Abstract
Three Holstein sires were identified as complex vertebral malformation (CVM) carriers using the polymerase chain reaction–primer introduced restriction analysis (PCR-PIRA) method. Using the carriers as positive controls, the PCR mismatch amplification mutation assay (MAMA PCR) method was developed and validated by sequence analysis. With MAMA PCR, 154 Chinese Holstein sires were tested for CVM, among which 24 were confirmed to be CVM carriers. With DNA isolated from hair follicles, 10 daughters of one CVM-positive bull were detected, among which 7 were confirmed to be CVM carriers.
Keywords
Complex vertebral malformation (CVM) is a bovine autosomal recessive disorder that was first reported and characterized by Danish scientists in 2000. 2 Since then, the occurrence of the disease and the identification of CVM carriers has been reported in other countries.4,6,10,15,18 Studies have revealed that the disorder is caused by a single-base mutation of guanine to thymine at position 73 in the fourth exon of the gene SLC35A.3,20 The homozygous mutation is lethal and leads to the abortion of most of the affected calves before gestation day 260.1,20 Fetal mortality causes a decrease in milk production and an increase in the return-to-service rate of cattle, which ultimately leads to the culling of Holstein cattle. Genetic investigations have revealed that the bull named Penstate Ivanhoe Star was the common ancestor of CVM carriers. 6 Star and its descendents have been used widely in the world due to their excellent performance, which increased the frequency of occurrence of CVM carriers in Holstein sires to 10–30% in several countries.8,13,20
Imported bulls and semen from several countries are used internationally for the breeding and artificial insemination of Holsteins, respectively, without assessment for the presence of CVM. Therefore, CVM carriers may exist in many countries, and the proportion of carriers in the Holstein population may be increasing silently and rapidly. Similar to other developing countries, a large number of latent CVM carriers were brought into China and used extensively for cattle breeding. Therefore, some elite bulls and a number of Holstein cows carry CVM alleles in the Chinese Holstein population.6,7,22,23 In China, the high abortion rate has always been a major limitation to the healthy development of the dairy industry. The abortion rates observed in Holstein cattle at some of the best dairy farms in China are approximately 12%, and have sometimes reached 20%, while the normal abortion rate is approximately 3–5%. Since some CVM carriers have been extensively used for breeding in China, CVM could be a critical factor involved in the high abortion rate. Moreover, this situation could also be occurring in other developing countries. Therefore, there is an urgent need for China and other countries to detect this lethal mutation and take preventive measures to control the disorder.
Several methods have been developed to detect the mutation that causes CVM. The polymerase chain reaction–primer introduced restriction analysis (PCR-PIRA) is a useful method for screening the CVM allele, 12 but it cannot distinguish the CVM allele from the wild-type allele directly by PCR amplification and requires further digestion analysis with specific restriction enzymes. The allele-specific PCR (AS-PCR) has an advantage that it can detect the CVM gene directly by PCR amplification. However, this method requires 2 amplification reactions for each sample. 11 The PCR–single strand conformation polymorphism (PCR-SSCP) is also a useful method for CVM screening, but it requires both agarose gel electrophoresis and polyacrylamide gel electrophoresis for genotype analysis. 6 In the current study, a simple and direct point mutation analyzing method was developed to detect the CVM defect.
In order to identify latent CVM carriers in China, the pedigrees of 730 Chinese Holstein bulls that had been extensively used in China were collected. Pedigree analysis revealed that 14 CVM carriers were sires of 27 Chinese bulls, 2 carriers were dams of 2 Chinese bulls, 3 carriers were grandsires of 4 Chinese bulls whose sires had not been tested, and 18 carriers were maternal grandsires of 32 Chinese bulls whose sires or grandsires were not CVM carriers or had not been tested. Frozen semen samples of 154 Chinese Holstein bulls and hair follicles of 10 daughters of a CVM-positive carrier were collected. Four bulls whose sires were CVM carriers were initially tested for CVM.
Genomic DNA was extracted according to the following procedure. For semen, 1 µl of semen was diluted and lysed with 0.1 of ml alkaline lysis solution (200 mM KOH/50 mM dithiothreitol). After incubation at 65°C for 20 min, 0.1 ml of neutralization solution (900 mM Tris-HCl [pH 8.3], 1 mM ethylenediamine tetra-acetic acid) was added to the lysate, and the supernatant containing template DNA was used for PCR amplification or stored at –20°C.9,24 For hair follicle, 2 or 4 follicles were lysed with 15 µl of alkaline lysis solution. After incubation at 65°C for 20 min, 15 µl of neutralization solution was added to the lysate, and the supernatant containing template DNA was used for PCR amplification or stored at –20°C.
