Abstract
Pseudocowpox virus is a parapoxvirus frequently associated with papulovesicular and scabby lesions on the teats and udders of milking cows and is often transmitted to human beings. An unusual outbreak of skin disease in fattening calves in southern Brazil is described. Fourteen of 17 male cattle (82%), aged 6–48 months, feeding on grass pastures were affected. Animals developed papules, vesicles, and scabby proliferative lesions on the muzzle in a clinical course of approximately 10–15 days. The scabby lesions often presented with exudation and bleeding. Histological examination of mucocutaneous tissue in detached scabs revealed acanthosis with thickening of the corneal layer and premature keratinization (parakeratotic hyperkeratosis). The dermis had multifocal lymphoplasmacytic infiltrates. Electron microscopic examination of scab specimens revealed typical parapoxvirus particles: oval shaped (260 nm × 160 nm), enveloped, and covered with a helical layer. Polymerase chain reaction using a set of pan-parapoxvirus primers for the B2L gene amplified a 590-bp product out of DNA extracted from scabs. Nucleotide sequencing of the amplicons revealed a nucleotide homology of 97% with Pseudocowpox virus and lower homology with other parapoxviruses: Bovine papular stomatitis virus (84%) and Orf virus (94%). A phylogenetic tree based on the B2L sequence was constructed, showing that the virus clustered with Pseudocowpox virus isolates.
Pseudocowpox virus (PCPV, previously known as Parapoxvirus bovis 2) is one of 2 parapoxviruses (PPVs) of cattle, along with Bovine papular stomatitis virus (BPSV, previously known as Parapoxvirus bovis 1). Two other virus species, namely Orf virus of sheep and goats (ORFV, previously known as Parapoxvirus ovis) and Parapoxvirus of red deer in New Zealand (NZPV), complete the genus Parapoxvirus within the subfamily Chordopoxvirinae of the family Poxviridae. 29 Parapoxviruses are epitheliotropic viruses identified throughout the world as causing nonsystemic, vesicular, and eruptive skin disease in domestic and wild mammals, especially ruminants. 11 Individual PPV species usually display a narrow host range yet are occasionally transmitted to human beings, causing localized lesions on the hands.2,17 Human PPV infection is generally occupational, affecting milkers or other personnel in contact with affected animals, causing lesions called “milker’s nodule” or “pseudocowpox.”6,11
Information on PCPV infection in cattle is very scarce in the literature, possibly due to the mild presentation of the disease and the clinical similarity with BPSV and ORFV.6,20 In addition, many reports are restricted to clinical descriptions, electron microscopy (EM), and/or serology, without definitive agent identification.14,23 Pseudocowpox virus infection is distributed worldwide and affects mostly milking cows but presents limited sanitary and economic importance in most countries. In some dairies, however, the infection may assume some economic importance due to poor hygiene conditions. The agent is usually introduced to herds through infected animals and disseminates slowly among the animals. Transmission within herds occurs by direct and indirect contact. Indirect routes include calf suckling of multiple cows, flies, milking equipment, and inadequate milking/management procedures. 20 Parapoxvirus infection occurs through scarified or damaged skin (or occasionally through the oral mucosa), followed by virus replication in keratinocytes.6,18 Virus replication near the port of entry is accompanied by a well-characterized clinical course that progresses through the stages of macules, papules, vesicles, pustules, and scabs. 18 Vesicles and pustules covered by scabs are also frequently observed. In the absence of bacterial contamination or other complications (e.g., myiasis, mechanic trauma), scabs usually detach after a few days, and lesions rapidly heal. Similar lesions may occur on the muzzle of calves suckling affected cows. 20 On the other hand, BPSV infects cattle of all ages, and lesions are usually seen on the muzzle of calves 9 and less frequently on the teats and udders of cows. 6
Exanthematic and vesicular disease in dairy cattle, occasionally affecting human beings, has been frequently reported in southeast Brazil in previous decades, assuming some importance in animal and human health in some rural communities.7,10,24 Most reported cases and outbreaks have been associated with infection by Vaccinia virus (VACV)4,7,15,25,28 of the genus Orthopoxvirus. The origin of these Brazilian VACVs is still a matter of debate, and both native and smallpox vaccine–derived origins have been proposed.2,27 In addition, cases of vesicular lesions in the teats and udders of dairy cows, empirically called “bovine variola” or cowpox, without laboratory diagnosis, are commonly reported by veterinarians all over the country. On 2 occasions, the presence of PPVs was demonstrated. Parapoxvirus particles were demonstrated by EM in scabs obtained from teat lesions in dairy cows in an outbreak of cattle disease, with human involvement, in the state of Rio de Janeiro. 23 Definitive virus identification, however, was not performed. A human case of co-infection by VACV and PCPV was described in a patient with pustular lesions on the hands, probably acquired by milking affected cows. 5 An outbreak of eruptive skin disease in dairy cows in midwestern Brazil, with human cases, was diagnosed and attributed to BPSV infection through identification and sequencing of viral genes on material present on scabs. 22
The present article describes an outbreak of PCPV infection on a small farm in southern Brazil, with unusual clinical and epidemiological features. Severe vesicular, papulopustular, and proliferative scabby lesions were observed on the muzzle of a group of 14 fattening, crossbred calves with no apparent link with dairy herds, the usual source of this virus. Parapoxvirus particles were visualized by EM on scabs, and PPV sequences were amplified by polymerase chain reaction (PCR). Nucleotide sequencing of the amplicon demonstrated a high homology with PCPV sequences deposited in GenBank.
