Abstract
Betacoronavirus 1 (BCoV-1) is an important pathogen causing diarrhea in calves. In the current study, a novel reverse transcription loop-mediated isothermal amplification (RT-LAMP) assay for rapid detection of BCoV-1 was successfully developed. The primers were designed to target the highly conserved fragment of BCoV-1 nucleocapsid gene. The assay displayed high specificity detecting only BCoV-1 with no cross reaction with other viruses. When 418 clinical samples from 6 different geographical areas of Xinjiang province were tested by the RT-LAMP method, the results indicated that this test is a simple, rapid, accurate, and sensitive method for the detection of BCoV-1.
Keywords
Betacoronavirus 1 (BCoV-1; order Nidovirales, family Coronaviridae, subfamily Coronavirinae, genus Betacoronavirus) is one of the important agents of neonatal calf diseases 1 and has become a worldwide infectious agent in the cattle industry.1,3 First identified as a diarrheic agent in calves in 1973 in the United States, 10 BCoV-1 can cause diarrhea in calves, 2 winter dysentry 6 in adult cattle, and respiratory system infections in various aged cattle,8,14 resulting in serious economic losses.
Various molecular detection methods, such as semi-nested reverse transcription polymerase chain reaction (RT-PCR), 15 real-time RT-PCR, 5 and multiplex RT-PCR, 4 have been developed to detect the BCoV-1 infection. However, most of these assays require expensive and special instruments, and therefore may not be readily applicable, particularly in diagnostic laboratories in underdeveloped or developing countries or regions. Thus, it is necessary to develop new assays for the detection of BCoV-1 in clinical practice. In 2000, details of a novel nucleic acid amplification method, namely, the loop-mediated isothermal amplification (LAMP) technique, were published. 11 Compared to other molecular detection assays (single RT-PCR or nucleic acid hybridization), the LAMP technique involves reagents that react under isothermal conditions with high specificity, sensitivity, and rapidity 11 and has been used for the detection of different pathogens.7,13 In the current study, a novel and rapid reverse transcription LAMP (RT-LAMP) assay was developed to detect BCoV-1 from clinical samples.
