Abstract
Enterococci are major zoonotic bacteria that cause opportunistic infections in human beings and animals. Moreover, pathogenic strains can be disseminated between human beings and animals, particularly companion animals that come into frequent contact with people. Recently, Enterococcus faecium clonal complex 17 (CC17) has emerged as a pandemic clone. Most CC17 strains are ampicillin resistant and possess virulence genes such as esp and hyl. Despite the possible dissemination of CC17 between human beings and animals, prevalence data about CC17 in animals is limited. In the present study, the phenotypes and genotypes of antimicrobial resistance were compared, as well as virulence gene profiles from 184 enterococci strains isolated from chickens, pigs, companion animals, and human patients in Korea. Ampicillin-resistant E. faecium (AREF) strains were selected, and multilocus sequence typing was performed to investigate the dispersion of CC17 among animals and human beings. The companion animal and human isolates showed high resistance rates to ampicillin and ciprofloxacin, whereas food animal isolates showed high tetracycline and erythromycin resistance rates. Ampicillin-resistant E. faecium was only detected in human (21/21 E. faecium, 100%) and companion animal (3/5 E. faecium, 60%) isolates, and all human AREF strains and 1 canine AREF strain were confirmed as CC17. In conclusion, the occurrence of antimicrobial resistance and virulence genes, and the distribution of enterococcal CC17 in companion animal enterococcal strains were similar to those of human strains rather than to those of food animal strains.
Introduction
Enterococci are one of the major zoonotic bacteria that colonize the skin, mucosal membranes, and gastrointestinal tract of human beings and animals. 33 Furthermore, these bacteria cause a wide range of nosocomial infections including bacteremia, peritonitis, endocarditis, and device-related infections in human beings. 32 Most infections are opportunistic, and the high rates of antimicrobial resistance make it difficult to treat the diseases. 2 Therefore, concerns about public health issues evoked by exchanging antimicrobial-resistant and virulent enterococci between animals and human beings have increased.6,9
Enterococci can easily acquire antimicrobial resistance, which allows them to play a role as an antimicrobial resistance indicator. 32 Intrinsic resistance (e.g., against cephalosporins, β-lactams, sulfonamides, and low-level aminoglycosides) and acquired resistance are both problems when attempting to treat enterococcal infections. For example, vancomycin-resistant enterococci (VRE) have threatened human health, as vancomycin is a common therapeutic used for severe enterococcal infections. 3 Furthermore, high-level aminoglycoside-resistant enterococci, such as high-level gentamicin-resistant (HLGR; minimum inhibitory concentration [MIC] >500 µg/ml) and high-level streptomycin-resistant (HLSR; MIC >2,000 µg/ml) enterococci, have become a problem in human health. High-level aminoglycoside-resistant enterococci narrow the choice of therapeutics for treatment, because aminoglycosides and β-lactams are generally used for treating serious enterococcal infections. 40
Enterococcus faecium clonal complex 17 (CC17), one of the global epidemic clones of ampicillin-resistant E. faecium (AREF) receiving attention as a hospital-adapted strain, has ampicillin and ciprofloxacin resistance and can acquire vancomycin resistance genes and other virulence genes.35,39 Because the strong linkage between AREF and CC17 has been widely accepted, 20 monitoring AREF is as important as detecting CC17.
Virulence factors contribute to colonization and infection by enterococci. Several virulence factors have been reported, including aggregation substance, gelatinase, cytolysin, enterococcal surface protein (Esp), collagen-binding protein, and endocarditis antigen protein of E. faecalis and Esp and hyaluronidase (Hyl) of E. faecium.16,23 Expression of these factors is related to attachment, biofilm formation, invasion into the host, evasion from killing by neutrophils, and secretion of toxins that damage host defense systems.27,28,31 In particular, most CC17 possess esp and hyl genes, which provide advantages in the adaptation of enterococci to the hospital environment. 20
Antimicrobial-resistant and virulent enterococci can be transferred between human beings and animals via consumption of food animal products or through direct contact with companion animals.7,14,18 Despite their importance, few studies have been conducted to screen the distribution of CC17 in animals and compare them with human isolates. In the present study, the phenotypes and genotypes of antimicrobial resistance as well as virulence gene profiles were investigated in enterococcal strains isolated from chickens, pigs, companion animals, and human patients in Korea. The AREF strains were selected, and multilocus sequence typing (MLST) was performed to investigate the dispersion of CC17 among animals and human beings.