The disease affected the calves in June 2011 on a small fattening farm in Pinhal Grande County, Rio Grande do Sul state, Brazil. The farm held a total of 17 crossbred male cattle, 6–48 months old, and 14 animals developed lesions. Calves with lesions were in a group maintained on semi-extensive grass pasture. Three calves that were in another group remained healthy. All animals were acquired from other farms soon after birth and introduced to the group at different times. The interval between the acquisition of the last animal and the onset of clinical signs was approximately 4 months. The affected calves developed clinical signs within approximately 2 weeks. The main clinical signs were scabby proliferative lesions that oozed and bled frequently, found around the muzzle and along the mucocutaneous junction (Fig. 1). The lesions started as macules and papules that evolved to vesicles, pustules, and scabs, with a clinical course of approximately 10–15 days. Lesions spontaneously resolved after a few days. Two calves presented with mucopurulent nasal discharge, probably due to secondary bacterial infection. Animals did not show systemic signs. Likewise, no human cases were reported. The farm owner, a veterinarian, reported that he had never seen such a clinical presentation in more than 2 decades of veterinary practice.
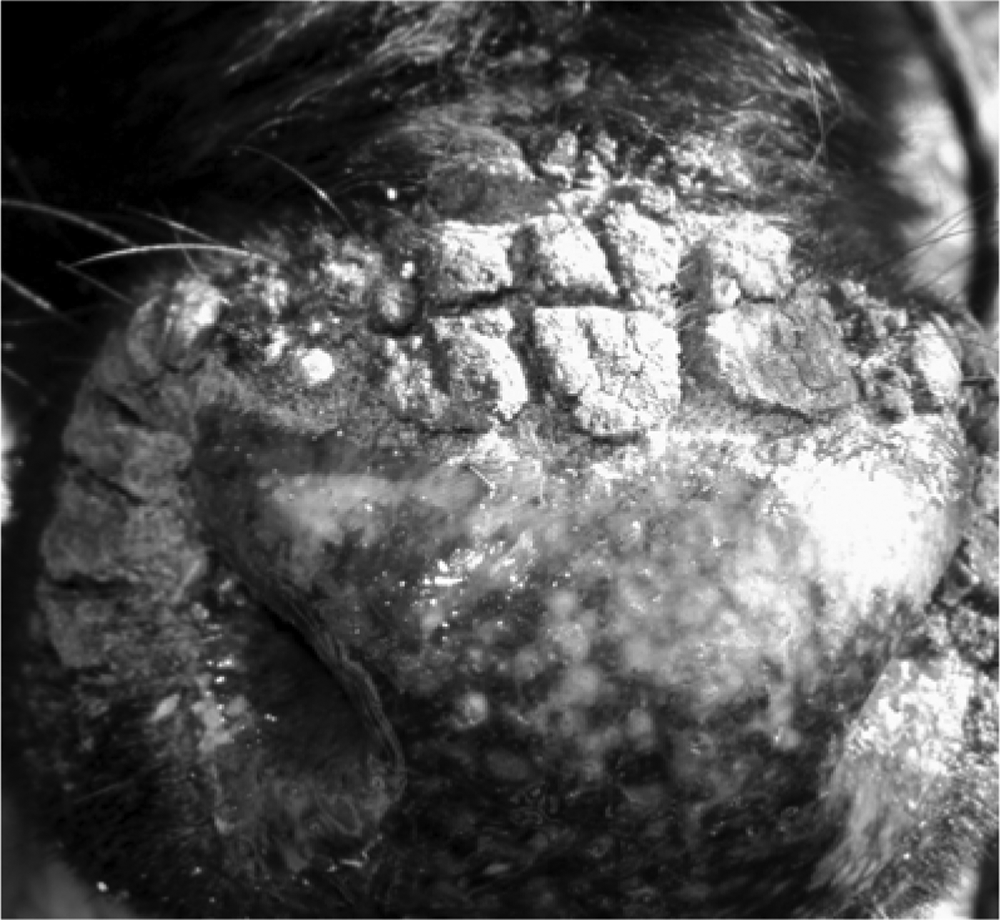

Calf muzzle exhibiting papules and vesicles, proliferative scabby lesions around the muzzle, and mucocutaneous junctions.
