Abstract
Cutaneous papillomatosis was diagnosed in an adult American beaver (Castor canadensis). Gross lesions included numerous exophytic, roughly circular, lightly pigmented lesions on hairless areas of fore and hind feet and the nose. The most significant histopathologic findings were multifocal papilliform hyperplasia of the superficial stratified squamous epithelium, with multifocal koilocytes, and multiple cells with large, darkly basophilic intranuclear inclusion bodies. A virus with properties consistent with papillomavirus (PV) was recovered by virus isolation of skin lesions, utilizing rabbit and feline kidney cell lines. The presence of the virus was confirmed by PV-specific polymerase chain reaction. The partial sequences of E1 and L1 genes did not closely match those of any PVs in GenBank, suggesting that this might be a new type of PV. Partial E1 and L1 nucleotide sequences of the beaver papillomavirus (hereafter, ARbeaver-PV1) were used to create a phylogenetic tree employing the complete E1 and L1 open reading frame nucleotide sequences of 68 PVs. The phylogenetic tree placed the ARbeaver-PV1 in a clade that included the Mupapillomavirus (HPV1 and HPV63) and Kappapapillomavirus (OcPV1 and SfPV1) genera. The present article confirms the papillomaviral etiology of cutaneous exophytic lesions in the beaver.
Keywords
The Papillomaviridae family includes a group of circular, double-stranded DNA viruses that possess a small genome (6.8–8.4 kb). Papillomaviruses (PVs) consist of a nonenveloped icosahedral capsid structure that is 50–55 nm in diameter. Current virus taxonomy of the International Committee on Taxonomy of Viruses divides Papillomaviridae into 29 genera, Alpha- through Dyoiotapapillomavirus. 1 Papillomaviruses cause proliferations of the stratified squamous epithelium of skin or mucosa in a wide variety of animals including domesticated and wild species.5,8,9,12,13,16 Transmission by direct contact is common among animals of the same or phylogenetically closely related species. Approximately 50 mammalian species are reported to be infected by PVs. 12
A single case of cutaneous papillomatosis in an American beaver (Castor canadensis) was documented almost 3 decades ago. 2 In addition to gross and histologic diagnosis, the viral etiology of tumors was supported by electron microscopy that identified numerous uniform, virus-like particles of approximately 46 nm in diameter within nuclei of superficial layers of the epidermis. However, a positive peroxidase–antiperoxidase test for PV-specific antigen only suggested that the lesions had been caused by PVs. Thus, to date, papillomatosis in the beaver has not yet been confirmed at the molecular level.
The purpose of the current article is to report a novel PV isolated from proliferative skin lesions of a wild American beaver (Castor canadensis) and confirmed by polymerase chain reaction (PCR). When the PCR amplicons were directly sequenced, the sequences did not closely match that of any PVs in GenBank, suggesting that this is a new type of PV. The isolated PV was tentatively designated “ARbeaver-PV1.” Furthermore, a maximum likelihood phylogenetic tree placed the ARbeaver-PV1 in a clade that included the Mupapillomavirus (HPV1 and HPV63) and Kappapapillomavirus (OcPV1 and SfPV1) genera.
In March 2011, fore and hind feet from an adult (>2 years old) wild American beaver were submitted to the Washington Animal Disease Diagnostic Laboratory (WADDL) from the Washington Department of Fish and Wildlife and U.S. Department of Agriculture–Animal and Plant Health Inspection Service–Wildlife Services in Washington state. The beaver had been lethally trapped in Western Washington and was in good body condition. On gross examination, hairless areas on the dorsal, and plantar and palmar aspects of the fore and hind feet, respectively, were severely affected by numerous, pinpoint to 2 mm in diameter, slightly raised, roughly circular, lightly pigmented lesions (Fig. 1). In addition to the lesions on the fore and hind feet, exophytic lesions were also observed on the nose of this animal (per submitter). Reportedly, another lethally trapped beaver from the same location had similar skin lesions, but was not submitted to WADDL.