Materials and methods
Bacterial isolation and identification
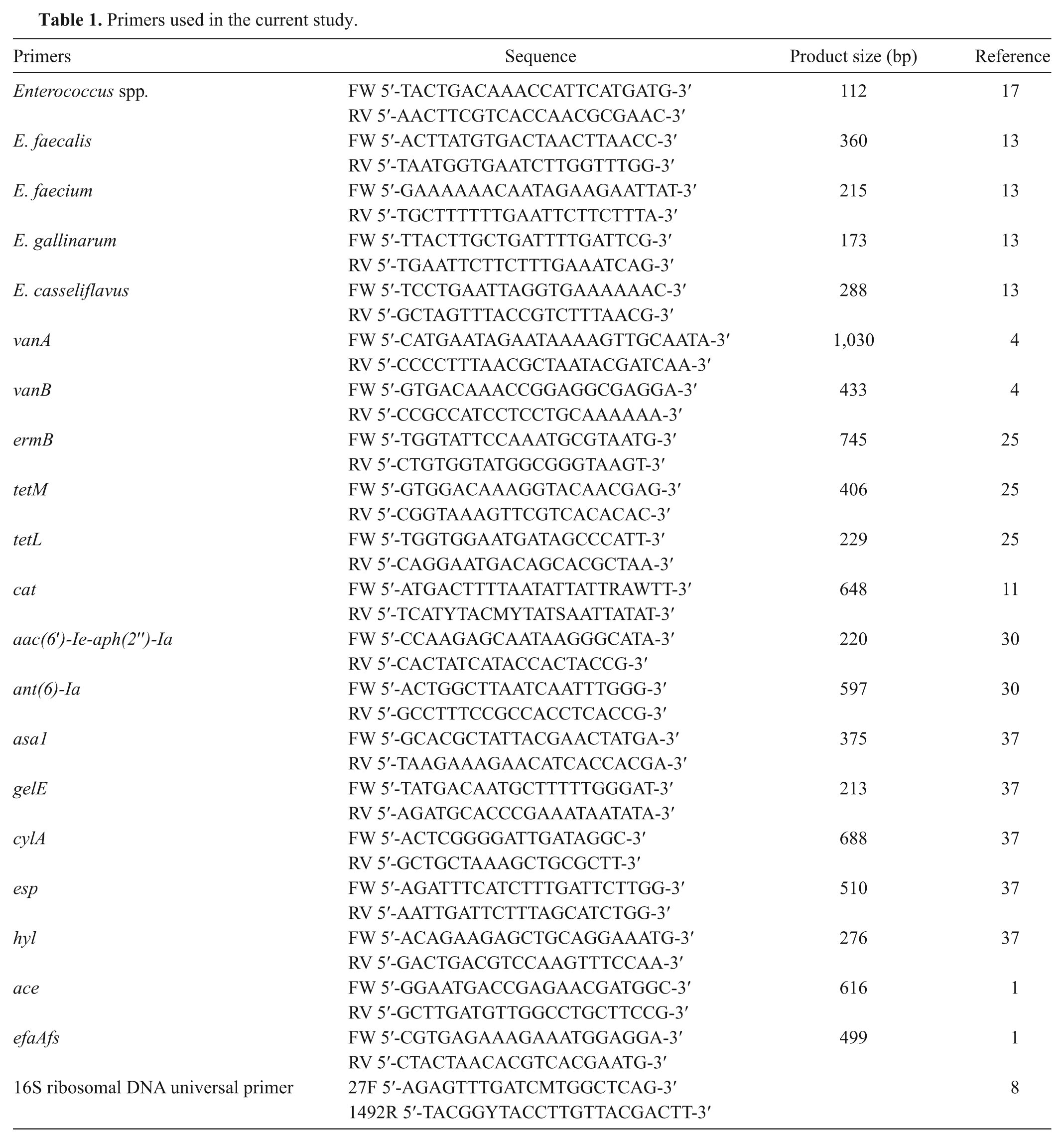
In total, 184 enterococcal isolates were collected. Forty-nine isolates were collected from chicken carcasses, and 52 isolates were collected from slaughtered pigs. Companion animal–originating isolates (44) were collected from swab samples of the skin and rectum of 43 dogs and 1 cat that visited 6 different veterinary clinics due to non-enterococci–associated diseases. Human isolates were provided by the Asian Bacterial Bank of the Asia Pacific Foundation for Infectious Diseases. The 39 human isolates were recovered from blood, peritoneal fluid, bile, and dialysate samples of enterococci-infected patients. All samples were collected between 2008 and 2010. All isolates were confirmed as enterococci by polymerase chain reaction (PCR) and identified as E. faecalis or E. faecium by species-specific PCR or other enterococcal species by 16S ribosomal RNA sequencing.8,13,17 The primers used in the current study are shown in Table 1.1,4,8,11,13,17,25,30,37
Primers used in the current study.
