Abstract
Bovine papillomaviruses (BPVs) are widespread pathogens mainly associated with benign, self-limiting, cutaneous lesions (warts). At least 8 viral types, defined by serology or nucleotide sequences of the L1 gene, have been identified to date. Different serotypes are associated with the specific type and morphology of the lesion and with particular geographical regions. This article describes the molecular identification of papillomaviruses from Brazilian cattle (n = 48) and horses (n = 1) through partial amplification and sequencing of the L1 gene. Bovine papillomavirus–1 (BPV-1) was identified in warts from 29 cattle (59%), BPV-6 from 9 cattle (18%), and BPV-2 in 8 lesions (16%). Warts of 2 cattle harbored L1 sequences of a new BPV type (BAA5), otherwise identified almost exclusively in healthy skin. The newly proposed BPV type “BR-UEL-4” was identified in a sarcoid tumor of a horse. Thus, the present report provides information on the main types of BPV involved in bovine papillomatosis in Brazil and reveals a new viral type associated with equine sarcoid, which to date has been attributed exclusively to BPV-1 and BPV-2.
Papillomaviruses (PVs) are small, nonenveloped, double-stranded DNA viruses with circular genomes that replicate in the nucleus of infected cells. 1 The Papillomaviridae family comprises 16 virus genera classified based on genomic DNA homology, especially of the late L1 structural protein gene. 1 Bovine papillomaviruses (BPVs) are frequently associated with tumors of cutaneous or mueosal epithelia, known as papillomas or warts. 10 Cutaneous papillomas of cattle are usually benign and tend to regress spontaneously. 1,4 Bovine papillomaviruses are classified into 6 distinct types (BPV-1 through BPV-6). 1 However, recent studies 3,6 indicate the inclusion of new viral types based on the variability of the L1 gene. Bovine papillomavirus–1 (genus Deltapapillomavirus) and BPV-2 infect the epithelium and dermis; BPV-3 (genus Xipapillomavirus), BPV-4, and BPV-6 are usually epitheliotropic; and BPV-5 (genus Epsilonpapillomavirus) infects the epithelia and dermis. 1
Papillomaviruses infect a large range of hosts in nature, yet each virus species is generally species-specific. 10 Exceptions include BPV-1 and BPV-2, which have been frequently associated with sarcoid tumors in horses. 10 In cattle, BPV infection has been associated with 4 main clinical presentations linked with specific viral types: 1) cutaneous fibropapillomas (BPV-1 and BPV-2); 2) cutaneous papillomas (BPV-3); 3) gastrointestinal papillomas (BPV-4) and teat fibropapillomas (BPV-5); and 4) cutaneous teat papillomas (BPV-6). 8,10
In Brazil, cutaneous papillomatosis (warts) is widespread among beef and dairy cattle, yet few studies 6,8,12,14 have focused on the identification of BPV types. Some studies 12,14,15 have identified BPV-1 and BPV-6 as the main types involved in cutaneous warts, whereas BPV-2 has been associated with enzootic hematuria. Recently, a new viral type (BPV-8) was identified in warts from cattle in the state of Paraná, Brazil. 7 The present study reports the molecular identification of BPVs from cutaneous warts in cattle in southern Brazil and from a sarcoid tumor of a horse from the southeast region of the country.
Forty-three bovine samples were submitted to the Virology Laboratory of the Universidade Federal de Santa Maria (Brazil) for the production of an autogenous vaccine; 5 papillomas were collected by veterinarians. The samples were received between 2007 and 2009 from herds in the states of Rio Grande do Sul (43 herds), Paraná (3 herds), and Santa Catarina (2 herds). Although many submissions contained papillomas from more than 1 animal from the same herd, a single sample per herd was included in the analysis. The equine sample was submitted from the state of Rio de Janeiro (2009) for the identification of the etiological agent.