The PCR-PIRA was used to identify CVM carriers with some modification. 12 The modified primers were named T primer pairs (5′-TCACAATTTGTAGGTCTCAATGCA-3′ and 5′-TCCACACTGATGGTTTGGTTTCTT-3′). a By PCR amplification with T primer pairs, a specific 106-bp band was obtained from all of the samples. After digestion with Nsil, b an 83-bp fragment and 106-bp fragment were observed in samples 2–4, while only a 106-bp fragment was observed in sample 1. These results indicated that sires 2–4 were CVM carriers, and sire 1 was CVM negative (Fig. 1).
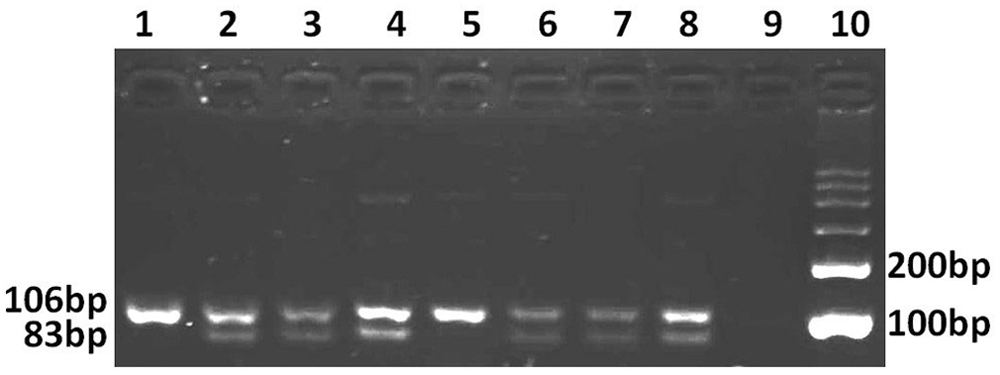
Analysis of complex vertebral malformation allele with the polymerase chain reaction (PCR)–primer introduced restriction analysis method. Lanes 1–8: PCR products with the T primer pair. The products were digested with Nsil (lanes 1, 5: sire 1; lanes 2, 6: sire 2; lanes 3, 7: sire 3; lanes 4, 8: sire 4); lane 9: negative control; lane 10: 100-bp marker. c
The PCR mismatch amplification mutation assay (MAMA PCR) is a reliable method for analyzing point mutants in DNA.5,14,16 The technique is based on the theory that the selective amplification of a gene with a primer containing a mismatch at the 3′ end has little effect on the extension of the primer by Taq polymerase and additional mismatches can enhance the specificity of amplification. 17 Studies have shown that a single T:C (primer/template) mismatch at the 3′ extremity of a primer has minimal effect on PCR product yield, and 2 consecutive mismatches located at the fifth and sixth residues from the 3′ end of a primer can also be efficiently extended by lowering the annealing temperature.14,17 However, the T:C mismatch at the 3′ end together with the 2 consecutive mismatches located at the fifth and sixth residues from the 3′ end may effectively restrain PCR amplification. Therefore, in order to specifically amplify the CVM allele, the 3′ end of the thymine-specific (TS) forward primer was designed to include the disease-causing mutation, with a T as the terminal base and an alteration of the fifth and sixth residue from the 3′ end (Table 1). Bovine-specific (BS) primers (5′-TGGAAGCAAAGAACCCCGCT-3′ and 5′- TCGTCAGAAACCGCACACTG-3′) a were designed according to the sequence of the bovine 1.715 satellite and used as an internal control (Bos taurus; V00124). Exon-specific (ES) primers (5′-TGTTTCTCTTTGTCCAGTG-3′ and 5′-CGAC CAAGTTGAATGTTTC-3′) a were designed for sequencing according to the sequence of the fourth exon of SLC35A3 (B. taurus; AY160683).
