Abstract
In dogs, papillomaviruses are thought to cause oral and cutaneous papillomas and pigmented plaques. Eight canine papillomaviruses have been fully sequenced to date. Four of these canine papillomaviruses, including
Papillomaviruses (PVs) are epitheliotropic, often species-and tissue-specific viruses within the family
Given the identification of different CPVs amplified from canine pigmented plaques, it is likely that there are additional PV sequences that have not been previously identified. The present study aimed to detect and sequence PV DNA from cutaneous pigmented plaques and identify the associated PV or novel PV sequences through nucleotide sequence analysis.
The pathology archives from the Veterinary Medicine Teaching Hospital (VMTH) at the University of California, Davis (Davis, CA), IDEXX Laboratories Inc. (West Sacramento, CA), Veterinary Diagnostics (VDx) Veterinary Pathology Services (Davis, CA), Yager-Best Histovet (Guelph, Ontario), and an independent biopsy service (Venice, CA) were searched for surgical biopsy specimens from dogs with the following diagnoses: viral plaque and pigmented plaque. In the study, 27 biopsies from 24 individual dogs were identified and included. All biopsies had been routinely formalin fixed and paraffin embedded (FFPE), and a 4-µm section cut for hematoxylin and eosin (HE) staining. The HE-stained sections were reviewed by 2 board-certified pathologists (Luff and Affolter) to confirm the diagnosis.
Immunohistochemistry was performed using a horseradish-peroxidase staining method on 4-µm FFPE sections of tissue from each biopsy. A rabbit polyclonal antibovine PV a antibody against the L1 capsid protein was used as the primary antibody at a dilution of 1:600 for 1 hr. The secondary antibody, biotinylated goat antirabbit IgG, b was used at a dilution of 1:500 for 30 min. The positive control was an oral papilloma. Omission of the primary antibody served as the negative control.
Genomic DNA (gDNA) was extracted from two 25-µm-thick sections of FFPE tissue from each case using a commercially available kit following manufacturer’s recommended protocol. c A 450-base pair region of the L1 ORF was amplified using a single set of degenerate consensus primers (MY09 and MY11). 9 PV DNA was not detected within 4 pigmented plaques (4/27) using the degenerate primer set. For these cases, specific primers for CPV-4 were used to increase the sensitivity for detection of PV DNA in these remaining cases. Specific primers for CPV-4 were chosen because this was the most commonly detected CPV based on initial sequencing results of the other 23 out of 27 pigmented plaques. The specific primers for CPV-4 have been previously published and amplify a 194-bp fragment. 11
Omission of gDNA template served as a negative control for all PCR reactions. The PCR reaction conditions used in the current study have been previously published. 9 All PCR products were electrophoresed through 1.5% agarose gel and stained with a commercially available nucleic acid stain. d The PCR products obtained from every sample (including those generated with specific primers) were purified using a commercially available kit following the manufacturer’s recommended protocol e and submitted to a commercial DNA sequencing laboratory following the laboratory specifications. f The nucleotide sequences generated from the sequencing laboratory were routinely trimmed and edited using commercially available software. g
Identification of new PV types is based on <90% shared DNA homology to any known PV type.
3
Therefore, identification of the PV type associated with each pigmented plaque was defined as having >90% shared nucleotide identity within the
Histologically, the pigmented plaques were characterized by locally extensive epithelial hyperplasia, marked orthokeratotic hyperkeratosis, prominent clumped keratohyalin granules, and hyperpigmentation within the stratum granulosum (Fig. 1A). Koilocytosis can be a subtle feature in pigmented plaques, 5 and it was identified within only 3 biopsies in the current study. Viral inclusion can be identified in pigmented plaques, 12 although this is not a consistent feature,11,15 and it was not identified in any of the biopsies in the current study. The clinical history, including breed of dog and location of the pigmented plaque, for all cases is listed in Table 1. Immunohistochemistry revealed positive nuclear staining in all pigmented plaques. In some cases, the staining was evident within the keratinocytes throughout the stratum spinosum in the proliferative epithelium (Fig. 1B), while in other cases, there were only 1 or 2 immunoreactive keratinocytes along the periphery of the acanthotic epithelium.