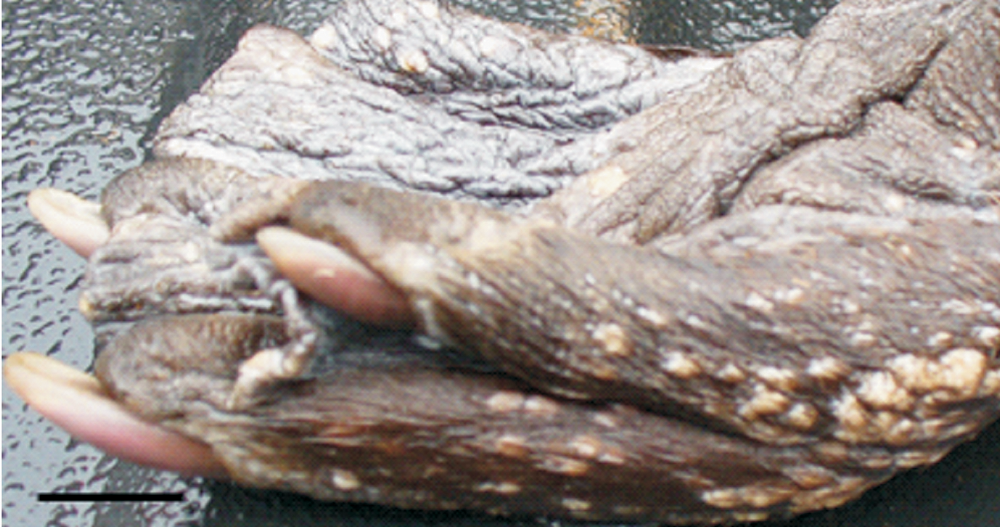

Gross pathology of the affected hind foot of the American beaver (Castor canadensis). The dorsal and plantar aspects of the feet were severely affected by numerous pinpoint to 2 mm diameter, coalescing, slightly raised, roughly circular, and lightly pigmented lesions. Bar = 1 cm.
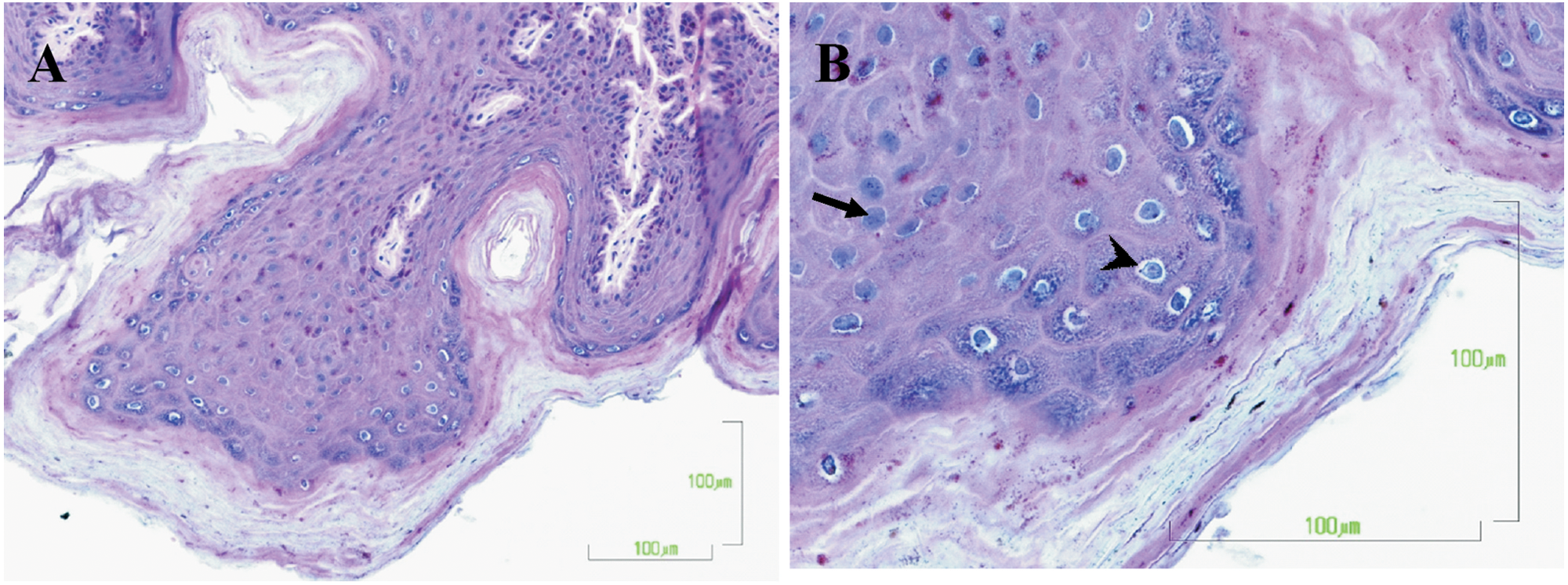
Microscopically, the cutaneous masses were composed of hyperplastic stratified squamous epithelium arranged in papilliform projections supported by thin stalks of fibrovascular tissue. The epidermis was multifocally acanthotic with prominent orthokeratotic hyperkeratosis. Often, individual or clusters of keratinocytes in the stratum spinosum and stratum granulosum had hyperchromatic nuclei and perinuclear clearing consistent with koilocytosis (Fig. 2a). Individual cells within the stratum granulosum had swollen, lightly amphophilic cytoplasm with nuclei filled with round to oval, 8–10 µm diameter, darkly basophilic inclusion bodies (Fig. 2b). There was intercellular edema with prominent intercellular bridges among epidermal cells (spongiosis). Bacterial culture detected moderate numbers of Corynebacterium sp. and Micrococcus sp., as well as numerous Propionibacterium sp., on the skin. Mycologic tests identified Acremonium sp. and Penicillium sp. No clinically important bacterial pathogens were identified.
American beaver (Castor canadensis).
Virus isolation was attempted on selected areas of affected skin tissues from feet. The tissues were soaked in minimal essential medium (MEM) with penicillin, streptomycin, gentamicin, and amphotericin B overnight at 4°C. The tissues were minced into 2–3 mm in diameter tissue explants and/or