Illustration of the TS forward primer and corresponding template sequence of the primer.*
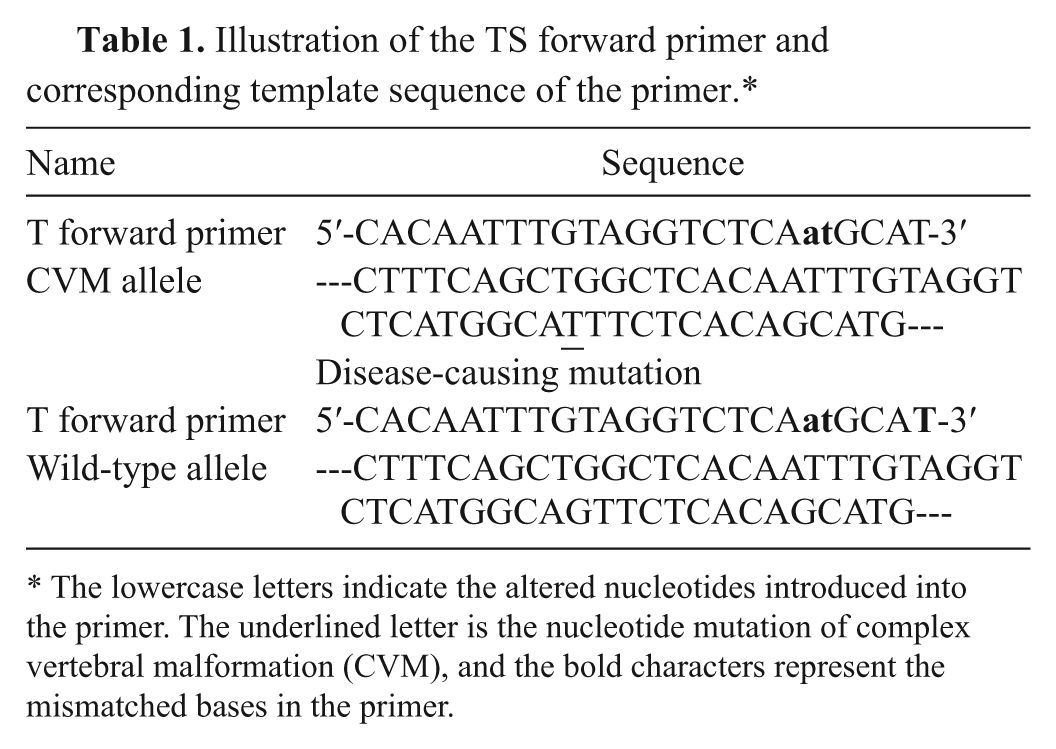
The lowercase letters indicate the altered nucleotides introduced into the primer. The underlined letter is the nucleotide mutation of complex vertebral malformation (CVM), and the bold characters represent the mismatched bases in the primer.
The MAMA PCR amplification was performed in a total volume of 20 µl containing 6.5 µl of double distilled water, 3 µl of 1 mM deoxyribonucleotide triphosphate, c 1.5 µl of 10× PCR buffer (500 mmol/l KCl, 100 mmol/l Tris-HCl, 15 mmol/l MgCl2, 0.1% gelatin), c 1.2 µl of 20 mM Mg2+, 1.2 µl of 2 µM BS primers, 1.8 µl of 2 µM TS primers (5′-CACAATTTGTAGGTCTCAATGCAT-3′ and 5′-TCCA CACTGATGGTTTGGTTTCTT-3′), a 0.8 µl of Taq polymerase (2.5 U/µl), c and 4 µl of genomic DNA. Samples were amplified for 31 cycles using the following PCR protocol: denaturation at 94°C for 10 sec, annealing at 51°C for 10 sec, and extension at 72°C for 10 sec. The protocol works well for mismatch PCR amplification when the rate of temperature change of the thermocycler is below 5°C/sec. The PCR products were detected on a stained 2% agarose gel. d
The MAMA PCR reaction efficiently detected the disease-causing mutation. The BS primers were used as an internal control of PCR amplification. The amplification of a 216-bp band indicated that the corresponding reactions were valid. In carrier samples, the 105-bp and 216-bp bands were both present. However, in samples that were CVM negative, only the 216-bp band was amplified. Using the MAMA PCR method, sires 2–4 were identified as CVM carriers, and sire 1 was CVM negative (Fig. 2). The data were completely consistent with the results of the PCR-PIRA experiment.
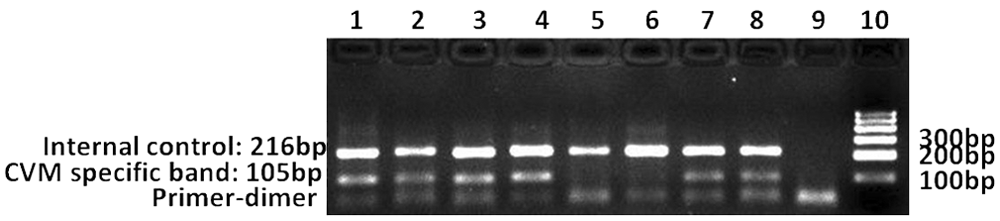
The polymerase chain reaction (PCR) mismatch amplification mutation assay reaction for diagnosis of wild-type and carrier bulls for complex vertebral malformation (CVM). Lanes 1–8: PCR products (lanes 1, 2: sire 2; lanes 3, 4: sire 3; lanes 5, 6: sire 1; lanes 7, 8: sire 4); lane 9: negative control (without template DNA in PCR reaction system); lane 10: 100-bp marker. c
Using the ES primer pair, PCR products of 192 bp were sequenced to confirm the test results from the MAMA PCR. The sequencing data revealed that sires 2–4 were heterozygous for T and G, while sire 1 was homozygous for G, which was consistent with the results obtained with the MAMA PCR and the PCR-PIRA methods.
