Abstract
Complex vertebral malformation (CVM) is a monogenic autosomal recessive hereditary defect of Holstein dairy cattle. It is caused by a point mutation from G to T at the nucleotide position 559 in bovine solute carrier family 35, member 3 gene (SLC35A3), which changes the amino acid sequence of uridine 5′-diphosphate-N-acetylglucosamine transporter protein from a valine to a phenylalanine in position 180. The elite U.S. Holstein sire Penstate Ivanhoe Star was identified as the common ancestor of the current CVM carriers. Because his offspring, mainly those of Carlin-M Ivanhoe Bell, were used in many countries, CVM has potentially spread into China. In the present study, using the polymerase chain reaction-single-stranded conformational polymorphism (PCR-SSCP) technique, 10 CVM carriers were found among 68 at-risk Chinese Holstein bulls, and 282 carriers were found among 602 at-risk cows. The results of this study indicate that the CVM gene exists in the Chinese Holstein population.
Keywords
Complex vertebral malformation (CVM), a recessively inherited disorder in Holstein calves, was first reported in Denmark in 2001. Shortly thereafter, reports documented the presence of CVM in the United States, 3 the United Kingdom, 5 The Netherlands, 7 and Japan. 4 The affected calves are characterized by complex anomalies, including proportional dwarfism, symmetrical arthrogryposis of anterior and posterior limbs, and multiple malformations of the cervical and thoracic parts of the vertebral column. At the same time, the fertility traits of cows, such as nonreturn rate, calving interval, and aborting proportion, are significantly affected by CVM. Because approximately half of the fetuses with CVM are aborted 100 to 260 days after conception, cows cannot maintain a high yield until the following calving. These problems bring severe economic losses to farmers. A previous study revealed that the genetic mechanism of CVM is the single base mutation from G to T at the nucleotide position 559 of the bovine solute carrier family 35, member 3 gene (SLC35A3), which alters the amino acid sequence of uridine 5′-diphosphate-N-acetylglucosamine transporter protein from a valine to a phenylalanine in position 180. 6
It has been reported that Penstate Ivanhoe Star was the common ancestor and that CVM was spread throughout the world mainly by his son, Carlin-M Ivanhoe Bell. Because Bell and his offspring were used in many countries, including China, it was presumed that CVM would occur in China. According to the list of CVM carrier sires published by the United States and Canada, as well as the Chinese Holstein sire pedigrees, 68 sires from Northern China were considered at-risk sires (i.e., potential CVM carriers). The aim of the present study was to detect the distribution of CVM in Chinese Holstein cattle.
Frozen semen samples of the 68 at-risk Chinese Holstein bulls were collected from the Beijing Dairy Center, the Tianjin Dairy Cattle Development Center, and the Hebei Breeding Bull Station, which are the main distribution centers for Chinese Holstein bulls and cows in Northern China. Genomic DNA (gDNA) was isolated according to a procedure previously described 2 and was adjusted to a final concentration of 200 ng/μl.
Oligo Primer Analysis Software Version 6.41 a was used to design a pair of primers around the mutation of site 559 according to the genomic DNA sequence of SLC35A3 in GenBank (accession number: AY160683). The forward and reverse primer sequences were 5′-AGCTCTCCTCTG TAATCC-3′ and 5′-TCTCAAAGTAAACCCCAG-3′. Polymerase chain reaction (PCR) was performed in a final volume of 20 μl containing 100 ng gDNA and 1 μM each of the following primers: 2.5 mM MgCl2, 200 μM dNTPs, and 1U Taq polymerase. b The mixture was incubated at 95°C for 5 min followed by 35 cycles of 94°C for 30 sec, 58 °C for 30 sec, and 72°C for 30 sec, with a final extension at 72° C for 7 min. Three microliters of PCR products and 7 μl of loading buffer were mixed and denatured at 98 °C for 10 min, followed by a rapid chill on ice for 10 min, then electrophoresed on 14% Polyacrylamide gel and detected by silver staining. One PCR product for each genotype was excised from 1.0% agarose gel and sequenced on the ABI 3730XL DNA Sequencer using the BigDye Terminator Cycle Sequencing Kit. c
By PCR amplification with the pair of primers, a specific 249-bp fragment of SLC35A3 was obtained. Partial results of PCR products by 2% agarose gel electrophoresis are shown in Figure 1. With single-stranded conformational polymorphism (SSCP) analysis, 2 types of band patterns were found corresponding to 2 kinds of genotype, named AA and AB in this study, respectively (Fig. 2). Two samples were randomly selected from each genotype and sequenced. Figure 3 gives the sequences around the mutation position, indicating both G and T bases for genotype AB, with 1 G base for genotype AA at position 559. This is consistent with a previous study. 6
All 68 at-risk individuals were tested with the above method, which confirmed that 10 bulls were carriers of recessive abnormalities and the remaining 58 were not. The frequency of heterozygotes was 15%. There are approximately 290 sires in the regions of Beijing, Tianjin, and Hebei, which indicates that the average percentage of CVM carrier sires is 3.5% (3.5 of 100 sires). Subsequently, 602 female offspring of the 10 confirmed bulls were selected randomly from the 3 regions; by PCR-SSCP detection, 282 of the 602 cows were carriers.