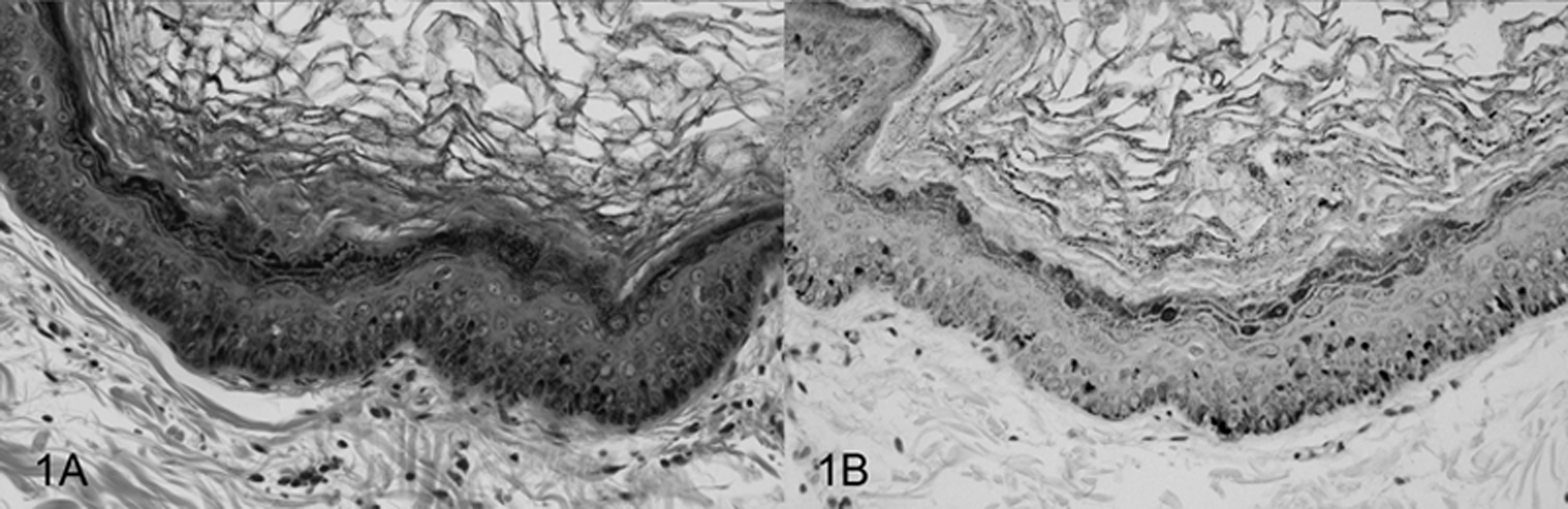
Photomicrographs of a canine pigmented plaque. A, locally extensive epithelial hyperplasia and orthokeratotic hyperkeratosis with prominent keratohyalin granules. Hematoxylin and eosin. B, demonstration of strong nuclear staining for papillomavirus antigen within keratinocytes in the proliferative epithelium. Anti-bovine papillomavirus antibody. Streptavidin–biotin staining method.
Specific Canis familiaris papillomavirus (CPV) or novel papillomavirus sequence (designated Novel CPV-A through F), breed of dog, and location of pigmented plaque detected for all biopsies included in the current study.

Different biopsies from one dog.
DNA from 10 different PVs was identified within the pigmented plaques (listed in Table 1). CPV-4 was detected in 11 out of 27 pigmented plaques, CPV-5 in 2 out of 27, and CPV-3 in 1 out of 27. PV DNA sequences with 99% identity over 342 base pairs to the partial L1 PV sequence (GU220384) previously identified 10 were detected in 2 out of 27 of the pigmented plaques in the present study (JQ040505). The 11 remaining pigmented plaques contained 6 previously unreported PV DNA sequences (designated novel CPV-A through F for the purpose of this study; listed in Table 1). The results of the percentage pairwise nucleotide identities between the 6 novel PV sequences identified in the present study and the most closely related CPVs are shown in Figure 2. Novel CPV-A (JQ040499) is most closely related to CPV-3 over 355 base pairs, novel CPV-B (JQ040500) with CPV-4 over 359 base pairs, novel CPV-C (JQ040501) with CPV-5 over 373 base pairs, and novel CPV-D (JQ040503) with GU220384 over 364 base pairs. 10 Both novel CPV-E (JQ040503) and CPV-F (JQ040504) were most closely related to CPV-8 over 342 and 372 base pairs, respectively. The nucleotide sequence for novel CPV-E was identical to a currently unpublished nucleotide sequence listed in the NCBI database (JF418155.1).