All 154 bulls were tested for CVM allele with MAMA PCR, which confirmed 24 bulls were CVM carriers. The frequency of heterozygotes was 15.58%. Pedigree analysis revealed that all the 24 positive CVM carriers were descendants of Star, the common ancestor of all CVM carriers, which further confirmed the accuracy of test results. The analysis results also show that the bull Eng-Amer Ivanhoe Jerry (1626173), which was the son of Star and the common ancestor of 8 positive CVM carriers, should be a CVM carrier.
When genomic DNA isolated from hair follicles were used to test cows for CVM, it was revealed that for cow 1 (CVM carrier), the more hair follicles used for DNA isolation the more efficiently the TS primers extended (Fig. 3). However, for cow 2 (CVM free), a very weak band of approximately 105 bp was amplified from DNA isolated from 8 hair follicles (Fig. 3). The amplification results showed that genomic DNA isolated from 4 hair follicles was suitable for testing the CVM allele with MAMA PCR. Subsequently, 10 female offspring of 1 CVM-positive carrier were tested, among which, 7 cows were confirmed to be CVM carriers.
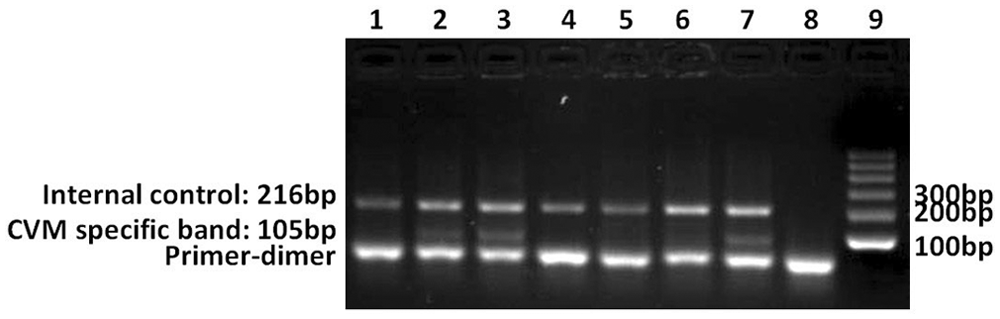
The amplification results from DNA isolated from hair follicles. Lane 1: 2 hair follicles of cow 1; lane 2: 4 hair follicles of cow 1; lane 3: 8 hair follicles of cow 1; lane 4: 2 hair follicles of cow 2; lane 5: 4 hair follicles of cow 2; lane 6: 8 hair follicles of cow 2; lane 7: positive control (sire 4: a complex vertebral malformation [CVM] carrier); lane 8: negative control; lane 9: 100-bp marker.
In the present study, 2 consecutive mismatches at the fifth and sixth bases from the 3′ end of the T forward primer were introduced to enhance the specificity of amplification from the CVM allele. Because of the mismatches between primer and template, the DNA polymerase used should lack high proofreading activity and the annealing temperature should be lower than regular PCR amplification. The amplification reaction is specific at annealing temperatures between 49°C and 51°C. The internal control is introduced to indicate the validity of PCR amplification, but it does not prevent false-negative results if too little template DNA is used. In order to enhance the reliability of MAMA PCR for CVM detection, the concentration of DNA template should be optimized.
The MAMA PCR method has been used widely in analyzing point mutations in DNA5,14,16,17 and has been shown to be simple, rapid, reliable, and economical. When compared to existing detection methods for CVM, the MAMA PCR method developed in the current study has 2 advantages. First, the MAMA PCR can directly identify the CVM gene in 1 amplification reaction and does not require an additional digestion reaction. Second, the protocol for both the DNA extraction and the PCR amplification developed is easier to perform than other methods for testing the CVM allele.12,21
If a lethal recessive mutation is not associated with any economically important trait, it would be much easier to eliminate the allele in the sires. However, the CVM mutation is thought to be linked to milk quality and quantity, which is a major economic factor in dairy cows. 19 Therefore, control of the disease allele might be preferable to its elimination. To control the disease gene, extensive screening is necessary for detection of carriers, especially cows. 12 The MAMA PCR method developed in the current study appears to be a reliable and rapid method for the detection of CVM in Holstein cattle.
Footnotes
a.
Invitrogen (Beijing) Ltd., Beijing, China.
b.
New England Biolabs (Beijing) Ltd., Beijing, China.
c.
Dongsheng Biotech Co. Ltd., Guangzhou, China.
d.
GOLDViewna™ I/II, Beijing Nucleic-Protein Bio-tech Co. Ltd., Beijing, China.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
This work was supported by China Agriculture Research System (CARS-37) and the Genetically Modified Organisms Breeding Major Projects of China (2009ZX08011-031B, 2009ZX08010-015B).