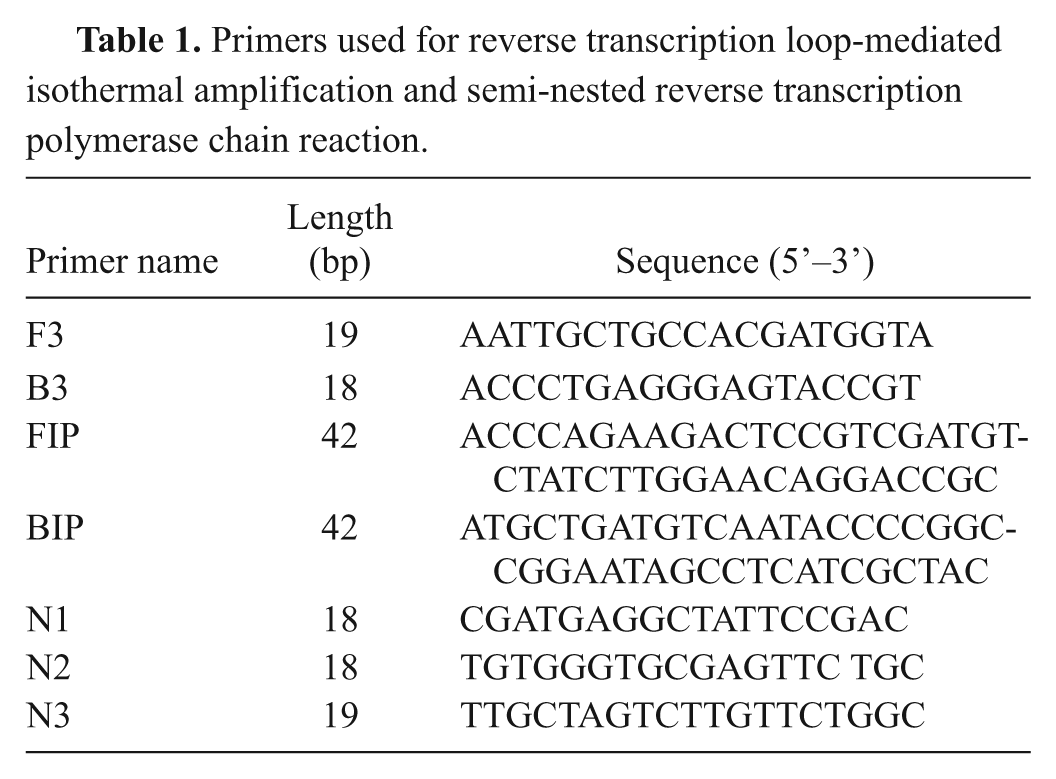
A set of RT-LAMP primers was designed targeting a highly conserved region located in the nucleocapsid (N) protein gene, with the region being selected based on alignment analysis. The analysis involved 24 genomic sequences retrieved from the GenBank database and aligned using the ClustalV method. a Two inner primers, forward inner primer (FIP) and backward inner primer (BIP), and 2 outer primers (F3 and B3) were designed by using the Primer Explorer version 4 software. b All primer sequences were analyzed with BLASTn c for specificity. The primers are shown in Table 1.
Primers used for reverse transcription loop-mediated isothermal amplification and semi-nested reverse transcription polymerase chain reaction.
Total RNA was extracted from BCoV-1–infected human rectal tumor (HRT-18G) cells with a commercial RNA kit, d according to the manufacturer’s instructions. The RNA was eluted in 20 μl of RNase-free water containing 0.04% sodium azide and was stored at −80°C until use. The RNA concentration (ng/μl) was measured with a spectrophotometer. e
The RT-LAMP reaction was carried out in a 25-μl volume containing 12.5 μl of LAMP buffer (20 mmol Tris–HCl [pH 8.8], 10 mmol KCl, 8 mmol MgSO4, 10 mmol (NH4)2SO4, 0.1% Triton X-100, 0.8 M betaine, and 1.4 mmol each of deoxynucleoside triphosphates), 2 μl of primer mixture (40 pmol each of FIP and BIP, and 5 pmol each of F3 and B3), 1 μl of Bst DNA polymerase f (8 U), 1 μl of Avian myeloblastosis virus reverse transcriptase g (40 U), 2 μl of template RNA, and 6.5 μl of distilled water. The reaction mixture was incubated at 63°C for 60 min in a heating block, followed by heating at 80°C for 2 min to terminate the reaction. After the reaction, a 10-μl aliquot of the RT-LAMP product was subjected to electrophoresis in 2.0% agarose gel and visualized by staining with 0.5 μg/ml of ethidium bromide under ultraviolet light. For visual fluorescence detection, 1 μl of a 10-fold dilution of SYBR Green I h (1,000×) was added into the reaction mixture, and the positive amplification was determined according to the color change (orange and green in a negative and a positive reaction, respectively).
