Abstract
During 2003–2005, 399 abortion samples (315 fetuses and 84 placentae) were collected from 107 ovine and caprine farms in northern Sardinia. Tissues from aborted fetuses and placentae were examined by PCR assay to detect DNA from Coxiella burnetii, Chlamydophila abortus, Salmonella enterica Serovar abortusovis, Toxoplasma gondii, and Neospora caninum. The DNA from at least 1 of these 5 infectious agents was amplified in 41% of ovine fetuses, while only 17% of the caprine fetuses yielded a positive amplification result for at least 1 of the 5 agents. Out of a total of 366 ovine aborted samples, T. gondii DNA was detected most frequently (18.1% of fetuses and 13.1% of placentae), followed by S. abortusovis (13% of fetuses and 14.4% of placentae), C. burnetii (10.9% of fetuses, of 9.2% placentae), C. abortus (2.4% of fetuses, 6.5% of placentae), and N. caninum (2% of placentae). In 33 fetuses and 9 placentae, the simultaneous presence of pathogens with different associations was detected. Out of a total of 31 caprine aborted samples, T. gondii was detected most frequently (13% of fetuses and 25% of placentae), followed by C. abortus (12.5% of placentae), C. burnetii (12.5% of placentae), and N. caninum (8.6%).
Sardinia, an island located in the middle of the Mediterranean, has approximately 3 million milking Sarda sheep, corresponding to one third of the total Italian stock and about 0.2 million goats. The economic losses due to lamb mortality and missed lactation are estimated to be about 10 million euro/year.
In the authors' previous studies, the prevalence of abortion and the role of different infectious agents in abortion were investigated. The data indicated that Toxoplasma gondii plays an important role in ovine and caprine abortion. In fact, 11.1% and 6.4% of ovine and caprine fetuses, respectively, tested positive by PCR assay during 1999–2002. 11 Furthermore, other agents that play a relatively marginal role in abortion have been reported: Coxiella burnetii, 12 and Chlamydophila abortus. 13
The isolation of these pathogens from aborted samples (fetuses and placentae) represents the gold standard for definitive diagnosis. However, isolation requires obtaining samples in optimal conditions (they must be fresh, with little or no contamination, and free of toxic factors) that contain a threshold number of live and viable microorganisms. In fact, contamination with other bacteria, inadequate transport conditions, autolysis, and other factors may all adversely affect isolation. 14 DNA detection is more rapid than isolation and can be considered a useful technique for diagnosing the previously mentioned pathogens. 2,5,6,9 In the present study, PCR assay was applied in routine detection of C. burnetii, C. abortus, Salmonella enterica Serovar abortusovis, T. gondii, and Neospora caninum in fetal tissues from naturally aborted ewes and goats in 2003–2005.
A total of 315 fetuses (292 ovine and 23 caprine) and 84 placentae (including 76 ovine and 8 caprine samples) were analyzed during 2003–2005 (Table 1). Following macroscopic examination, the brain, skeletal muscle, liver, spleen, and abomasum were removed from each fetus. However, the condition of some of the fetuses was such that not all tissues could be collected. From the placental samples, homogenates were prepared using the cotyledon and intercotyledon regions. Fetal tissues and placental samples were then washed with phosphate buffered saline (PBS) containing 1,000 U/ml of penicillin and 1,000 U/ml of streptomycin sulphate and then digested with 2% trypsin for 3 hours at 37°C. When the fetus was too small, it was digested in toto. The brain was digested using a 0.6% trypsin concentration. Digested tissues were filtered through sterile gauze and centrifuged at 3,000Xg for 10 minutes. After 3 washes, pellets were resuspended in Feral Bovine Serum (FBS) containing 10% dimethylsulphoxide, aliquoted, and stored at −20°C until use.
Positive control genomic DNA was extracted from C. burnetii Nine mile/I/EP1 (ATCC VR-615), C. abortus S26/3, T. gondii (ATCC 40050), N. caninum (BPA1, ATCC 75710), and S. abortusovis. The first 4 micro-organisms were propagated in specific cell cultures at 37°Cwith5% CO2 while S. abortusovis SS44, an isolate from sheep originating from Sardinia, was grown in Luria-Bertani broth a at 37°C. 3 DNA was extracted with phenol-chloroform-isoamyl alcohol, purified following a previously described procedure, 1 and used as positive control in PCR amplification. DNA concentration was determined by ultraviolet (UV) absorption.