homogenized with tissue grinder or tissue stomacher prior to inoculation onto 70–85% confluent cell culture of Madin–Darby canine kidney (MDCK), Crandell feline kidney (CRFK), rabbit kidney–derived (RK)-13B, or African green monkey kidney epithelial (Vero) cells. The cell cultures were kept at 37°C with 5% CO2. At day 5 postinoculation (pi), the cell cultures were harvested for subsequent passage. To liberate the suspected cell-associated virus, the cells were frozen at –60°C for 1 hr, and then quickly thawed in a 37°C water bath. Tissue explants and cellular debris were pelleted at 3,000 × g for 5 min, and supernatants were transferred to 24-hour-old cell cultures (passage 2) and observed for 5–10 days. Passages 3 and 4 were performed only on RK-13 and CRFK cell cultures because no cytopathic effect (CPE) was observed in Vero or MDCK cells (Fig. 3). Uninfected cell lines (negative controls) did not show any CPE. Cytopathic effect was progressive over 5–7 days and was characterized by enlarged, rounded, and refractile cells arranged in loose clusters. Infected RK-13 cells had a propensity to exhibit CPE upon passage much earlier, usually within the first week pi, whereas CPE on the infected CRFK cell line appeared later, at 3–4 weeks pi. (Long-term [>2 weeks] maintenance of infected cell cultures was achieved by replacement of the culture media with MEM with 5% fetal bovine serum every 4–6 days.) Chloroform sensitivity was performed at passage 3 by mixing CHCl3 with tissue supernatant at a 1:1 ratio for 10 min at room temperature. The treatment with chloroform had no effect on virus viability in RK-13 or CRFK cells, indicating that the virus was nonenveloped and consistent with properties of PVs.