A fragment of approximately 100 mg of each papilloma was minced with a sterile razor blade and subjected to DNA extraction using 900 μl of lysis buffer (10 mM Tris, 10 mM ethylenediaminetetraacetic acid [EDTA], 0.5% Nonidet, a 1% sodium dodecy1 sulfate, 0.001 mg/ml proteinase K), followed by 1 hr of incubation at 56°C and phenol:chloroform extraction. After extraction, the DNA was solubilized in Tris–EDTA buffer and stored at −20°C until testing. The DNA concentration was measured by ultraviolet (UV) absorbance at 260 nm. For the L1 gene amplification, the following sets of degenerate primers were used: FAP59/FAP64 (5′-TAACWGTIGGI-CAYCCWTATT-3′ and 5′-CCWATATCWVHCATIT-CICCATC-3′), generating a product of 478 base pairs (bp) 9 ; and MY09/MY11 (5′-GCMCAGGGWCATAAYAATGG-3′ and 5′-CGTCCMARRGGAWACTGATC-3′), with an expected product of 450 bp. 11 DNA from all papillomas was initially analyzed by polymerase chain reaction (PCR) with a set of FAP primers. Samples that were not amplified by these primers were then analyzed by PCR using MY primers. Both sets of primers amplify sequences within the L1 gene and have been widely used for molecular identification and phylogeny of BPVs. 9,11
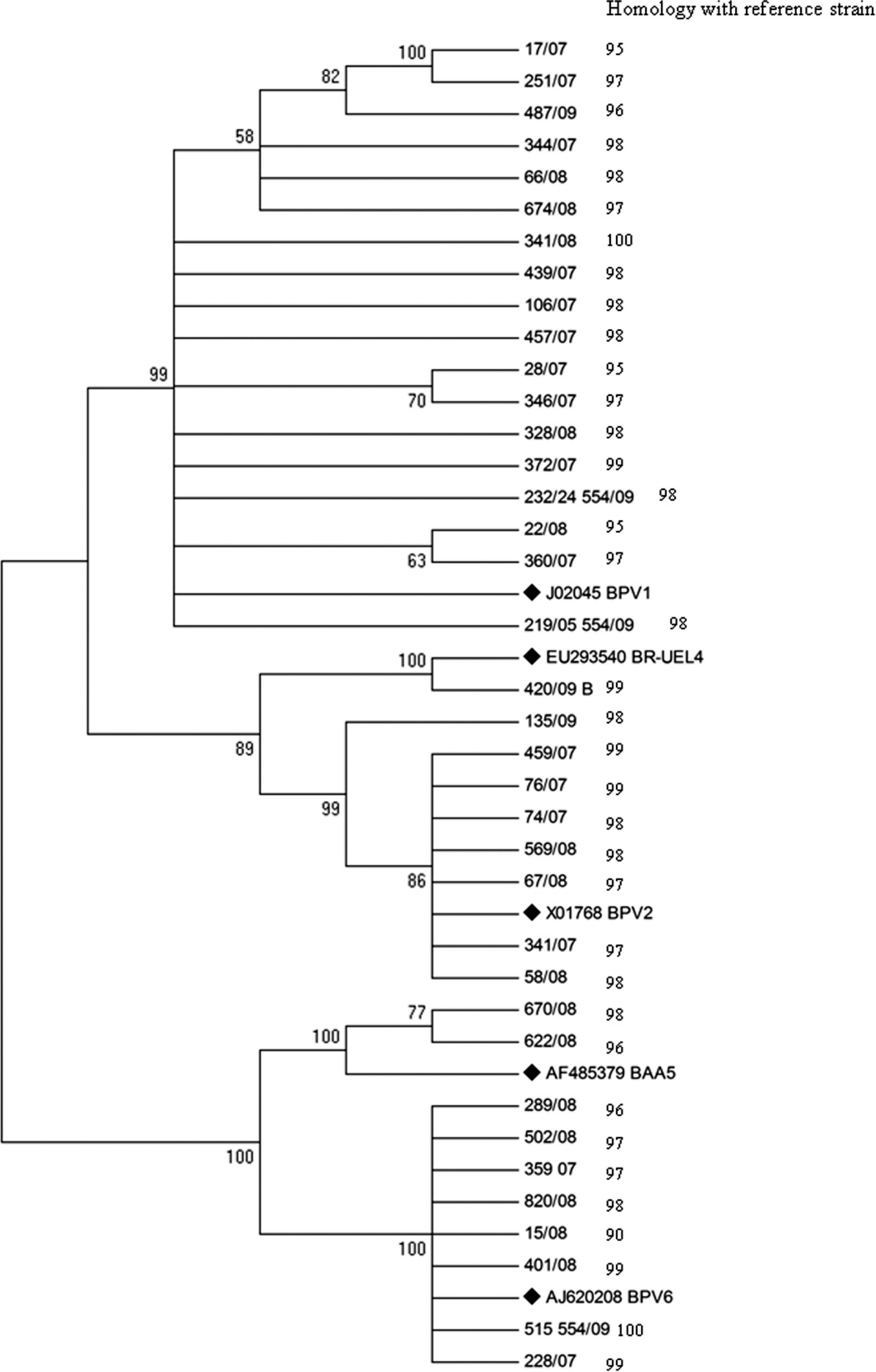
Neighbor-joining phylogenetic tree constructed with bovine papillomavirus (BPV) sequences obtained using primers FAP59/ FAP64 and reference strains for each viral type. The numbers in the branches represent the bootstrap support values determined for 1,000 replicates. The reference strains for each viral type are indicated by ♦ The following reference sequences were used: BPV-1 (GenBank accession no. J02045); BPV-2 (accession no. X01768); BPV-6 (accession no. AJ620208); BR-UEL-4 (accession no. EU293540); and BAA5 (accession no. AF485379). The number on the right side represents the percentage (%) of nucleotide homology between each sequence and the respective reference strain.