DNA was extracted from the digested tissue samples of brain, skeletal muscle, liver, spleen, abomasum, and placenta using the DNeasy Tissue Kit. b The primers used in this work, the expected product sizes, and corresponding bibliographic references are listed in Table 2. All PCR reactions were performed in a GeneAmp PCR System 9700. c Positive control DNA and water samples were included in all amplifications. PCR products were resolved on a 1–1.5% agarose gel in 1 × TAE buffer (0.04 M Tris-acetate, 0.001 M EDTA). After electrophoresis at 100 volts for 60 minutes, gels were stained with ethidium bromide and examined over UV light in an ImageMaster VDS-CL System. d
Total number of fetuses and placentae collected during 2003–2005 in 107 farms.
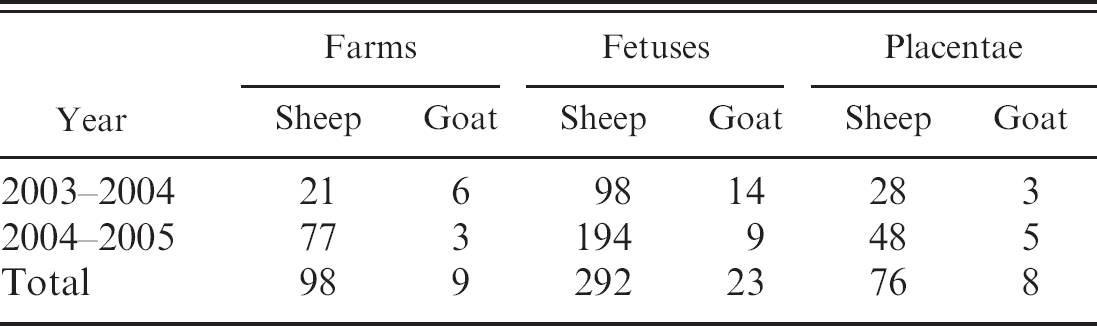
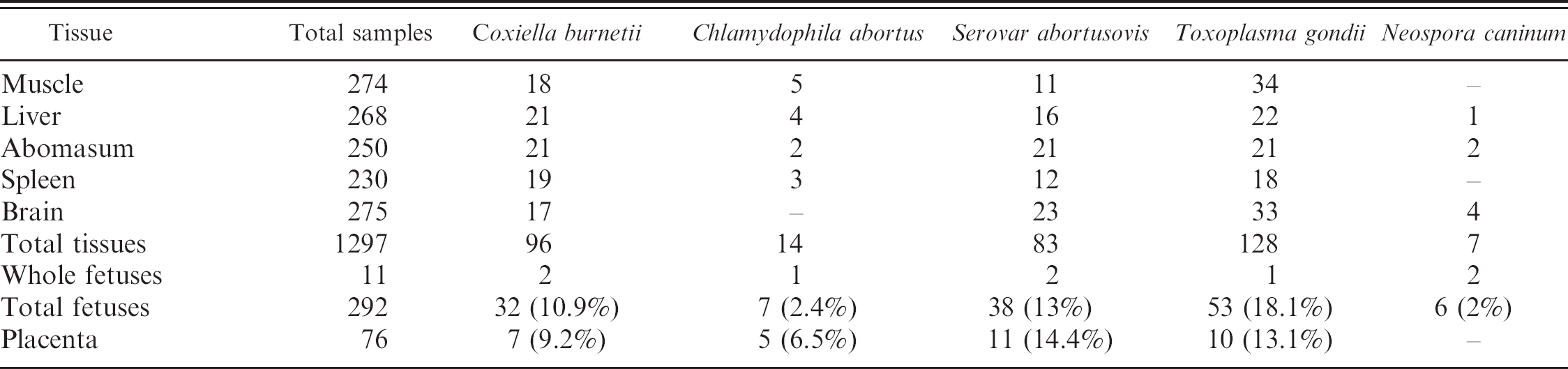
During 2003–2005, 107 (98 ovine and 9 caprine) farms located in northern Sardinia submitted at least 1 abortion specimen to the authors' laboratories for diagnostic testing (Table 1). Table 3 shows the results obtained from tissues, fetuses, and placentae analyzed by PCR assay utilizing the primer sets listed in Table 2. Out of a total of 292 ovine fetuses, we found 53 fetuses to be T. gondii PCR positive, corresponding to 18.1%, followed by S. abortusovis with 38/292, corresponding to 13%;by C. burnetii with 32/292 (10.9%); by C. abortus with 7/292 (2.4%); and finally by N. caninum with 6/292 (2%).