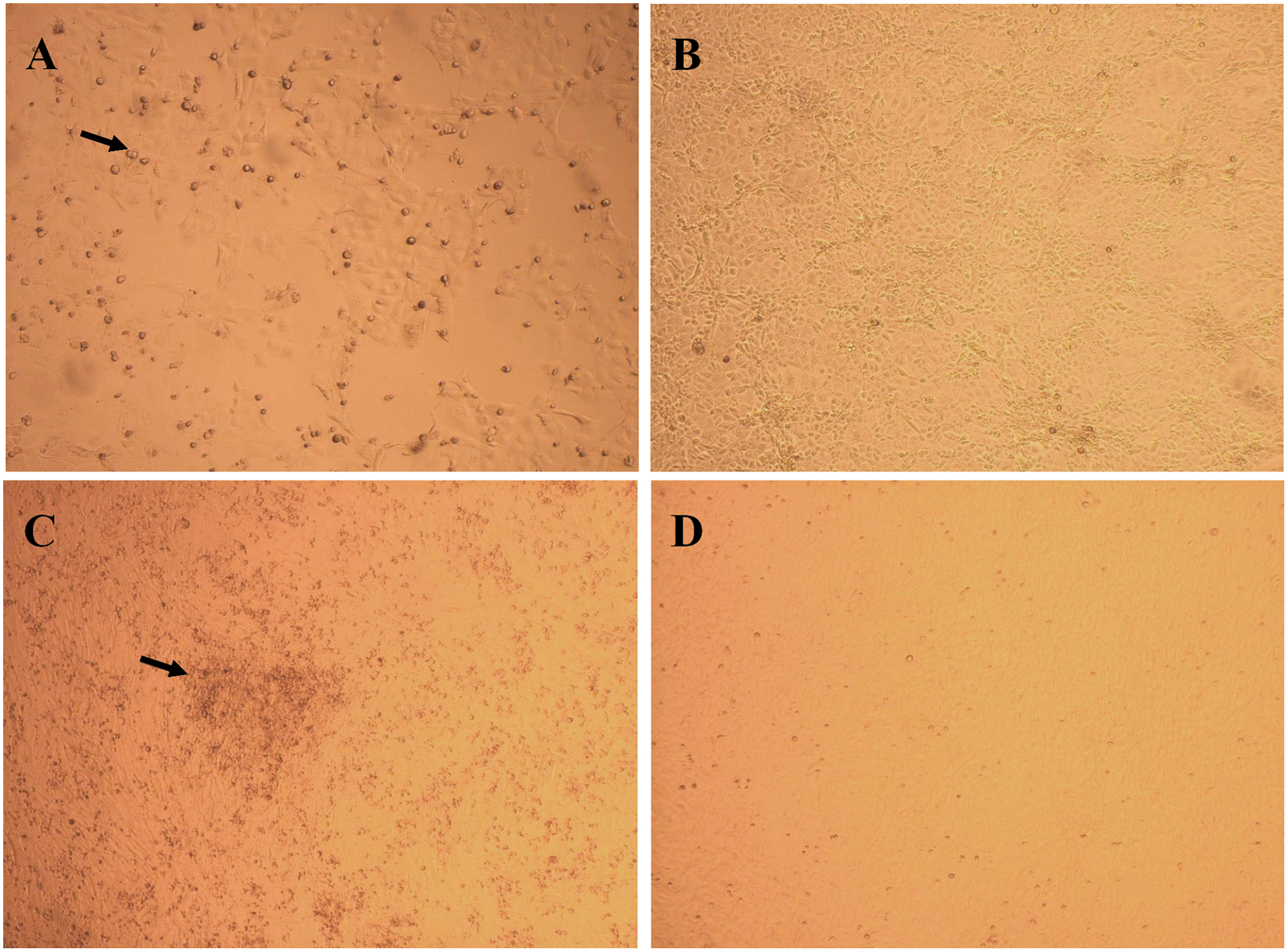
Virus isolation on rabbit kidney–derived (RK)-13B and Crandell feline kidney (CRFK) cell lines.