Detached scabs from 2 calves were collected for laboratory examination (histopathology, EM, virus isolation, and PCR). Samples of fresh tissue were fixed in 10% buffered formalin, processed for routine histopathological examination, and stained with hematoxylin and eosin. Scabs from 2 calves were processed for negative-staining EM. For virus isolation, scabs collected from these calves were homogenized, resuspended to 10% (weight-volume) in culture medium, and inoculated onto monolayers of Madin–Darby bovine kidney and Vero (African green monkey kidney epithelial) cells. The cultures were monitored for cytopathic effect during 3 passages of 5 days each. Total DNA was extracted from scabs of 2 animals using DNAzol reagent.a Initially, DNA was examined by PCR for VACV using previously described primers. 1 No VACV sequences were detected in the scabs. Next, DNA was examined using a PCR designed to detect the B2L gene of PPVs with the following primers: PPP1 forward (5′-3′GTCGTCCACGATGAGCAG) and PPP4 reverse (5′-3′TACGTGGGAAGCGCCTCGCT). 13 Total DNA extracted from baby hamster kidney (BHK)–21 cells infected with the ORFV strain IA-82 9 was used as a positive control. 3 As the set of primers was expected to amplify all PPVs, 13 the amplicons were sequenced to identify the specific PPV. The PCR products were purified using a commercial kita according to the manufacturer’s instructions and sequenced.b The obtained sequences were analyzed by the Staden package, 26 aligned using BioEdit Sequence Alignment Editor software (version 7.0.5.3), 12 and compared with other PPV sequences deposited in GenBank. The sequence was then deposited in GenBank as SV285/11 (accession no. JN638992).
Pathological examination of material from scabs showed the superficial epidermis covered by thick serocellular crusts. Squamous keratinocytes were hyperplastic with thickening of the corneal layer and premature keratinization (parakeratotic hyperkeratosis). The epidermis was acanthotic, and multifocal lymphoplasmacytic infiltrates were observed (Fig. 2).
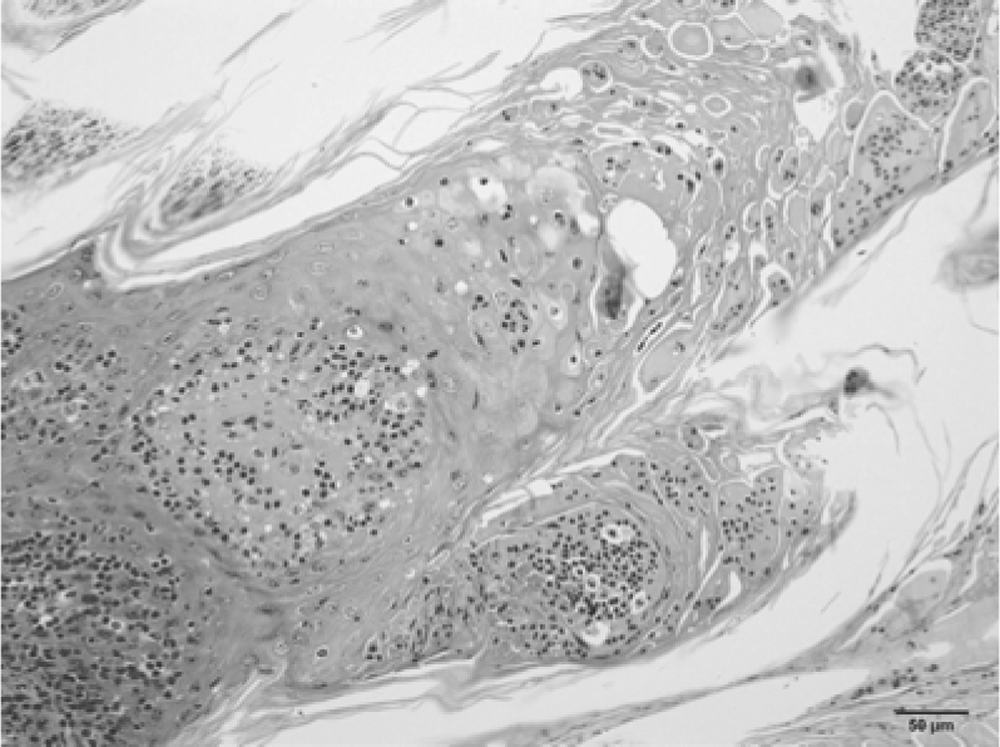
Calf; skin and muzzle. Severe diffuse proliferative (acanthosis) and necrotizing dermatitis with moderate lymphoplasmacytic infiltration. Note ballooning degeneration of superficial keratinocytes. Bar = 50 µm.