Antimicrobial susceptibility tests
Antimicrobial susceptibility tests were performed for 6 antibiotics (vancomycin, erythromycin, tetracycline, chloramphenicol, ampicillin, and ciprofloxacin) by the disk diffusion method. 5 The agar dilution method was performed according to the Clinical and Laboratory Standards Institute guideline to determine HLGR and HLSR. 5 Enterococcus faecalis ATCC (American Type Culture Collection) 29212 and E. faecalis ATCC 51299 were used as reference strains. A multidrug-resistant (MDR) isolate was defined as an isolate that was resistant to a minimum of 3 antibiotic categories. 24
Detecting antimicrobial resistance and virulence genes
The 7 antimicrobial resistance–associated genes and 7 virulence-associated genes were investigated by PCR using primers (Table 1) from previous studies,1,4,11,25,30,37 including vancomycin resistance (vanA and vanB), erythromycin resistance (ermB), tetracycline resistance (tetM and tetL), chloramphenicol resistance (cat), HLGR (aac(6′)-Ie-aph(2′′)-Ia) and HLSR (ant(6)-Ia), aggregation substance (asa1), gelatinase (gelE), cytolysin (cylA), esp, hyl, collagen-binding protein (ace), and E. faecalis endocarditis antigen (efaA fs ).
Multilocus sequence typing
The 24 AREF isolates detected in the current study (3 isolates from companion animals and 21 isolates from human beings) were subjected to MLST as previously described. 10 Briefly, 7 housekeeping genes (atpA, ddl, gdh, purK, gyd, pstS, and adk) were amplified and sequenced. Allele analysis and sequence typing were performed at the relevant website (http://www.mlst.net).
Statistical analysis
Statistical analysis was conducted with the chi-square test using the commercial software. a A P value < 0.05 was considered significant.
Results
Enterococcus spp. detected from animals and human beings
The enterococcal species identified are summarized in Table 2. Enterococcus faecalis and E. faecium were detected in all samples. Other than these 2 major species, the other 6 minor species (E. hirae, E. gilvus, E. avium, E. canintestini, E. saccharolyticus, and E. pseudoavium) were detected in companion animals (4 species only), pigs (3 species only), and human beings (1 species only).
Enterococcus spp. isolated from chickens, pigs, companion animals, and hospitalized human beings.*
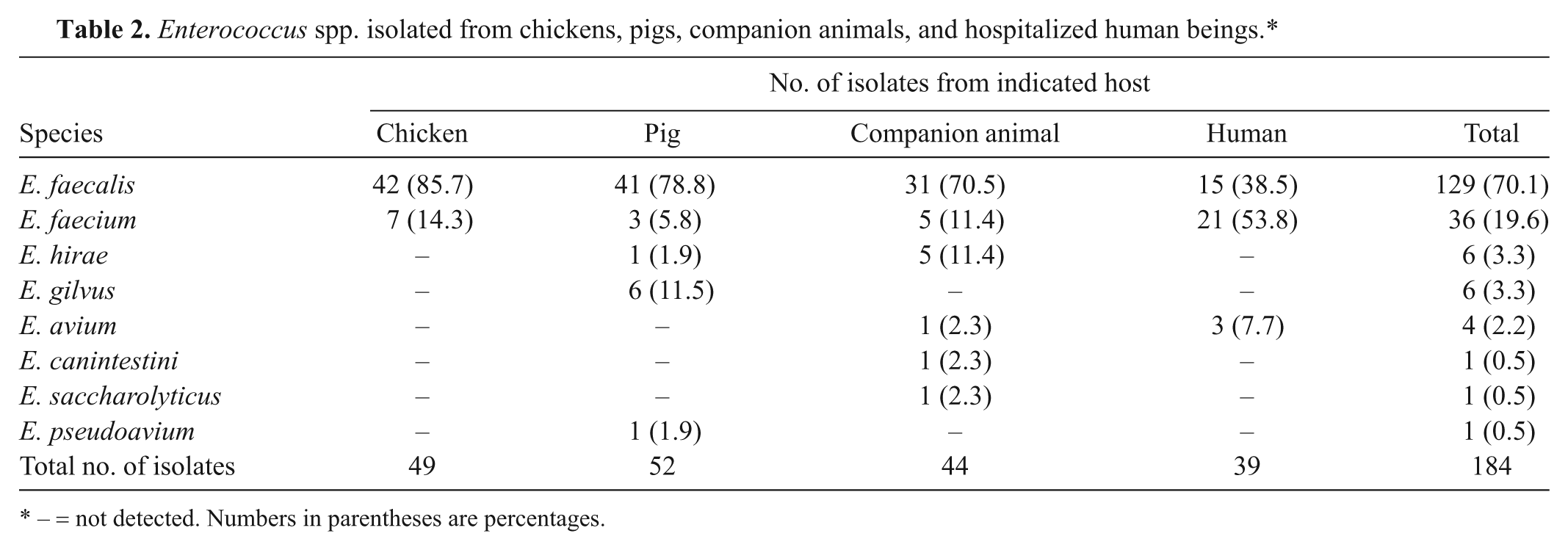
– = not detected. Numbers in parentheses are percentages.