The supernatant fluids from infected RK-13 and CRFK cell cultures, fresh (frozen) skin, and formalin-fixed, paraffin-embedded sections of skin were subjected to multiplex hot start polymerase chain reaction (PCR) followed by sequencing. Specifically, DNA was extracted by using a commercial kit a as previously described. 15 Two sets of previously described PV-specific degenerate primers, AR-E1F1/AR-E1R4 and AR-L1F1/AR-L1R3, were used for amplification of partial E1 and L1 sequences, respectively. 10 Each 50-µl PCR reaction consisted of 5 µl of 10× PCR buffer with NH4(SO)4, b 4 µl of 25 mM Mg2Cl, 4 µl of 2.5 mM deoxyribonucleotide triphosphates, 1.5 µl of 40 µM primer stocks, 0.5 U of Taq polymerase, b 24 µl of sterile diethylpyrocarbonate (DEPC)-treated water, and 9.5 µl of template. Extracted sterile DEPC-treated water was used as a negative extraction control for the PCR reaction. The PCR cycle was composed of 3 min of denaturing at 95°C, followed by 40 amplification cycles consisting of 1 min of denaturing at 95°C, 1 min annealing at 54°C, and 1 min extension at 72°C, with a final extension of 7 min at 72°C. Positive results were obtained from all the tissues and infected cell lines as compared to the negative control. The bands of 551 bp and 610 bp corresponding to the expected sizes of E1 and L1 amplicons of PVs were visualized and directly sequenced in both directions as previously described. 15 The partial sequences of E1 and L1 genes did not closely match that of any PVs in GenBank, suggesting that this is a new type of PV with 72% and 71% sequence identities to Rabbit oral papillomavirus (GenBank accession no. AF227240.1) and HPV63 (GenBank accession no. X70828.1), respectively.
Additional assays were performed in order to rule out poxvirus. Two pan-pox universal PCR assays were performed as previously described, 7 using Orf virus as a positive control and water as a negative control. Neither of these assays detected poxvirus from any of the cultured, fresh, or tissue block samples.
Partial E1 and L1 nucleotide sequences of the beaver PV were used to create a phylogenetic tree employing the complete E1 and L1 open reading frame (ORF) nucleotide sequences of 68 PVs representing each species (see Table 3 1 ). The nucleotide sequences of each ORF were aligned using the corresponding aligned amino acid sequences (MAFFT v6.864b 6 ) using SeaView version 4.3 3 and then concatenated. Maximum likelihood (ML) trees were constructed using RAxML MPI v7.2.8.27 11 and PhyML MPI v1.4.3 4 and were visualized with MEGA5. 