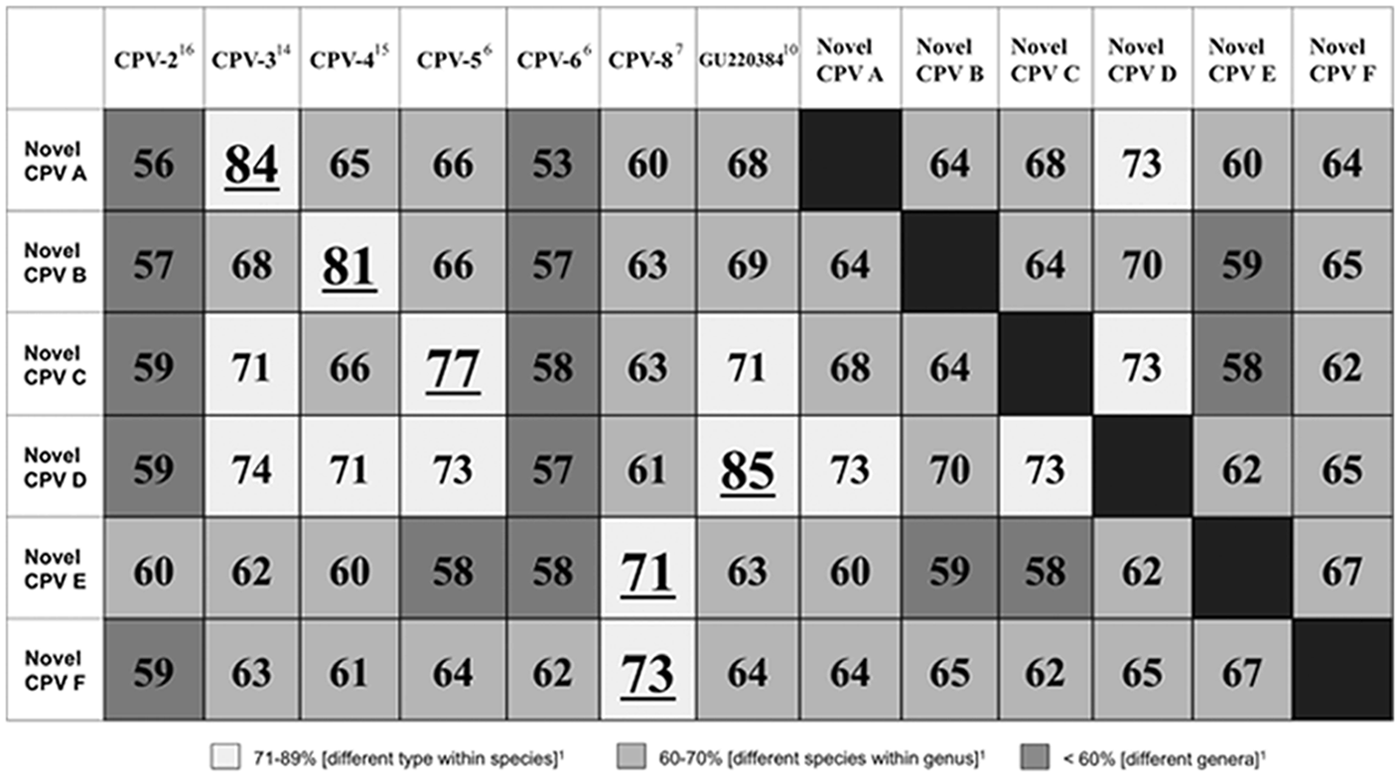
Percentage pairwise nucleotide identities comparing the 6 novel
The neighbor-joining phylogenetic tree generated from the alignment of an approximately 330-bp segment of the
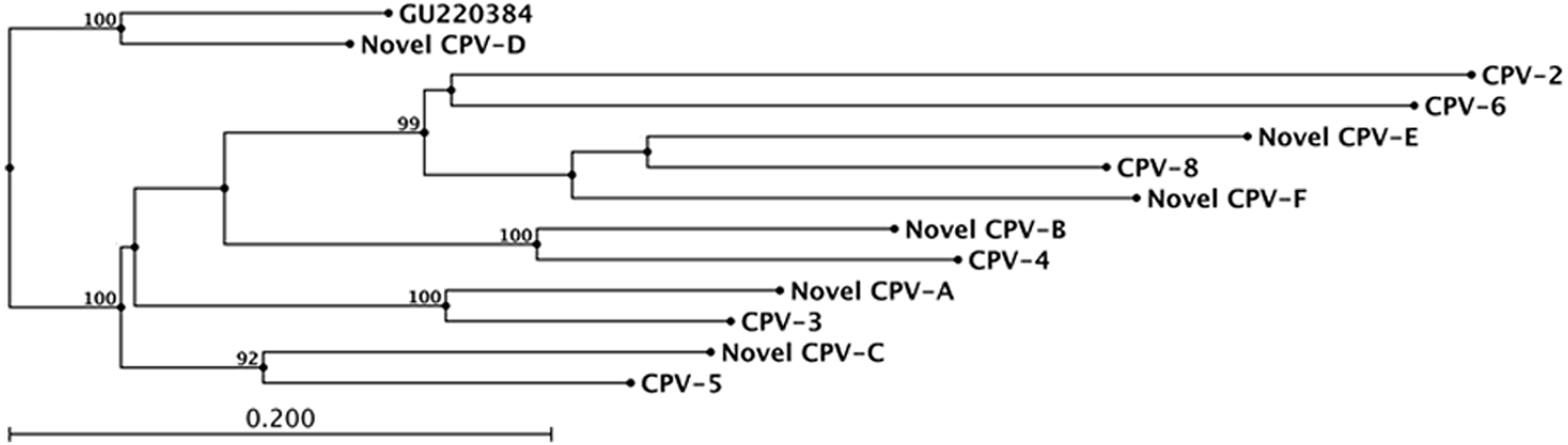
Neighbor-joining phylogenetic tree generated from an alignment of the nucleotide sequences (approximately 330 base pair segment) of the
CPV-8 DNA was not detected within any pigmented plaque in the current study. However, the degenerate primers used were able to successfully amplify DNA from the most closely related CPVs, including the novel PV sequences, and it is likely that these primers would have been able to amplify DNA from CPV-8. In addition, when the degenerate primers were aligned with the published nucleotide sequences for CPV-8, CPV-3, CPV-4, and CPV-5, there are similar numbers of mismatches at similar locations between each of the target sequences and the primers. Given that CPV-3, CPV-4, and CPV-5 were successfully amplified with these mismatches, it is likely that these same primers should be able to amplify CPV-8. However, a coinfection with CPV-8 cannot be completely excluded.
A causal association between the PV detected and plaque formation cannot be definitively proven because PV DNA can be detected on nonlesional skin as well. 8 However, the presence of abundant PV L1 protein identified within the proliferative epithelium using immunohistochemistry is suggestive for causality. In addition, PVs on nonlesional skin are maintained at low copy numbers (when compared to the large numbers of viral particles present in a productive PV infection in a papilloma or pigmented plaque), making is less likely that a rare PV associated with nonlesional skin would be amplified over the abundant PV DNA present within lesional skin from a single biopsy. 13 Furthermore, the study that detected PV DNA on normal skin extracted DNA from fresh samples, not FFPE biopsies, and the sensitivity for detection of a latent PV infection in formalin-fixed skin is unknown.