Antimicrobial resistance and associated genes
The phenotypes and genotypes of the antimicrobial-resistant enterococcal isolates are shown in Tables 3 and 4. Vancomycin resistance only occurred in 6 human E. faecium isolates (15.4% among all the human isolates), and the vanA gene was detected in 1 of these isolates. No vancomycin-resistant or vanA gene–possessing strains were isolated from animals. The vanB gene was not detected.
Antimicrobial resistance rates of the enterococcal isolates.*
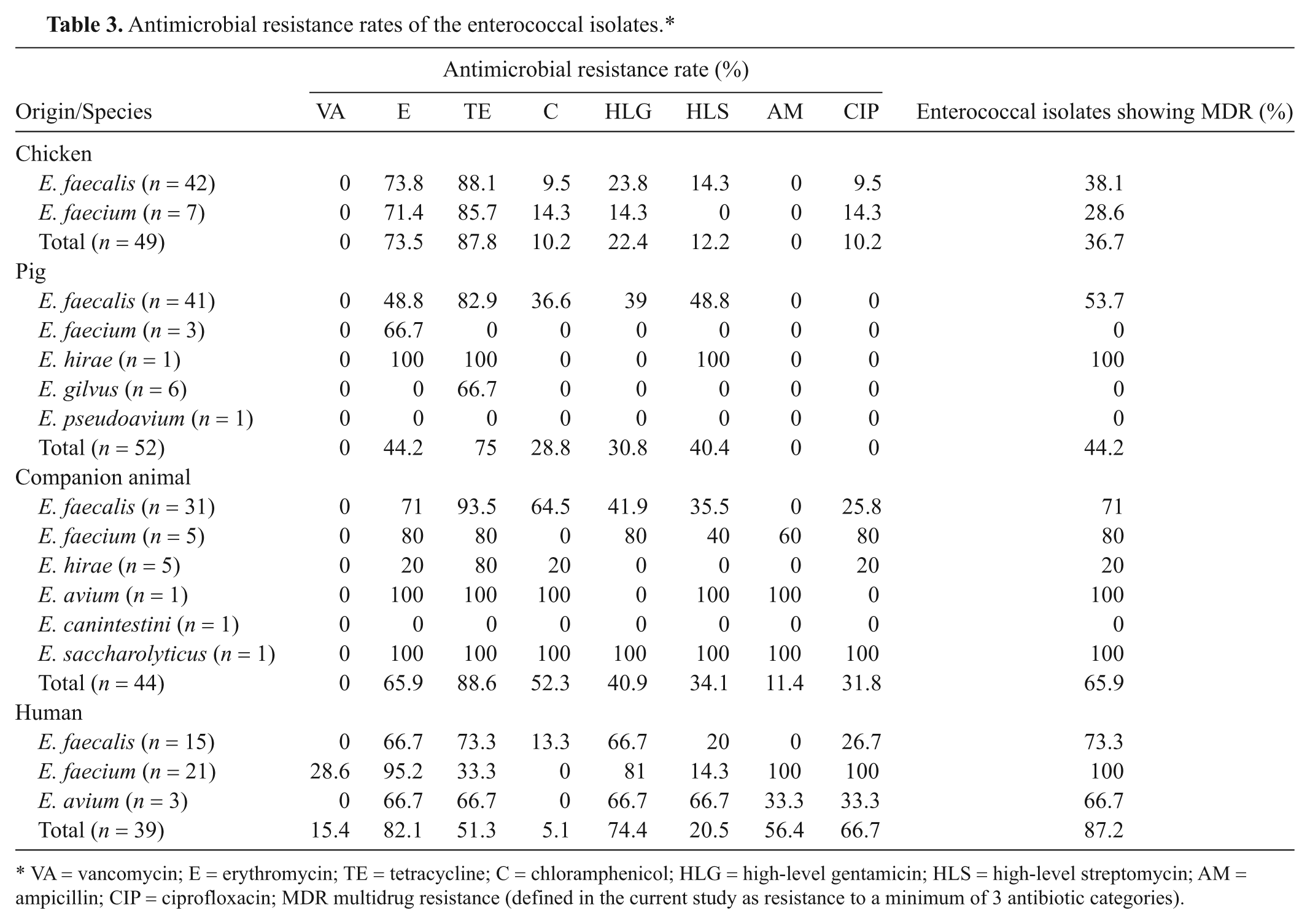
VA = vancomycin; E = erythromycin; TE = tetracycline; C = chloramphenicol; HLG = high-level gentamicin; HLS = high-level streptomycin; AM = ampicillin; CIP = ciprofloxacin; MDR multidrug resistance (defined in the current study as resistance to a minimum of 3 antibiotic categories).