Out of a total of 23 caprine fetuses, 3 (13%) tested positive for T. gondii and 2 (8.6%), for N. caninum. We did not find any amplification band against the other abortifacients. Regarding the caprine placentae, out of a total of 8 examined samples, 2 (25%) tested positive for T. gondii, and 1 (12.5%), for C. burnetii and C. abortus. DNA of S. abortusovis and N. caninum was not detected in caprine placenta (data not shown). Table 4 shows the percentage of fetuses and placentae positive for single or multiple infectious agents.
Of a total of 33 ovine fetuses and placentae, we detected the DNA of different abortifacients concurrently, with the following combinations: 11/33 fetuses were S. abortusovis and C. burnetii PCR positive; 5/33, S. abortusovis plus T. gondii; 4/33, C. burnetti plus T. gondii; 3/33, T. gondii plus N. caninum; 1/33, S. abortusovis plus N. caninum;1, C. burnetii plus C. abortus;1. C. burnetii and N. caninum; and at last 1, C. abortus and T. gondii. Some fetuses were simultaneously positive for 3 aborting agents: 2 for S. abortusovis-C. burnetii-C. abortus, 2 for S. abortusovis-T. gondii-C. burnetii, and 1 for S. abortusovis-C. burnetti-N. caninum. In 1 fetus, we detected the DNAs of all agents except that of N. caninum. Regarding the 9 positive placentae, in 4, we detected the DNA of S. abortusovis and C. burnetii;in 2, C. abortus and T. gondii; in 1, C. burnetii and C. abortus; in 1, C. burnetii and T. gondii; and last in 1, T. gondii and N. caninum. In a single caprine fetus, we found the simultaneous presence of T. gondii and N. caninum DNA (data not shown).
Primer sequences used in PCR amplification.
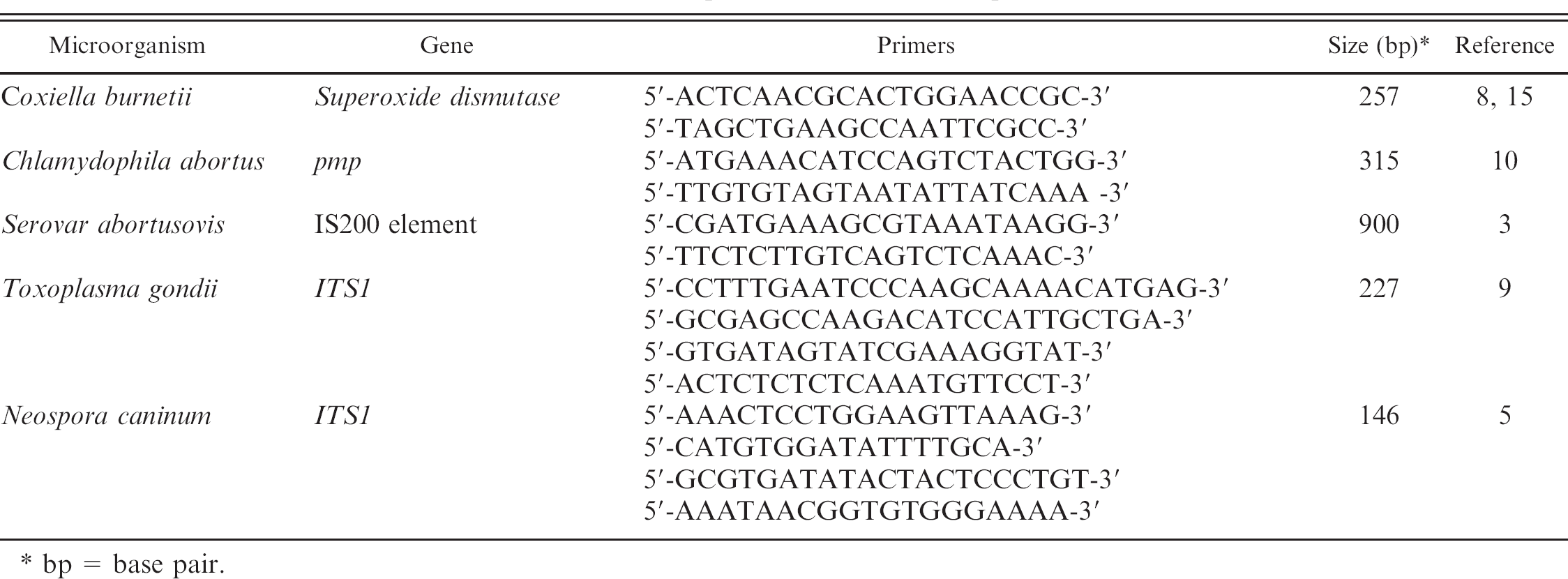

bp = base pair.
