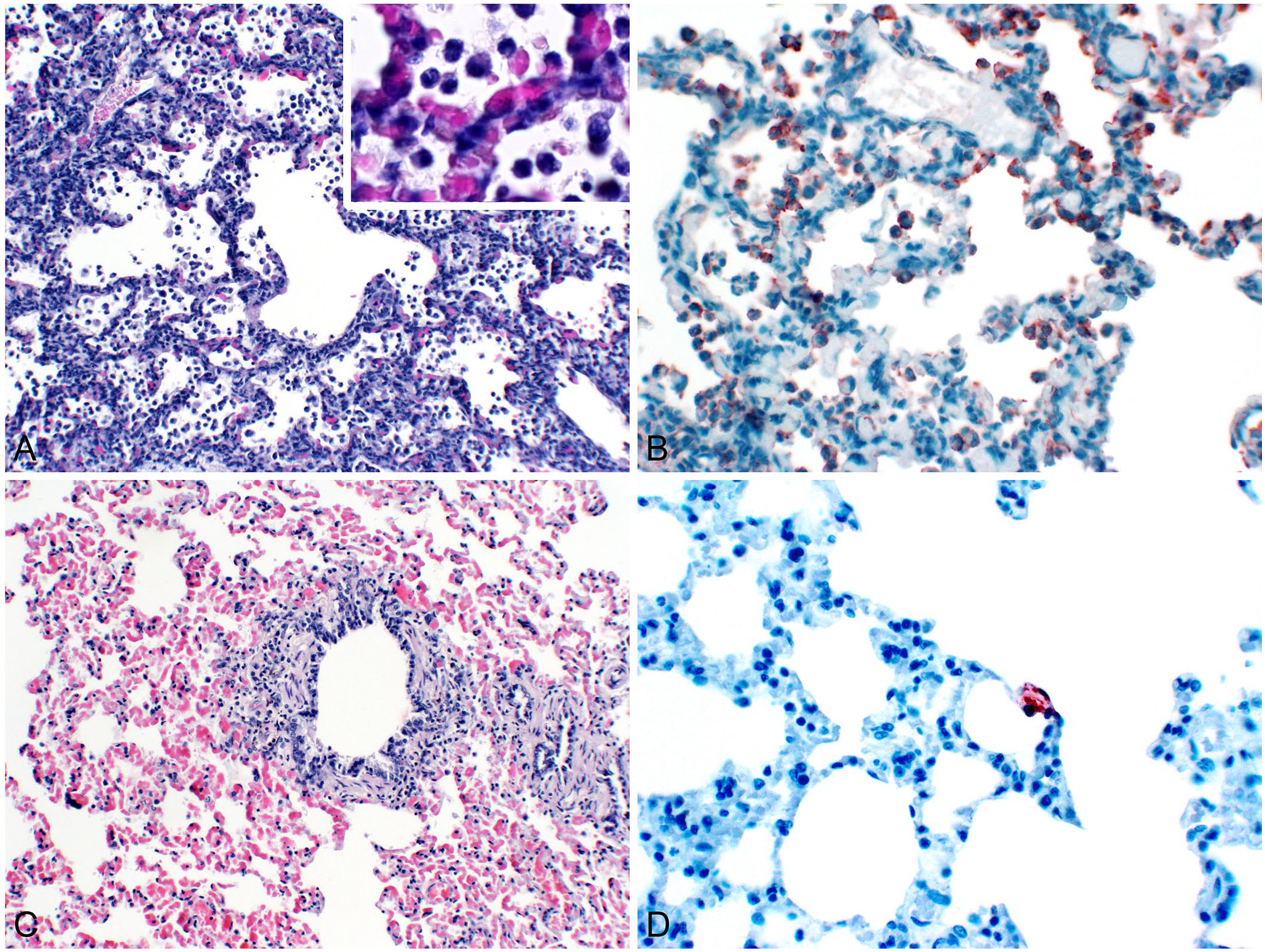
Interstitial pneumonia caused by influenza A virus (IAV) in a mink kit, and bronchointerstitial pneumonia caused by IAV in an adult mink.
Mink and other mustelids are susceptible to influenza viruses. 1 The interstitial pneumonia in kits suggested a viral infection, and IAV was detected by rtPCR, immunohistochemistry, and virus isolation. Death from the infection occurred solely in kits that were ≤14-d-old, which is consistent with a previous report of A(H1N1)pdm09 virus infection of mink in Norway. 1 This difference in susceptibility is most likely related to development of an adequate immune response. The producer reported that some kits that were >14-d-old exhibited mild respiratory signs and subsequently recovered. Canine distemper and Aleutian disease were the main differential diagnoses, but the clinical history and autopsy findings did not support infection with the respective viruses.2,12
The source of the IAV infection is speculated to be one of the farm employees who had respiratory signs at the time of the outbreak. The owner of the mink farm reported that one employee had a cough before the outbreak, but the employee did not seek medical care and a specific diagnosis was never made. Employees wore an unspecified type of respirator, and access to the affected barn was limited after the onset of respiratory signs to reduce the risk of zoonotic spread. An additional possible source of infection is through feed material. Beef byproduct and fishmeal were the main feed provided during breeding and parturition to avoid introduction of Salmonella sp. from spent hen feed material. This feed choice decreases the possibility of exposure to pathogens from poultry and swine in feed, which are the most likely species to be infected with IAV. Mink farms in the region use the same feed provider, and no other facilities reported an outbreak. Therefore, feed material is not the likely source of infection.
Pandemic IAVs have zoonotic and reverse zoonotic potential, and caution is warranted for agricultural workers and veterinarians. As in most published case reports of A(H1N1)pdm09 virus in animals, the source of infection was not determined, but is thought to be the result of zooanthroponosis. Food-borne transmission is possible but considered unlikely. Phylogenetic analysis of the virus suggests a human source in our case. Even after the pandemic ended, localized outbreaks continued to occur, and A(H1N1)pdm09 virus continues to circulate seasonally in the human population. 3 Domestic and wild animals can act as sentinel species for viruses and are a potential reservoir for IAV. Our case emphasizes the need for close monitoring for interspecies transmission of IAV and for safe work practices in farm and diagnostic laboratories. Prompt diagnosis is critical in cases where there is potential for significant spread of disease within the human population.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387211052966 – Supplemental material for Pandemic lineage 2009 H1N1 influenza A virus infection in farmed mink in Utah
Supplemental material, sj-pdf-1-vdi-10.1177_10406387211052966 for Pandemic lineage 2009 H1N1 influenza A virus infection in farmed mink in Utah by Michael J. Clayton, E. Jane Kelly, Marta Mainenti, Amanda Wilhelm, Mia Kim Torchetti, Mary Lea Killian and Arnaud J. Van Wettere in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared that they had no conflict of interests with respect to their authorship or the publication of this article.
Funding
The authors prepared this article as part of routine duties of employment and received no outside financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