Detection of antimicrobial resistance genes among enterococcal isolates.
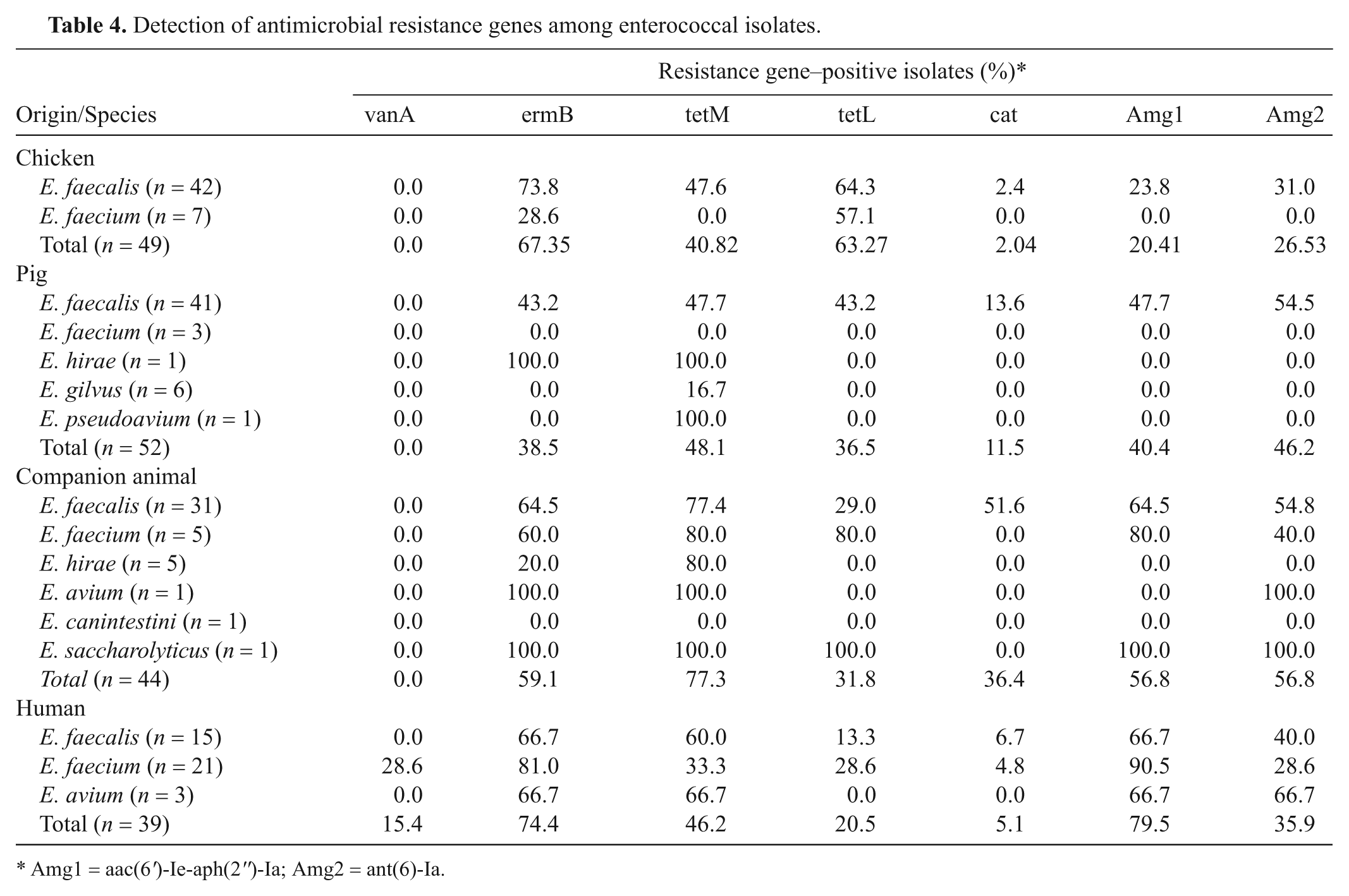
Amg1 = aac(6′)-Ie-aph(2′′)-Ia; Amg2 = ant(6)-Ia.