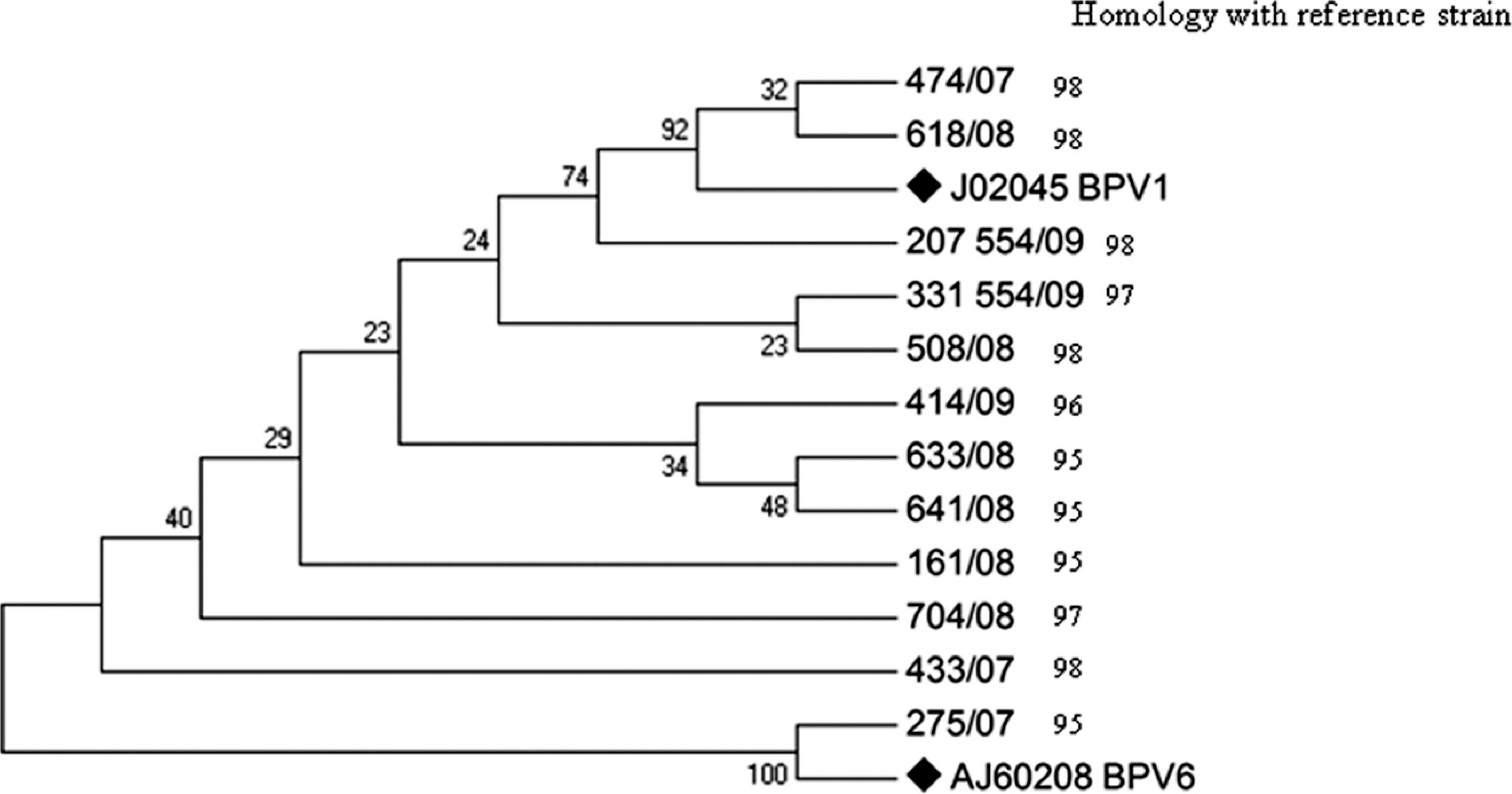

Neighbor-joining phylogenetic tree constructed with bovine papillomavirus (BPV) sequences obtained using primers MY09/MY11 and reference strains for each viral type. The numbers in the branches represent the bootstrap support values determined for 1,000 replicates. The reference strains for each viral type are indicated by ♦. The following reference sequences were used: BPV-1 (GenBank accession no. J02045) and BPV-6 (accession no. AJ620208). The number on the right side represents the percentage (%) of nucleotide homology between each sequence and the respective reference strain.