Classification of PVs into different genera, species, and types is generally based on the L1 nucleotide sequence identities, although interpretation is also based on genome organization and biology of the virus.1,2 The 8 known CPVs identified to date are divided into 3 distinct genera.
1
CPV-1 and CPV-6 are classified into the
In conclusion, canine pigmented plaques were associated with a large diversity of viruses in the current study, including CPV-3, CPV-4, CPV-5, 6 novel PV sequences not previously reported, and a seventh putatively novel CPV identified in a previous study.
10
These 6 novel PV sequences were most closely related to members of the
Footnotes
Acknowledgements
The authors are grateful for the cases provided by IDEXX Laboratories Inc. (West Sacramento, CA), Dr. John Peauroi at Veterinary Diagnostics (VDx) Veterinary Pathology Services (Davis, CA), Dr. Julie Yager at Yager-Best Histovet (Guelph, Ontario), and Dr. Emily Walder at an independent biopsy service (Venice, CA).
a.
Dako North America Inc., Carpinteria, CA.
b.
Vector Laboratories, Burlingame, CA.
c.
DNeasy tissue kit, Qiagen Inc., Valencia, CA.
d.
Cybr gold, Invitrogen Corp., Carlsbad, CA.
e.
Promega Wizard SV Gel and PCR Clean-Up System, Promega Corp., Madison, WI.
f.
DNA sequencing facility, University of California, Davis, CA.
g.
Vector NTI Software sequence analysis software, Invitrogen Corp., Carlsbad, CA.
h.
CLC Bio’s free CLC Work Bench, Germantown, MD.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding for this study was provided by the Center for Companion Animal Health, School of Veterinary Medicine, University of California, Davis, CA.