Erythromycin and tetracycline resistance detection rates were generally high among all sources. The chloramphenicol resistance rate was higher in companion animals than that from the other sources (P < 0.05). The HLGR rates of E. faecium from companion animal and human strains were higher than that from food animals (P < 0.05).
The ampicillin resistance rate was 56.4% in human beings (22/39 isolates) and 11.4% in companion animals (5/44 isolates), whereas it was not detected in chicken or pig isolates. Among E. faecium isolates, 24 AREF isolates were detected in companion animals (3/5 isolates, 60%) and human beings (21/21 isolates, 100%). Ciprofloxacin resistance was detected in human isolates (66.7%), companion animal isolates (31.8%), and chicken isolates (10.2%), but not from pig isolates. Interestingly, all of the AREF isolates from companion animals and human beings were resistant to ciprofloxacin.
Multidrug-resistant enterococci
The enterococci MDR rates are summarized in Table 3. The MDR rates for the E. faecalis and E. faecium strains in companion animal (71.0% and 80.0%, respectively) and human (73.3% and 100%, respectively) isolates were higher than those in chicken (38.1% and 28.6%, respectively) and pig (53.7% and 0%, respectively) isolates (P < 0.05). In agreement with the MDR phenotype results, the number of isolates possessing multiple antimicrobial resistance genes was higher in companion animals and human beings than that in the chicken and pig (P < 0.05, data not shown). The most prevalent E. faecalis antimicrobial resistance gene patterns from companion animals and human beings were ermB/tetM/aac(6′)-Ie-aph(2′′)-Ia/ant(6)-Ia/cat (25.8%) and ermB/tetM/aac(6′)-Ie-aph(2′′)-Ia/ant(6)-Ia (33.3%). These 2 patterns were similar except for the cat gene. The sole E. saccharolyticus isolate, obtained from a companion animal, was resistant to all the antibiotics tested except vancomycin and also had 5 antimicrobial resistance genes.
Detection of virulence genes
The detection rates of virulence genes are shown in Table 5. Generally, most of the virulence genes, except the hyl gene, appeared in E. faecalis. Among E. faecium strains, only human isolates were associated with the esp (76.2%) and hyl (66.7%) virulence genes. In other species, E. avium strains isolated from companion animals (1/1 isolate) and human beings (2/3 isolates) contained esp. The E. saccharolyticus strain isolated from companion animals (n = 1) contained hyl. Enterococcus hirae, E. gilvus, and E. pseudoavium, but E. canintestini did not have any virulence genes.
Incidence of virulence genes among enterococcal isolates.*
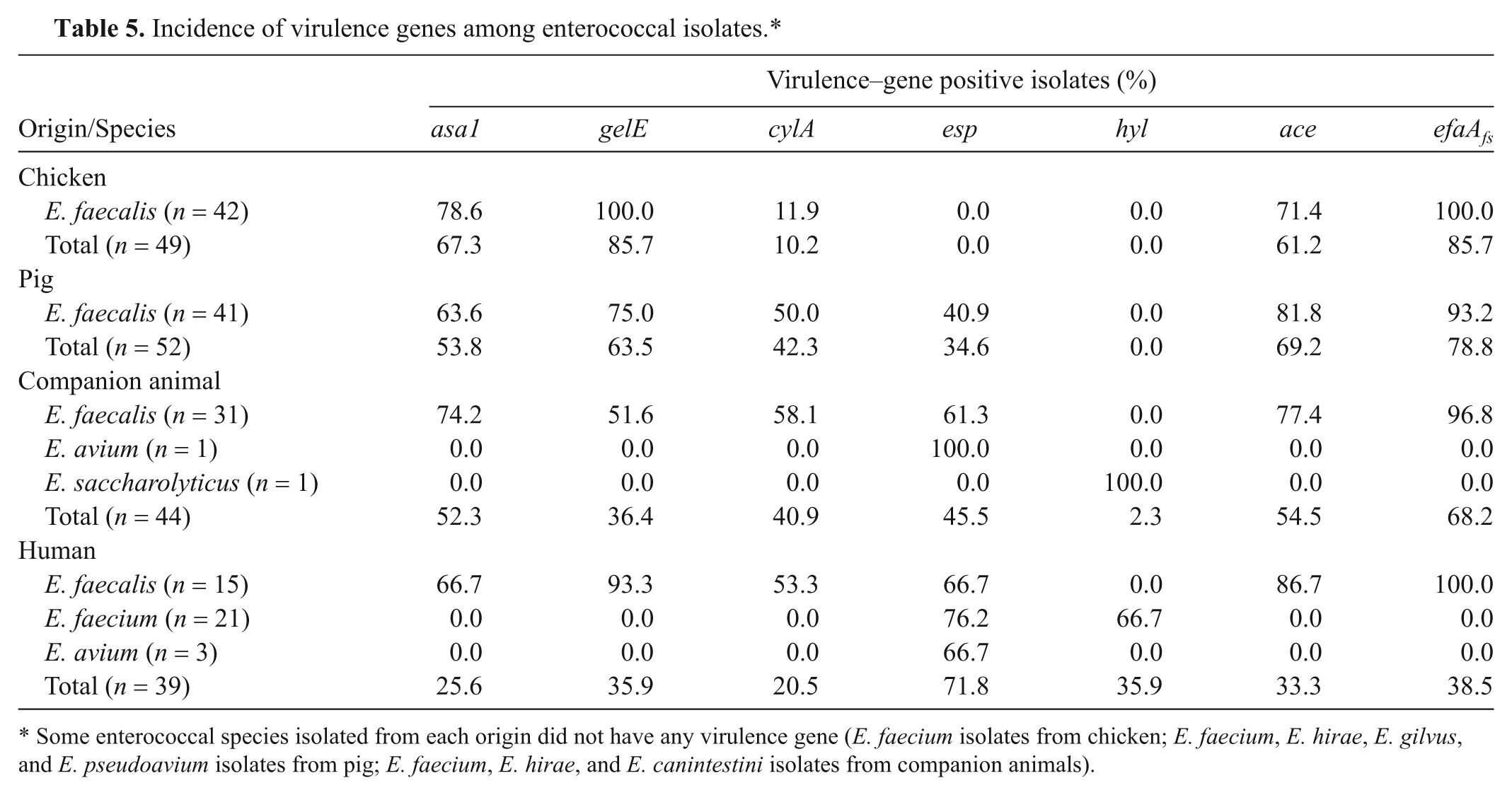