Electron microscopic examination of scabs revealed typical PPV particles: oval shaped (260 nm × 160 nm), enveloped, and with a helical structure (Fig. 3). However, the samples were negative for infectious virus, as ascertained by the absence of cytopathic effect in inoculated monolayers. The failure to isolate the virus may be related to inappropriate handling of clinical samples, resulting in virus inactivation, or to relative insensitivity of the cell line used.
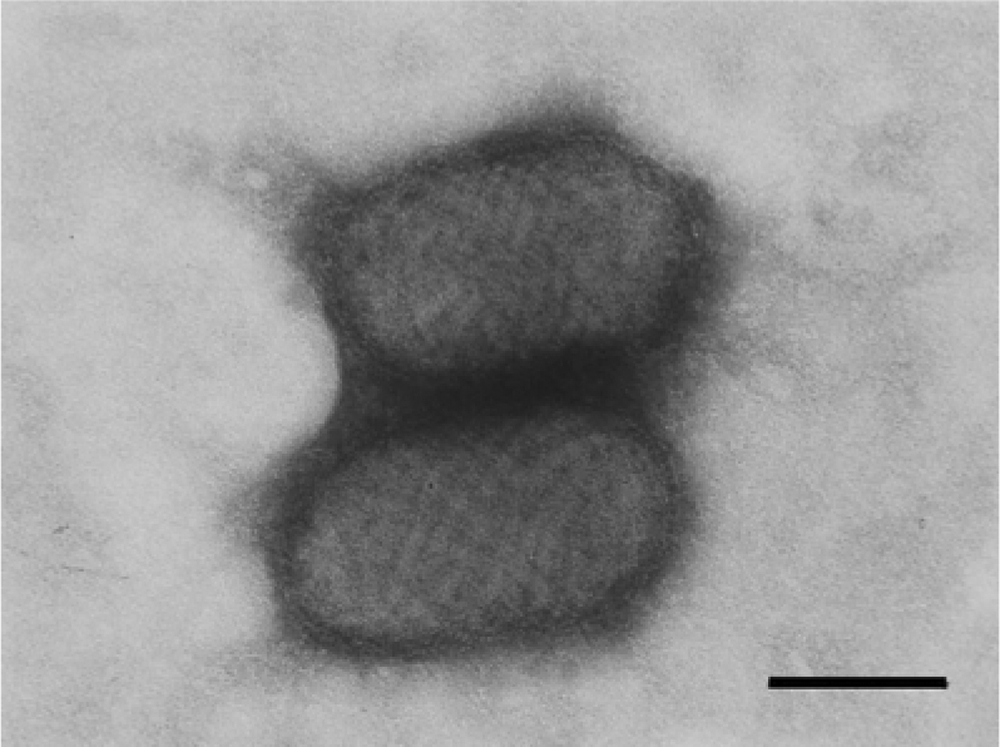
Calf. Parapoxvirus particles from scab samples processed for negative staining for electron microscopy. Oval-shaped (260 nm × 160 nm) particles, enveloped, with a helical structure typical of the Parapoxvirus genus.
