Abstract
Chlamydia spp., Coxiella burnetii, and Neospora caninum are responsible for reproductive diseases and are closely linked with high abortion rates in ruminants. Furthermore, C. burnetii and Chlamydia spp. have zoonotic potential. A real-time polymerase chain reaction (PCR) assay was developed for the simultaneous detection of Chlamydia spp.,C. burnetii, and N. caninum. The detection of beta-actin as internal control in the same PCR reaction provides additional information about sample quality by detecting the presence of PCR inhibitors. The multiplex real-time PCR developed in the current study shows a greater sensitivity compared to previously used single-target PCR reactions with a reproducible detection limit of 0.13 plasmid copies per PCR for each target. Additional parallel amplification of all detectable pathogens did not adversely impact sensitivity. This new multiplex PCR allows the highly sensitive, cost-effective, and rapid detection of 3 important pathogens and has the potential to be a useful time-saving tool in the routine diagnosis of abortion cases in ruminants.
Abortion in ruminants is a common economic problem worldwide,16,19 with Neospora caninum causing an estimated annual loss of $546.3 million in the U.S. dairy industry alone. 16 Three significant infectious agents causing abortion in ruminants are Chlamydia spp.,4,13 Coxiella burnetii, and Neospora caninum.1,12 In addition to impairing animal health, C. burnetii and several members of the genus Chlamydia are responsible for zoonotic diseases such as abortions in human beings.10,13 The enormous risk potential of C. burnetii being spread by goats was shown by the Dutch outbreak of 2007–2010, which caused more than 4,000 human infections. 6
Because these 3 pathogens cause similar symptoms, such as abortion, stillbirth, and the delivery of weak offspring,7,12 diagnostic investigations of ruminant abortions should include at least these pathogens. For rapid and specific diagnosis, direct detection by polymerase chain reaction (PCR) has been proven to be an important tool.5,11,14 To better use diagnostic resources, the aim of the current study was to develop a multiplex real-time PCR for the simultaneous and rapid detection of Chlamydia spp., C. burnetii, and N. caninum.
For the multiplex real-time PCR developed in the current study, previously described probe and primers for Chlamydia spp. 14 and N. caninum were used. 5 For C. burnetii, a previously described assay that targets the iso citrate dehydrogenase gene (icd) 11 was used. The newly developed multiplex real-time PCR was compared to the previously used single-target PCR protocols for Chlamydia spp., 14 C. burnetii, 8 and N. caninum. 2 To monitor amplification conditions and to detect PCR inhibitors, an internal control based on the housekeeping gene beta-actin 18 was integrated into the multiplex PCR.
DNA was extracted from the clinical specimen and isolates of bacteria (listed in Table 1) using a commercial DNA extraction kit a according to the manufacturer’s instructions. The multiplex PCR reaction was carried out in a 25-µl reaction volume. For a single reaction, the PCR mastermix contained 0.8 µl of each primer (10 pmol/µl working solution, final concentration of 320 nM), 0.4 µl of each probe (10 pmol/µl working solution, final concentration of 160 nM), 12.5 µl of PCR buffer containing DNA polymerase, b and 0.5 µl of RNase-free water. Finally, 4.0 µl of DNA template was added. The PCR was carried out in a commercial real-time PCR system c with an initial denaturation for 15 min at 95°C, followed by 45 cycles of 1 min at 94°C and 90 sec at 60°C.