Some enterococcal species isolated from each origin did not have any virulence gene (E. faecium isolates from chicken; E. faecium, E. hirae, E. gilvus, and E. pseudoavium isolates from pig; E. faecium, E. hirae, and E. canintestini isolates from companion animals).
Multilocus sequence typing
Among the 3 AREF isolated from companion animals (dogs), sequence type (ST)202 (n = 1), ST590 (n = 1), and ST591 (n = 1) were detected, with ST202 being part of CC17. Twenty-one human AREF isolates represented the following 6 sequence types: ST78 (n = 9, 42.9%), ST192 (n = 7, 33.3%), ST17 (n = 2, 9.5%), ST18 (n = 1, 4.8%), ST323 (n = 1, 4.8%), and ST631 (n = 1, 4.8%). All the sequence types identified in human-sourced AREF belonged to CC17.
Discussion
Enterococcus faecalis and E. faecium were detected more than the other species, except in pigs where E. faecium was the third most detected species. Enterococcus faecalis is the major cause of enterococcal infection in human beings, and E. faecium, such as CC17, has also emerged as an epidemic pathogen.34,36 Enterococcus faecalis is the most frequently isolated species in animals as well.12,14,18 In the current study, the human isolates were collected from enterococcal-associated infections while the animal isolates might have included normal flora isolates. Indeed, diverse species, an expected finding with normal flora, were identified from pigs and companion animals, while only E. faecalis and E. faecium were detected in chickens. This difference in the species diversity may have been influenced by the isolation and identification methodology, the age of the animals, feed, and geographical differences.12,18
Vancomycin-resistant enterococci harboring vanA were only detected in human isolates (15.4%), which was consistent with studies in other countries, including Japan, Italy, and Portugal where no VRE has been reported among animal enterococcal isolates.18,26,29 Vancomycin-resistant enterococci detection rates have gradually decreased among food animal isolates after a ban on avoparcin as a feed additive. In Korea, the use of avoparcin in feed was banned in 1998. 22 In 2002, the incidence of VRE was reported as 16.7% in chickens and 1.9% in pigs. 22 In 2003, 7.7% of enterococci from Korean poultry were VRE. 15 As VRE were not detected among animal isolates (12 years after the banning of avoparcin), it appears that the prohibition on the use of glycopeptides in food animals might be effective in Korea. The absence of VRE detection among companion animal isolates could be related to the rare use of glycopeptides in companion animal medicine. 38 In contrast, VRE have been steadily isolated from hospitalized human patients at rates of 5.7–32%,2,21,41 and were also detected in the current study. It appears that a decrease in VRE among animal enterococci might not immediately contribute to a decrease in detection in human beings.