14 The GTR + gamma model was set for among-site rate variation and allowed substitution rates of aligned sequences to be different. Data were bootstrap resampled 100 times in PhyML. The constructed trees place the ARbeaver-PV1 in a clade that includes the Mupapillomavirus (HPV1 and HPV63) and Kappapapillomavirus (OcPV1 and SfPV1) genera (Fig. 4).
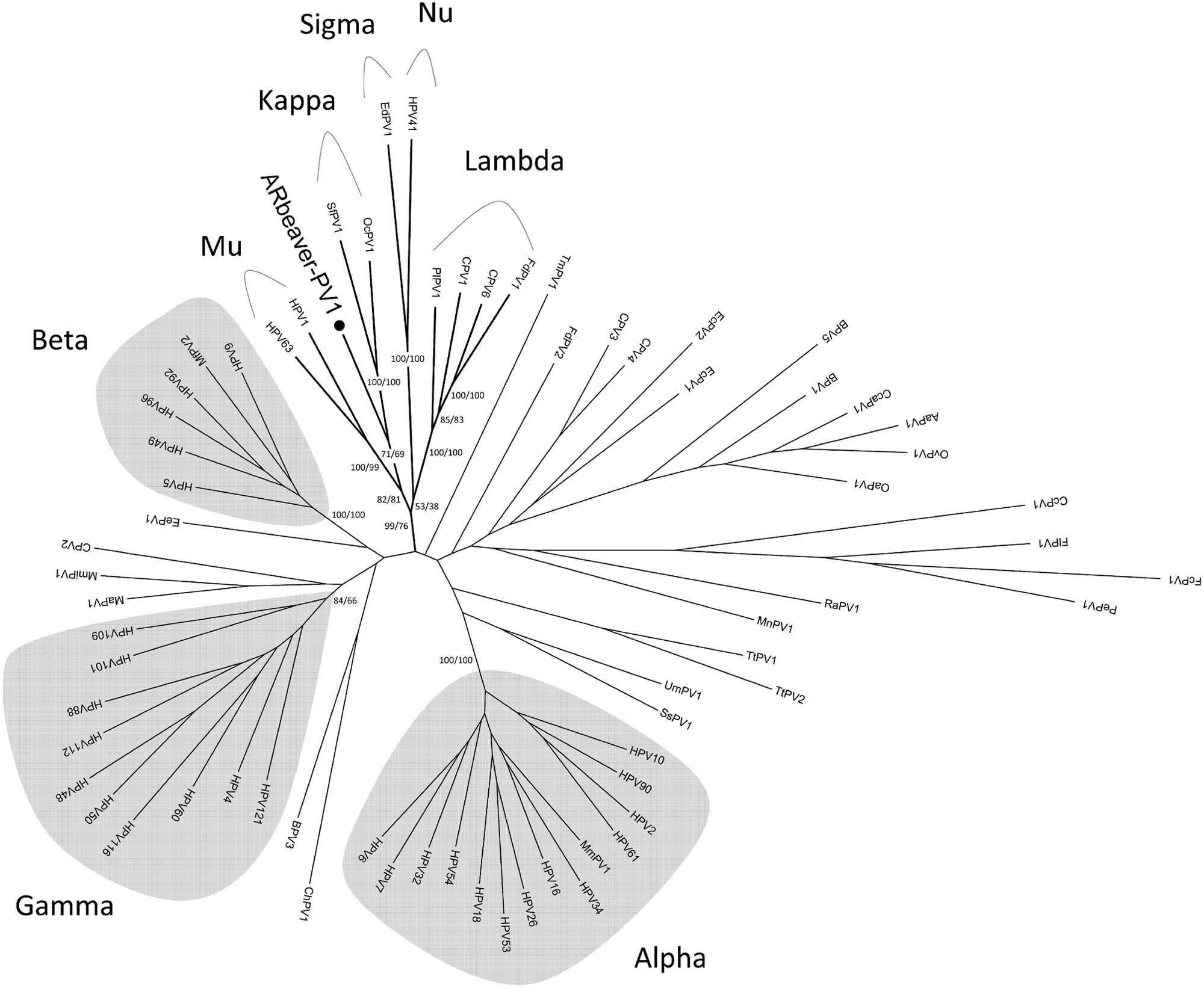
Phylogenetic position of a beaver papillomavirus (ARbeaver-PV1). Maximum likelihood trees were constructed using RAxML and PhyML inferred from the concatenated E1 and L1 nucleotide sequences as described in the Materials and Methods section. The topology was taken from the RAxML tree, with RAxML and PhyML bootstrap support values (percentages) of key taxa groups displayed on or near the branch nodes. The names of papillomavirus genera are shown distal to the relevant clade groups. The Human papillomavirus–containing genera are highlighted in gray.
The present study describes a case of papillomatosis in a wild American beaver, which manifested as exophytic skin lesions on hairless areas of feet and nose. A single documented case of proliferative skin lesions in Castor canadensis only suggested a papillomaviral etiology. 2 The gross and histopathological findings in the current report are consistent with the previous description of cutaneous form of PV infection in the beaver as well as other animal species.2,12,13 A virus with properties consistent with PVs was isolated utilizing RK-13 and CRFK cell lines. Polymerase chain reaction amplification and sequence analysis of the PV-specific genes identified a novel PV in the skin lesions and infected cell lines. The current report confirms the papillomaviral etiology of cutaneous exophytic lesions in the beaver and molecularly characterizes the isolated virus through phylogenetic analysis.
Footnotes
Acknowledgements
The authors would like to thank Dr. Kathleen Potter and Ms. Lorraine Tanaka for taking pictures of histological sections and tissue cultures, respectively. Mr. Mat Craig is acknowledged for reporting the lesions.
a.
Qiagen Inc., Valencia, CA.
b.
Fermentas Inc., Glen Burnie, MD.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The current study was funded by the Washington Animal Disease Diagnostic Laboratory, Pullman, WA, and the Washington Department of Fish and Wildlife.