Results showing the analytical specificity of the real-time multiplex polymerase chain reaction (PCR) assay by analyzing different targets.*
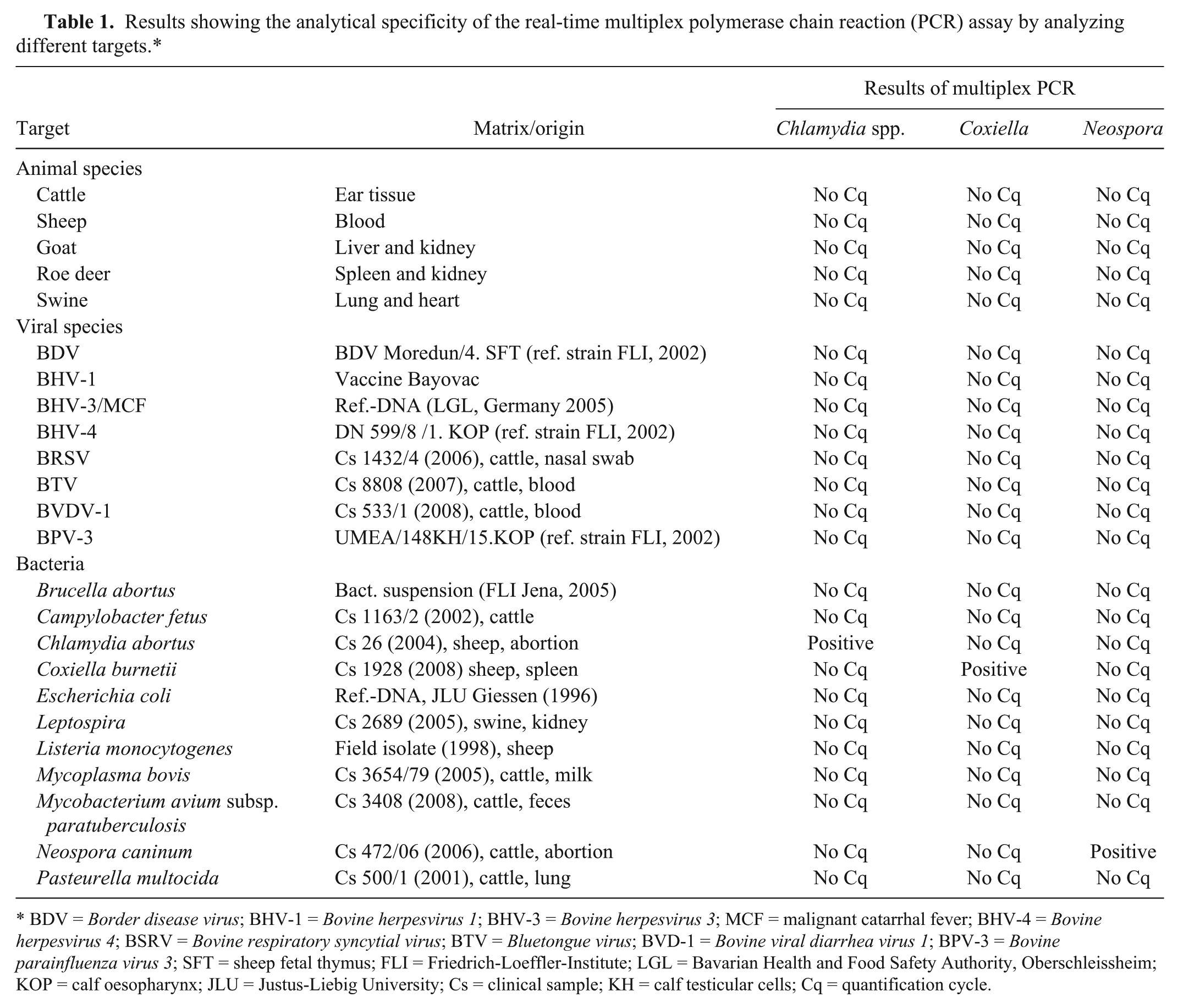
BDV = Border disease virus; BHV-1 = Bovine herpesvirus 1; BHV-3 = Bovine herpesvirus 3; MCF = malignant catarrhal fever; BHV-4 = Bovine herpesvirus 4; BSRV = Bovine respiratory syncytial virus; BTV = Bluetongue virus; BVD-1 = Bovine viral diarrhea virus 1; BPV-3 = Bovine parainfluenza virus 3; SFT = sheep fetal thymus; FLI = Friedrich-Loeffler-Institute; LGL = Bavarian Health and Food Safety Authority, Oberschleissheim; KOP = calf oesopharynx; JLU = Justus-Liebig University; Cs = clinical sample; KH = calf testicular cells; Cq = quantification cycle.