In a PCR for the B2L gene of PPVs, a band of approximately 590 bp, corresponding to the expected size of the amplicon, was amplified from DNA extracted from scabs of 2 animals. Nucleotide sequencing of the amplicons and sequence alignment showed that the amplified region of the B2L gene of SV285/11 had a nucleotide homology of 97% with PCPV and lower homology with BPSV (84%) and ORFV (94%). In general, the nucleotide differences between SV285/11 and the other PPV sequences were scattered throughout the sequenced region (Fig. 4). A phylogenetic tree based on the B2L sequence was constructed using the maximum parsimony method with 1,000 bootstrap replicates implemented by MEGA3.1, as previously described.1,5 The tree is depicted in Figure 5 and shows SV285/11 clustered together with several PCPV isolates.
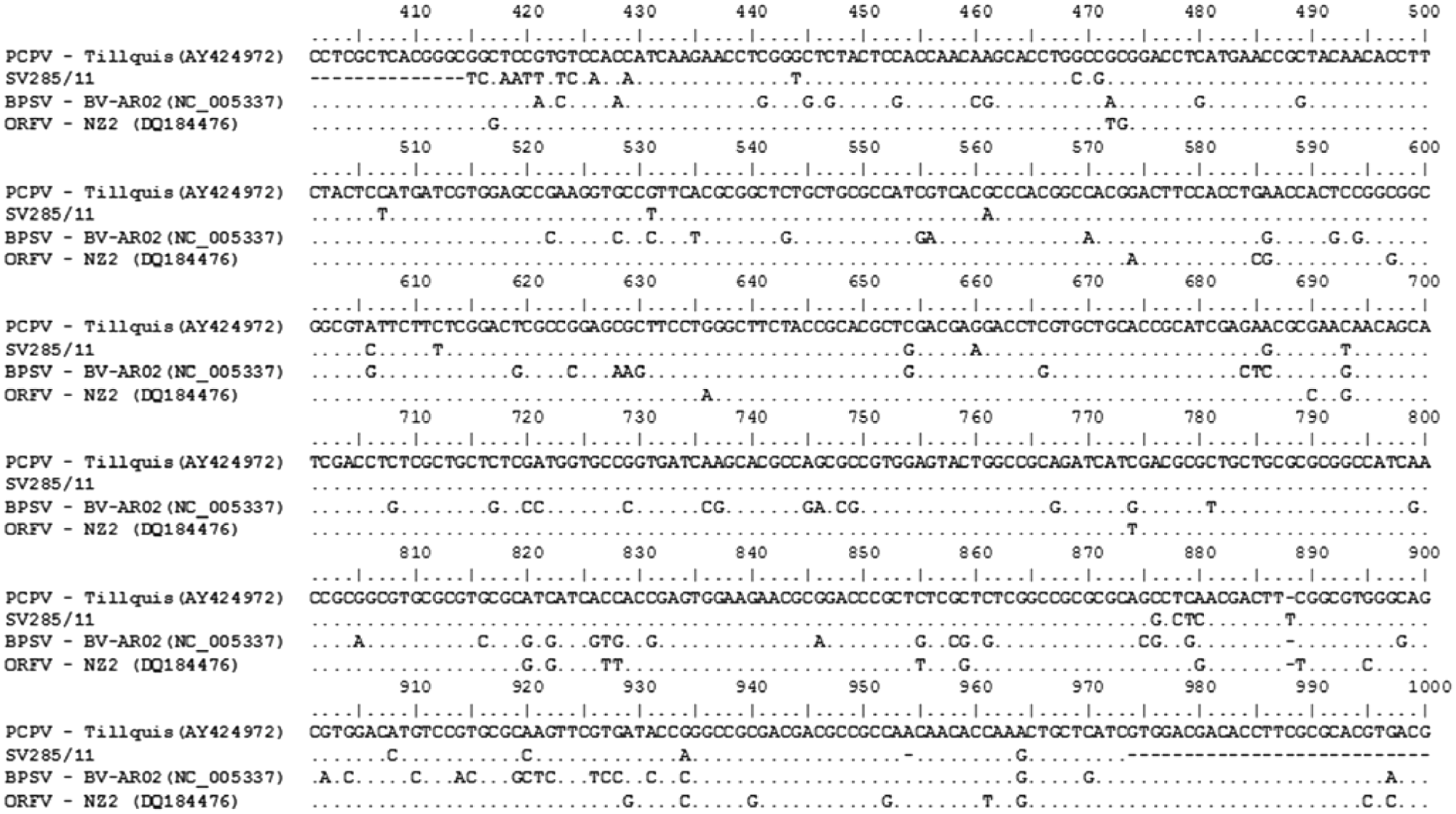

Alignment of nucleotide sequences of a segment of B2L gene of parapoxvirus and sample SV285/11 of Pseudocowpox virus (PCPV). Sequences of PCPV, Bovine popular stomatitis virus (BPSV), and Orf virus (ORFV) were retrieved from GenBank and were aligned using the BioEdit program. The strain PCPV (accession no. GQ329670) was used as a reference sequence.