Positive PCR results obtained from total ovine fetuses, placentae, and tissues during 2003–2005.

Comparison of percentage of positive fetuses and placentae by PCR assay against 1 or more infectious agents. *
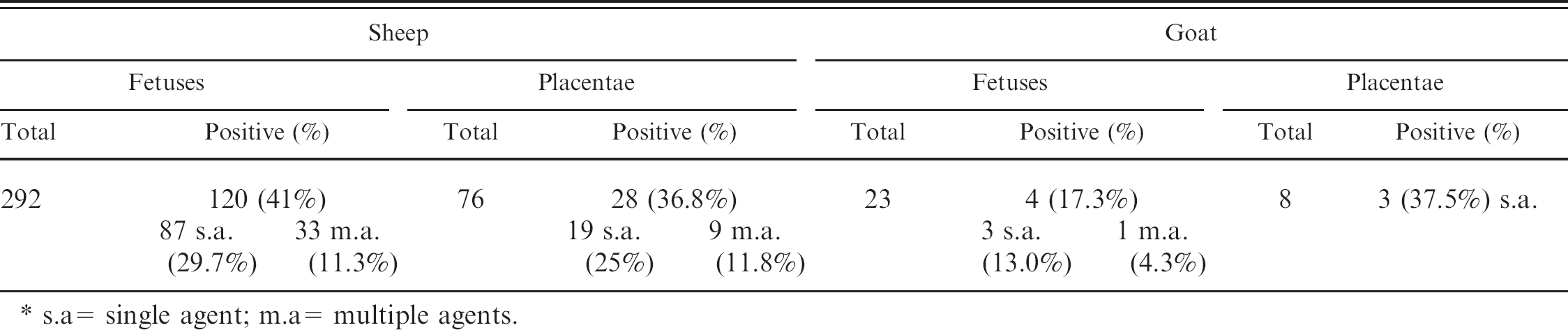

s.a= single agent; m.a= multiple agents.
This study is one of the most thorough of its kind reported in the literature, considering the number of fetuses and placentae tested for the presence of DNA from C. burnetii, C. abortus, S. abortusovis, T. gondii, and N. caninum in ovine and caprine abortion products. The results indicate that T. gondii DNA was the most frequently detected in ovine fetuses (18.1%), and S. abortusovis DNA was most frequently detected in placentae (14.4%) (Table 3). On the other hand, in goats, T. gondii was detected in 13% of fetuses (data not shown). These data are further confirmation of our previous results from Sardinia. 11–13
The first report of N. caninum related to abortion in naturally infected sheep was published in 2003, 7 and studies regarding the prevalence of this protozoa in sheep are few. In this study, N. caninum DNA was detected in only 2% of ovine fetuses and 8% of caprine fetuses. Moreover, 33 fetuses (11.3%) and 9 ovine placentae (11.8%) yielded positive PCR results for 2 or more infectious agents; the simultaneous detection of S. abortusovis and C. burnetii was the most prevalent. Brodie et al. 4 described the simultaneous detection of T. gondii and C. abortus in an abortion outbreak in a Scottish sheep flock, while Szeredi and Bacsadi 16 observed simultaneous infection with C. abortus and T. gondii in 4 ovine placentae, even though they caused lesions in separate locations of the cotyledons. Hässig et al. 7 described the presence of both of toxoplasmosis and neosporosis problems in the same flock. They also detected simultaneous infections with N. caninum and C. psittaci in 3 fetuses.
In conclusion, 41% (120 out of 292) ovine fetuses tested positive for T. gondii, S. abortusovis, C. burnetii, C. abortus, or N. caninum; the remaining 59% were not positive for any of the 5 agents tested. In caprines, the percentage of abortion samples yielding no positive results for any of the 5 agents was 82.7%. Further studies are in progress to determine the possible role of other bacterial and viral agents in ovine and caprine abortion.
Footnotes
a.
Difco, Sparks, MD.
b.
Qiagen, Valencia, CA.
c.
Applied Biosystems, Foster City, CA.
d.
Amersham Biosciences Europe GmbH, Milano, Italy.