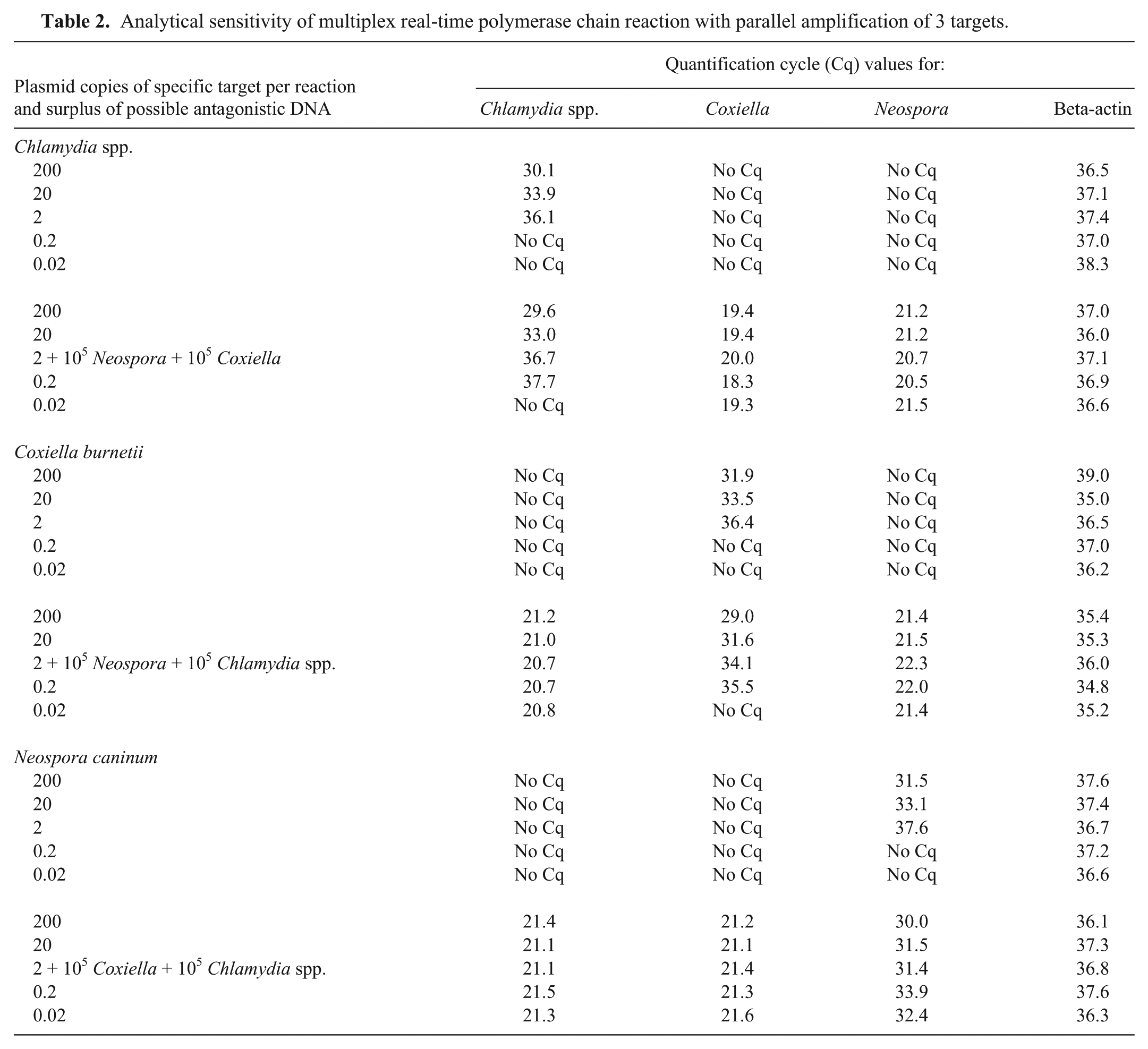
For analytical specificity, samples from different animal species and DNA of various viral and bacterial pathogens were examined. The samples showed no signs of nonspecific amplification (Table 1). Analytical sensitivity was determined by using a 10-fold dilution series of target-specific plasmid DNA d ranging from 109 to 100 copies per ml. Dilution series of each plasmid were examined 3-fold; mean values and standard deviation were calculated. 15 For each target, a strong correlation to the respective quantification cycle (Cq) value was found. The reproducible detection limit was 0.13 copies per reaction for each target (Fig. 1). To exclude the possible inhibition of a surplus of competitive DNA, serial dilution ranging from 0.02 to 200 plasmid copies of each target were analyzed with and without a fixed concentration of 105 plasmid copies of the respective competing agent. The addition of 105 copies of each competing target per reaction did not adversely impact the detection limits for the specific agent in any case. Therefore, inhibition of amplification due to a high surplus of certain targets can be excluded (Table 2). To compare the sensitivity between the multiplex real-time PCR and respective single-target PCR for Chlamydia spp., N. caninum, and C. burnetii, dilution series of 1 defined clinical reference sample for each target were examined in quadruplicate with multiplex real-time PCR and in triplicate with previous PCR methods. This comparison revealed identical and reproducible detection limits with the tendency for higher sensitivity of the multiplex assay (Table 3).