This study identified the existence of CVM allele in Chinese Holstein. According to the pedigree analysis of bulls, the proportion of CVM carriers should be 20% (13 of 68 sires). This number is approximately the same as the percentage (10/68 or 14.7%) detected in the present study. These 68 at-risk sires are the top sires in China and have been widely used throughout the country. Meanwhile, many female offspring of the 10 sires are in the same herds. The abortion rate is >13% in many dairy farms in Beijing compared with 2% to 3% in other countries. Perhaps the high abortion rate is partly the result of this hereditary disease.
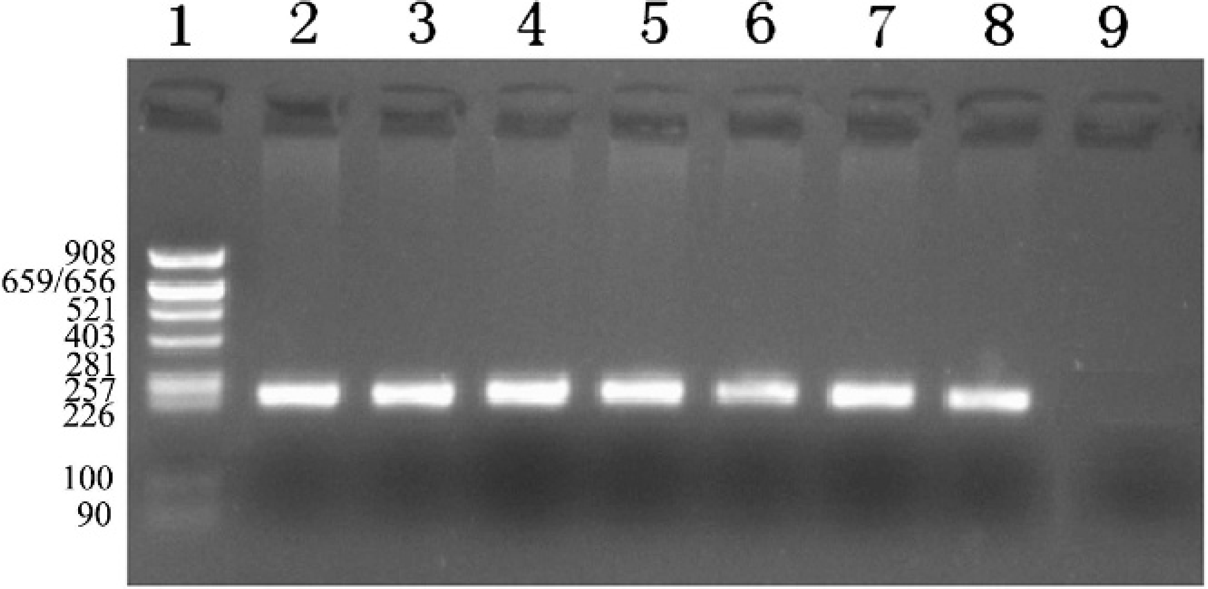
Polymerase chain reaction (PCR) products of SLC35A3 in Chinese Holstein cattle. Lane 1: pBR322AluI marker; lanes 2–8: PCR products; lane 9: negative control (without template DNA in the PCR reaction system).
The authors believe that it is necessary to have all top bulls in China screened, especially those that have CVM carriers in their pedigrees. Subsequently, all suspicious cows will also need to be examined. Because it is not economically beneficial to cull carrier bulls, the pedigree of the cows will need to be examined to avoid using carrier bulls on those cows whose pedigree suggests that they might be carriers. This will help to reduce the negative influence on breeding and prevent the occurrence of malformed calves.
Acknowledgements. This work was supported by the State Major Basic Research and Development Project of China (2006CB102107), National Key Technologies R&D Program (2006BAD04A01), National High-tech R&D Program (2007AA10Z157), and National “948” Project (2006–G48).