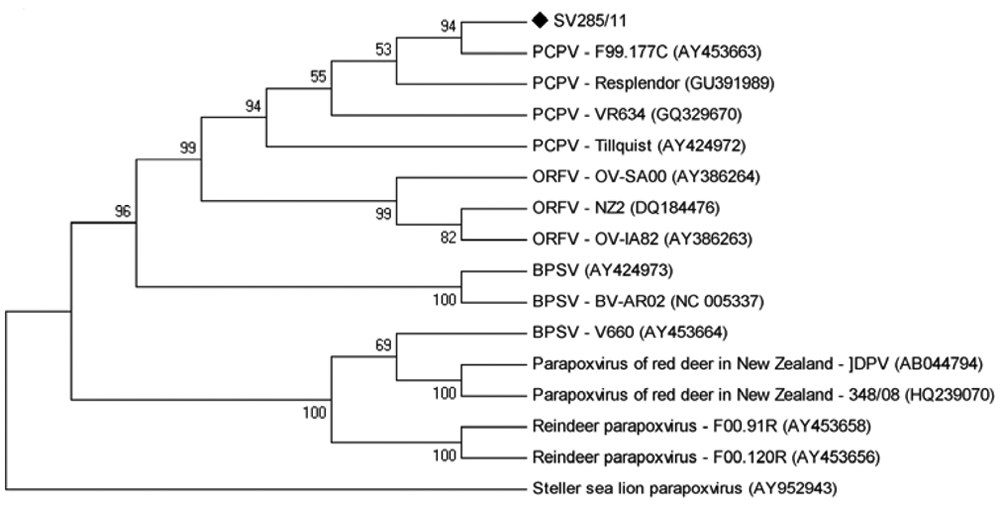
Phylogenetic tree based on the nucleotide sequences of the parapoxvirus B2L gene. The tree was constructed using the maximum parsimony method with 1,000 bootstrap replicates implemented by MEGA5.03. Values >50% are shown. Pseudocowpox virus SV285/11 sample is identified with a black marker.
The morbidity in the described outbreak was 82% (14/17 calves). The origin of the virus is unknown, as no similar disease has been noticed in this herd or in nearby farms. The main form of introduction of PCPV in herds appears to be the purchase/acquisition of infected animals. 20 However, the long time interval between the introduction of the last animal into the herd and the outbreak, approximately 4 months, makes this route unlikely. The high morbidity and quick spread of the virus within the herd (a point epidemic curve) are unusual and suggest a recent introduction of the virus.
Pseudocowpox virus was originally isolated from lesions on teats of dairy cows and from the mouth of their suckling calves. 19 Similar scabby lesions in cattle can be caused by other poxvirus infections, such as BPSV and VACV. Although clinically similar, BPSV infection is associated with lesions in the muzzle of young calves, and PCPV is more often involved in scabby lesions (papulovesicular and pustular) on the udder and skin of dairy cows. 6 Thus, the similarity of clinical presentations of PCPV and BPSV and their identical virion morphology make the correct diagnosis very difficult. 20 Hence, the definitive diagnosis of PPV infection in cattle relies upon the identification of the agent, mainly by molecular means. 20
A number of reports have described the occurrence of poxvirus-associated disease in cattle in Brazil, with many of them reporting human disease as well, and PCPV and BPSV have been frequently suggested as the etiological agents.16,21,23 However, PCPV was identified in only one of these reports (a human case), probably derived from exposure to affected cows. 5 The present report describes bovine cases of pseudocowpox with definitive identification of the agent. However, VACV infection is fairly common in Brazil, and several outbreaks of cattle and human infections have been reported in the last decades.5,8,25 The current report contributes to the recognition and diagnosis of vesicular diseases in cattle in Brazil.
Footnotes
a.
PureLink PCR kit, Invitrogen Corp., Carlsbad, CA.
b.
MegaBACE sequencer, Amersham Biosciences, Piscataway, NJ.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) received no financial support for the research, authorship, and/or publication of this article.