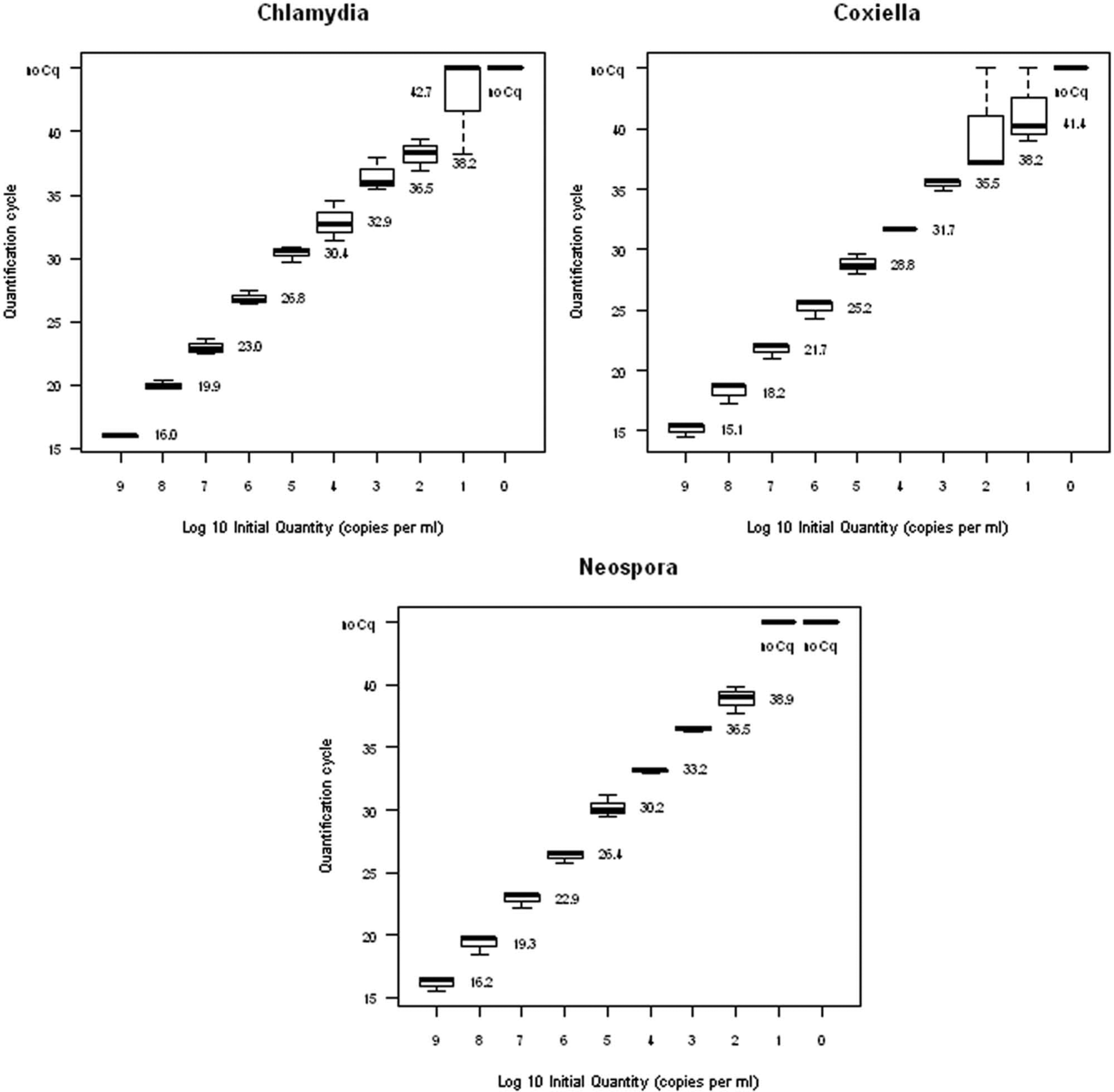
Analytical sensitivity of the assays included in the multiplex real-time polymerase chain reaction determined by dilution series of the respective synthetic plasmids. Mean quantification cycle (Cq) values of the replicates of each dilution step are depicted alongside each box plot. Boxplots were designed supported by the R software package. 15
Analytical sensitivity of multiplex real-time polymerase chain reaction with parallel amplification of 3 targets.