Erythromycin and tetracycline resistance rates were generally high in all sources, presumably reflecting the use of erythromycin and tetracycline in human and animal therapy. The incidence of chloramphenicol resistance was high only among companion animals, which may have been caused by the chloramphenicol ban in food animals in 1991 in Korea and the observation that it is used only for limited purposes in human medicine 19 and for treating companion animals and horses. The HLGR rates of E. faecium were high in companion animals and human beings but not in food animals (P < 0.05). This similarity between companion animal and human isolates might reflect the similar therapeutic antimicrobial use in human and companion animal medicine. Taken together, these results suggest that antimicrobials should be used more prudentially in companion animals as well as human beings.
Penicillin and ampicillin are important antimicrobials for enterococcal infections. In particular, the combination of β-lactams and aminoglycosides leads to synergistic bactericidal effects. 40 No ampicillin-resistant isolates were detected in food animals in the present study. However, companion animal and human isolates showed 11.4% and 56.4% ampicillin resistance rates, respectively. Ciprofloxacin resistance rates among companion animals and human beings were also higher than those of food animals (P < 0.05). Although there was no detection of ampicillin-resistant isolates from food animals in the current study, there has been a report of CC17 in food animal samples elsewhere. 8 Thus, screening of ampicillin-resistant enterococci and major human-adapted strains, such as CC17, continues to remain important in Korea.
Multidrug resistance rates of companion animal and human isolates were higher than those of food animals (P < 0.05). Moreover, the most prevalent antimicrobial resistance gene patterns of the isolates were similar between companion animals and human beings except for the cat gene, suggesting that MDR is associated with intensive antimicrobial therapy in human and companion animal medicine. Multidrug-resistant enterococci can be a serious problem not only in human hospitals but also in veterinary clinics, causing an increasing therapy failure rate.
In the current study, an E. saccharolyticus isolate from a companion animal sample was resistant to most of the antibiotics tested except vancomycin and harbored 5 resistance genes. It is presumed that minor enterococcal species such as this E. saccharolyticus isolate might be active reservoirs for antimicrobial resistance genes. More screening and investigations should be performed targeting minor enterococcal species as a pool of antimicrobial resistance genes.
Among virulence factors, almost all genes except for the hyl gene were detected from E. faecalis from all 4 sources. Interestingly, E. faecalis isolates with more than 4 virulence genes were detected in human beings and animals at similar rates, although the animal isolates probably included normal flora. The abundant number of virulence genes in E. faecalis might be needed for colonization and infection of the species in human beings and animals. In contrast, the esp and hyl genes among the E. faecium, E. avium, and E. saccharolyticus strains were only detected from companion animals and human isolates. The esp and hyl genes are linked with ampicillin- and ciprofloxacin-resistant enterococci, particularly in CC17. 35 Indeed, 12 out of 21 human CC17 E. faecium isolates in the current study had the esp and hyl genes (data not shown). Those 2 virulence factors seem to be related with the adaptation of E. faecium and other species to human and companion animal hospital settings.
Multilocus sequence typing was performed with 21 human and 3 companion animal AREF strains to detect the presence of isolates belonging to CC17. All human isolates and 1 companion animal isolate were classified as CC17. The CC17 isolate from a canine patient may have been transferred from a human being, reflecting frequent contact between human beings and animals. Alternatively, it is also possible that CC17 is adapting to veterinary hospital settings independently. Larger scale surveillance to search for CC17 among companion animals and studies comparing human isolates are required.
Fortunately, no VRE was detected in food and companion animal isolates in the current study. Enterococcus faecium isolate belonging to CC17 was detected in 1 companion animal and 21 human beings. As companion animals come in frequent contact with human beings and receive intensive drug therapy individually for disease treatment like human patients, more attention is required for the use of antibiotics in companion animal clinics as well as human hospitals. The epidemiological relationships among enterococcal isolates from food animals, companion animals, and human patients should be further studied in-depth.
Footnotes
Acknowledgements
The Asian Bacterial Bank of Asia Pacific Foundation for Infectious Diseases provided the human enterococci isolates.
Notes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, Forestry and Fisheries (IPET 110069-2 and 110032-3) and the Korea Food and Drug Administration (10162Antibiotics113 and 11162KFDA157). Additional support was provided by the Research Institute of Veterinary Science, Department of Veterinary Microbiology, College of Veterinary Medicine, and the BK21 Program for Veterinary Science, Seoul National University, Seoul, Korea.AQ:Please verify that this statement is accurate and correct.