Polymerase chain reactions were performed in a 50-μl volume, using approximately 1 μg of total DNA, 0.5 μM of each primer, 2.5 mM MgC12, 0.2 mM of each deoxyribonucleotide triphosphate, 1X reaction buffer, and 0.75 U of Taq b polymerase. The PCR conditions were initial denaturation (94°C/10 min), followed by 40 cycles (94°C/50 sec; annealing at 50°C, or 55°C/55 sec, respectively [FAP and MY]; 72°C/50 sec; and a final extension of 7 min at 72°C). The PCR products were visualized in a 1.5% agarose gel, stained with ethidium bromide, and visualized under UV light. The amplicons produced by FAP or MY primers were purified using a commercial kit, c according to the manufacturer's instructions. Each amplicon was sequenced 3 times using a dye terminator kit d and then analyzed with the Staden Package (http://staden.sourceforge.net/). The consensus sequence generated was compared online using the BLAST tool (http://www.ncbi.nlm.nih.gov/blast/Blast. cgi). The phylogenic trees were constructed using Molecular Evolutionary Genetics Analysis software version 4.1, 13 using the neighbor-joining method with 1,000 bootstrap replicates.
Thirty-seven out of 49 samples were amplified by using the FAP primers, and 12 required the use of the MY set. Twenty-nine bovine papillomas (59%) harbored BPV-1 sequences (>95% nucleotide [nt] homology with reference strain BPV-1; GenBank accession no. J02045); 8 (16%) harbored BPV-2 (>97% nt homology with a BPV-2; accession no. X01768), and 9 samples (18%) contained BPV-6 sequences (>95% nt homology with a BPV-6; accession no. AJ620208). Two papillomas (4%) harbored sequences very similar to a recently identified BPV (BAA5), 3 displaying a nucleotide homology of 96% and 98%, respectively, with the BAA5 sequence (accession no. AF485379).
Sequences obtained using FAP and MY primers were subjected to phylogenic analysis and used to generate 2 independent trees. Figure 1 shows the genetic relationship among sequences amplified with FAP primers and GenBank reference strains. Figure 2 shows the genetic relationship among sequences obtained by amplification and sequencing using the MY set of primers and reference strains. The nt homology between each sample and the respective reference strain is also shown.
Bovine papillomavirus-1 was the main viral type identified in bovine papillomas (59%), followed by BPV-6 (18%) and BPV-2 (16%). Even though papillomatosis is widespread among Brazilian cattle, few studies have identified the BPV types involved with the lesions. One 8 of these studies analyzed warts from 9 animals from 4 herds; another 6 analyzed 22 papillomas from 4 herds. Both studies 6,8 reported BPV-6 as the most prevalent BPV viral type, followed by BPV-1. In addition, 4 new putative BPV types were reported in one study, 6 including the proposed type BR-UEL-4 identified in skin papillomas of cows from 2 herds from the state of Paraná.
Two papillomas analyzed in the present study harbored sequences showing 96% and 98% homology, respectively, with the reference strain BAA5 (accession no. AF485379). This BPV type was first identified in 2002 in a swab collected from healthy bovine skin 3 and was subsequently identified in teat papillomas and healthy skin as well. 11 Even though the presence of BPVs is usually associated with warts, some reports have indeed identified genomic sequences of BPV-1, BPV-3, BPV-6, BAA1, and BAA5 in healthy teat skin, indicating the occurrence of latent and/or subclinical infections. 11
The present study analyzed the largest number of BPV types associated with warts in Brazil to date (48 bovine specimens; 48 different herds; 3 states). Although the samples were not obtained through a statistically designed sampling, the viral types identified probably reflect the approximate frequency and distribution of BPV types circulating in the field. Unfortunately, many specimens were not accompanied by a detailed description of the morphology, gross aspect, body location, and/or distribution of the lesions, precluding an in-depth analysis of possible associations of specific viral types with these aspects.
The horse sample analyzed herein was collected from an animal with lesions histologically confirmed as sarcoid (Anjos et al., unpublished data). Equine sarcoid is a frequent tumor, the etiology of which has been associated with BPV-1 and, less frequently, with BPV-2. 5 Interestingly, the horse BPV sequence in the present study displayed a 99% homology with another BPV type, BR-UEL-4, 6 recently identified in Brazil. In summary, data in the current study provide important information on the main BPV types associated with bovine papillomatosis in southern Brazil and highlight the involvement of an additional BPV type in equine sarcoid.
Footnotes
a.
Nonided P40, Roche Diagnostics, Mannheim, Germany.
b.
Taq DNA polymerase, Invitrogen Brasil Ltda., São Paulo, Brazil.
c.
PureLink, Invitrogen Brasil Ltda., São Paulo, Brazil.
d.
DYEnamic™ ET Dye Terminator, GE Healthcare, São Paulo, Brazil.