Quantification cycle (Cq) values and results for diluted clinical reference samples tested in quadruplicate with the multiplex real-time polymerase chain reaction (PCR) and in triplicate with a previously published PCR method.
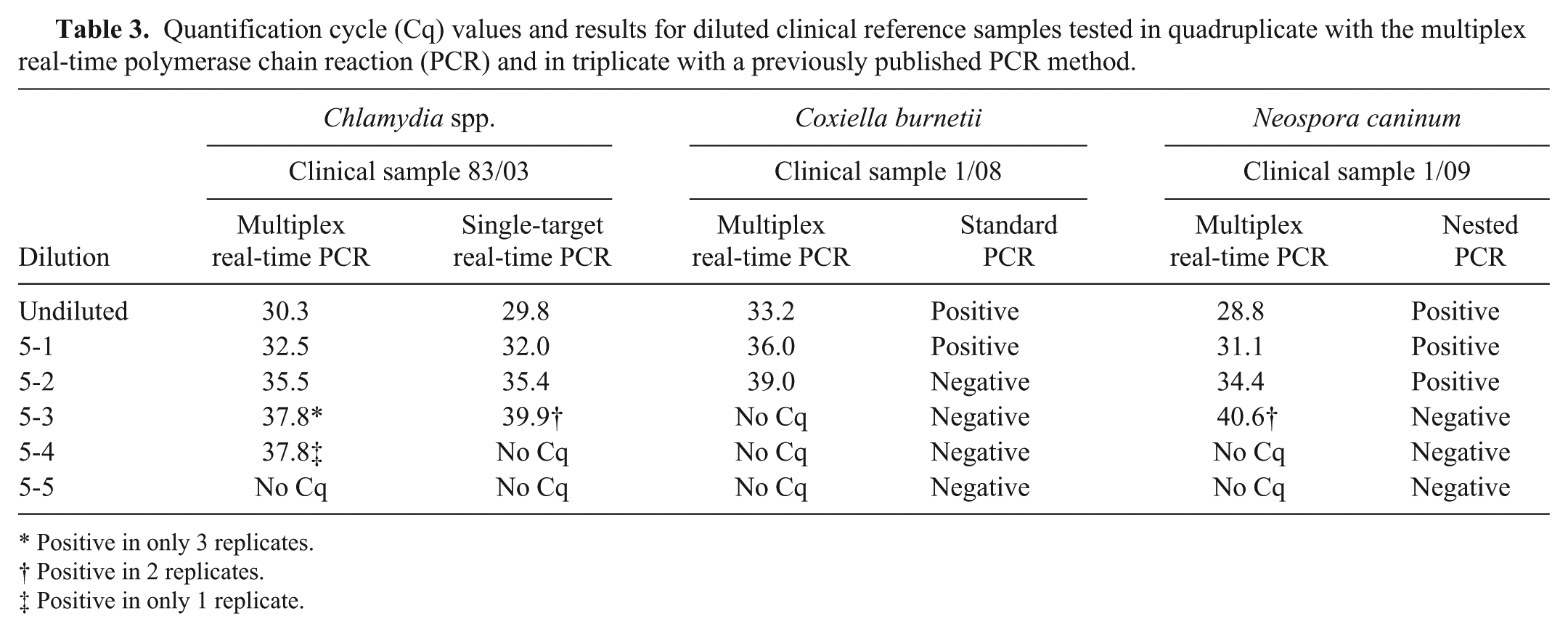
Positive in only 3 replicates.
Positive in 2 replicates.
Positive in only 1 replicate.