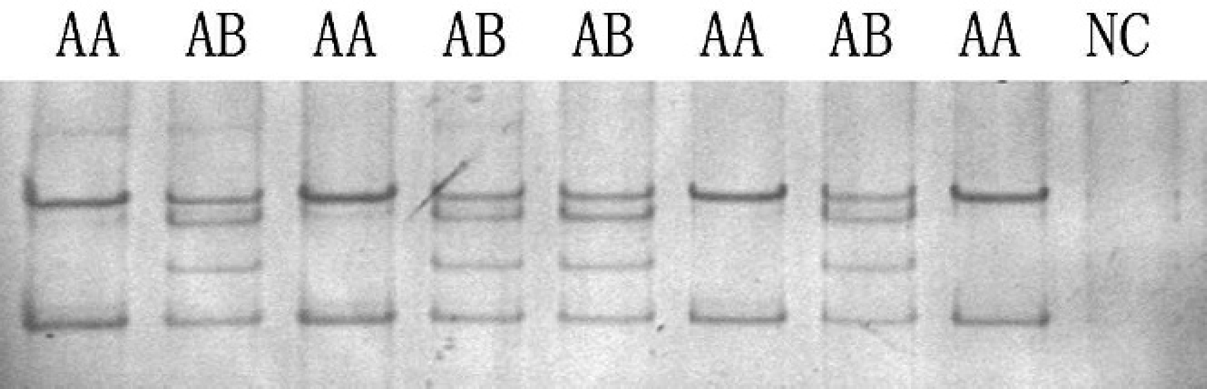
Polymerase chain reaction-single-stranded conformational polymorphism (PCR-SSCP) result of SLC35A3 in Chinese Holstein dairy bulls. NC = negative control (without template DNA in the PCR reaction system).
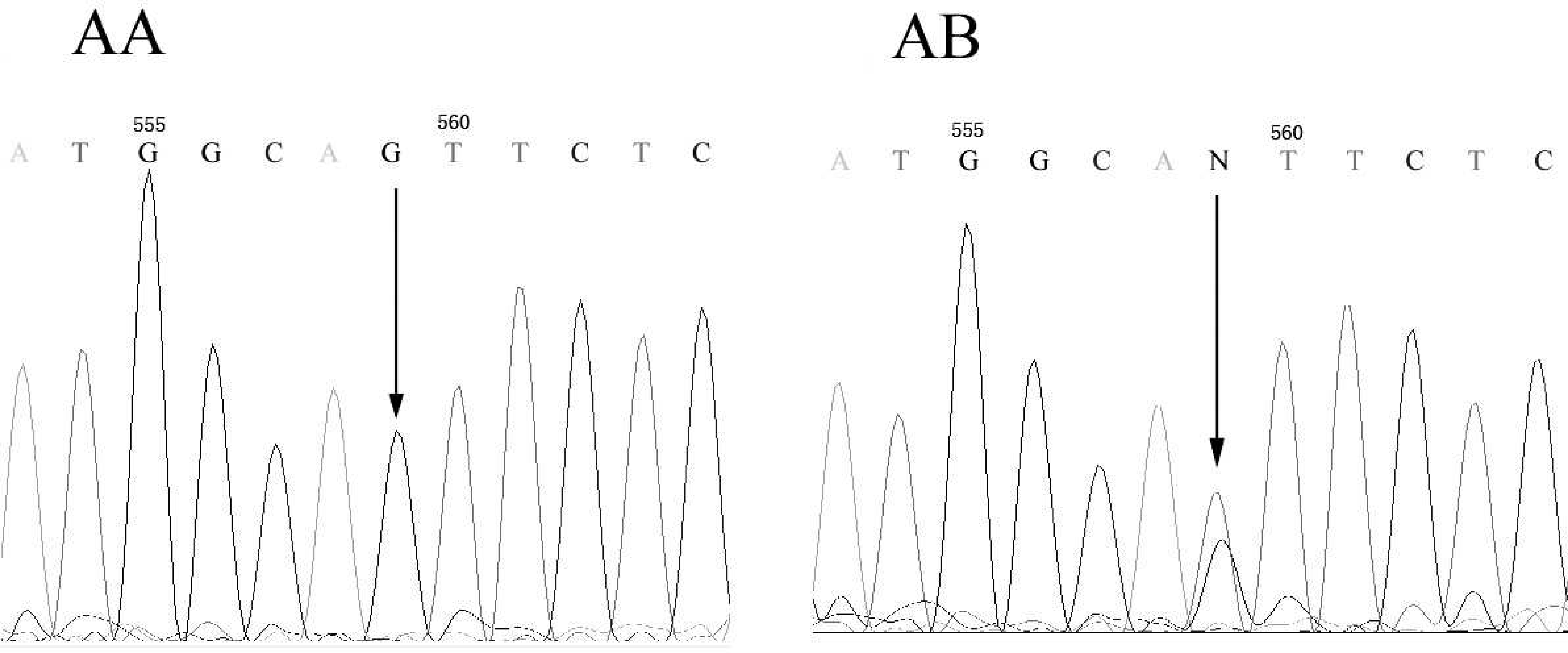
Partial sequences of SLC35A3 gene. Arrows indicate the polymorphic site in the genotypes AA and AB.
Footnotes
a.
Molecular Biology Insights, Cascade, CO.
b.
Tiangen Biotech (Beijing) Co. Ltd., Beijing, China.
c.
Applied Biosystems, Foster City, CA.