For diagnostic sensitivity and implementation, 91 samples positive for Chlamydia spp., obtained from Saxony-Anhalt (Germany), were analyzed (42 samples/abortion material; 49 samples/nonruminant or nongenital). The nongenital samples in the study included conjunctival and nasal swabs, bronchioalveolar lavage, feces, and tissue homogenates. Nonruminant samples originated from birds, horses, cats, and swine. Nineteen positive samples for C. burnetii were obtained from abortive material from the Bavarian Health and Food Safety Authority (Erlangen, Germany: 3 samples; Oberschleissheim, Germany: 16 samples). Thirty-eight samples positive for N. caninum were obtained from the Bavarian Health and Food Safety Authority (Oberschleissheim, Germany: 27 samples) and the Institute of Epidemiology (Friedrich-Loeffler-Institute, Wusterhausen, Germany: 11 samples). For the detection of Chlamydia spp., the multiplex real-time PCR was compared to the results obtained by single-target PCR. 14 Positive results were found with both methods for all 91 clinical samples (Fig. 2). Some of the Chlamydia spp.–positive samples have been further identified to species level by species-specific real-time PCR, sequencing, and/or microarray analysis 17 at the Institute of Molecular Pathogenesis of the Friedrich-Loeffler-Institute (Jena, Germany; data not shown). Chlamydia psittaci, Chlamydophila abortus, Chlamydophila caviae, Chlamydia pecorum, and Chlamydophila felis were identified. Nineteen C. burnetii–positive samples were analyzed with the multiplex real-time PCR in comparison to the single-target PCR. 8 All positive results were confirmed by the current multiplex real-time PCR (Fig. 3). Although the mean Cq value of the multiplex real-time PCR is increased by 6, the results of both assays are in strong linear correlation. All 38 analyzed N. caninum–positive field samples were confirmed by both multiplex real-time PCR and the previously used conventional semi-nested PCR. 2
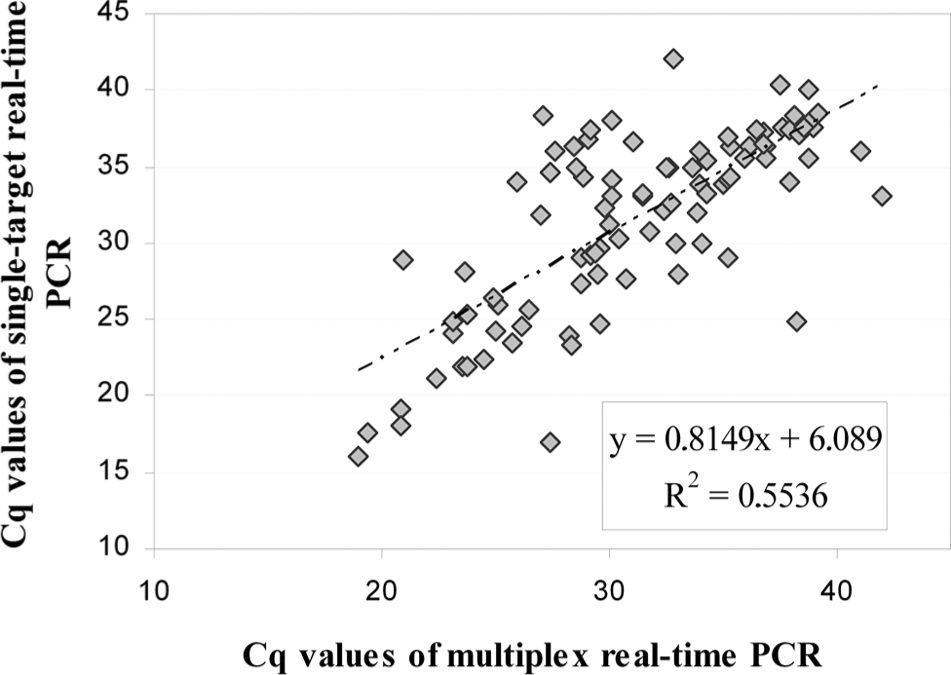
Correlation between the multiplex and single-target real-time polymerase chain reaction (PCR) detecting Chlamydia spp. for 91 positive samples obtained from various host species. Cq = quantification cycle.
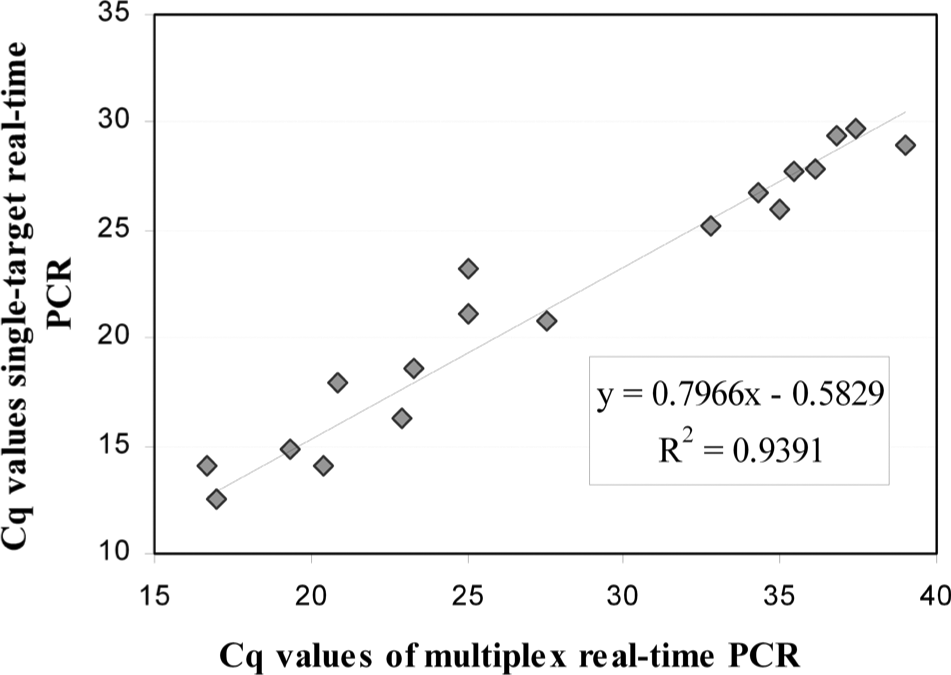
Correlation between the multiplex real-time polymerase chain reaction (PCR) and the single-target real-time PCR targeting the insertion sequence 111111 of Coxiella burnetii (results obtained by at the Bavarian Health and Food Safety Authority, Erlangen and Oberschleissheim, Germany) for 19 Coxiella burnetii–positive samples. Cq = quantification cycle.
The current study shows that the multiplex real-time PCR is able to simultaneously detect Chlamydia spp., C. burnetii, and N. caninum in abortive material of cattle and small ruminants as well as Chlamydia spp. in different sample types of ruminant and nonruminant species. Because of the diversity of Chlamydia spp. present in ruminants, the multiplex PCR was deliberately limited to screening for Chlamydia at the level of the genus.4,14 After positive detection of Chlamydia spp. DNA in clinical samples, further genotyping can be achieved by the application of species-specific real-time PCR or microarrays. 17 The application of only 1 fluorescence channel for Chlamydia spp. allows simultaneous detection of N. caninum as an additional, and important, pathogen, in contrast to conventional multiplex PCR assays previously described. 3
One common problem of multiplex PCR assays is the possible interaction of different oligonucleotides in the same reaction tube, leading to possible inhibition of the amplification reaction and subsequent reduced sensitivity of 1 or even all involved reactions. 9 In the current multiplex PCR, the sensitivity was not found to be influenced by the co-amplification of the 2 additional targets and achieved at least the same sensitivity as the respective single-target PCR when analyzing clinical samples. In conclusion, the current multiplex real-time PCR is an appropriate tool for the detection of Chlamydia spp., C. burnetii, and N. caninum in ruminants in a single reaction and may reduce the expenditures for biochemical reagents, equipment, and labor, resulting in a fast diagnosis for potential zoonotic infections.
Footnotes
Acknowledgements
The authors would like to thank Drs Juliana Drdlicek and Stephanie Christina Sörgel from the team of Dr Karl-Heinz Bogner (Bavarian Health and Food Safety Authority, Erlangen, Germany), Dr Gereon Schares (Institute for Epidemiology, Friedrich-Loeffler-Institute, Wusterhausen, Germany), and Dr Pia Zimmermann (Bavarian Health and Food Safety Authority, Oberschleissheim, Germany) for the delivery of DNA templates and their respective analytical results. The authors especially thank Dr Kerstin Wernike (Institute of Diagnostic Virology, Friedrich-Loeffler-Institute, Insel Riems, Germany) for critically reading the manuscript and preparing ![]() .
.
a.
High Pure PCR Template Preparation Kit, Roche Diagnostics GmbH, Mannheim, Germany.
b.
QuantiTect Multiplex PCR NoRox Kit, Qiagen GmbH, Hilden, Germany.
c.
Mx3005, Agilent Technologies, Santa Clara, CA.
d.
Gene Express, Berlin, Germany.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and publication of this article.