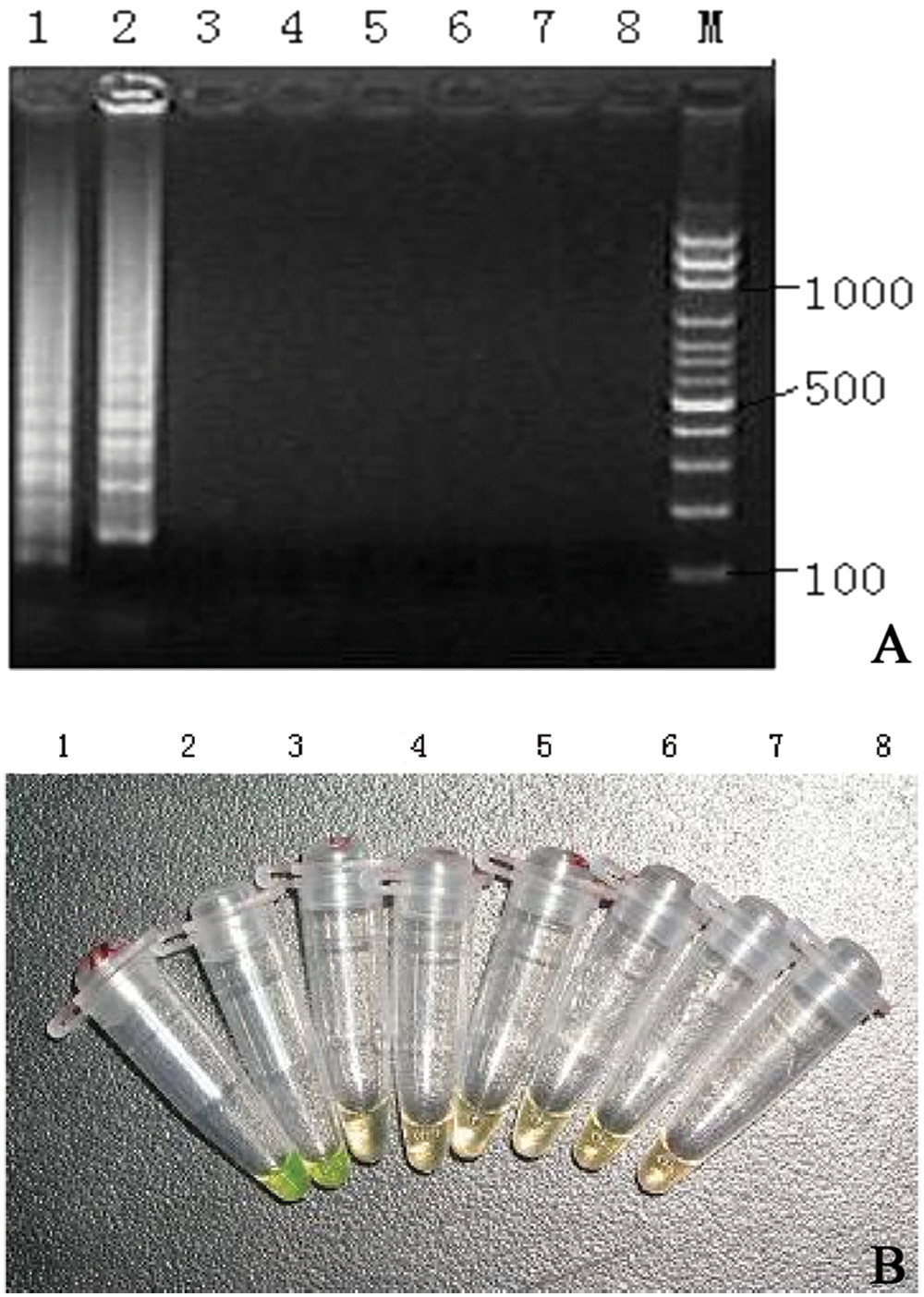
To evaluate the specificity of RT-LAMP, viruses known to cause similar clinical symptoms in newborn calves (Rotavirus A strain 10733 i ; Bovine viral diarrhea virus strain NADL j ) or to be related genetically to BCoV-1 (Canine coronavirus strain INSAVC-1 k ; Transmissible gastroenteritis virus strain Purdue l ; Porcine epidemic diarrhea virus strain CV777 m ) were tested in the present study. Total RNA was isolated from the different cell culture supernatants as described above and subjected to RT-LAMP assay. The specificity of the amplicons was confirmed by digestion with restriction enzyme SphI and sequencing. The results showed that only BCoV-1 was detectable by RT-LAMP assay, and the amplified products were shown as the typical ladder-like bands in 2% agarose gels (Fig. 1a). Follow SphI digestion, the amplification products appeared as 2 fragments of 48 and 153 bp in size. In contrast, none of the other viruses tested in the study were positive. All positive results by gel electrophoresis were also positive by SYBR Green I staining (Fig. 1b). Results confirmed the high specificity of the RT-LAMP assay for BCoV-1.
To analyze the sensitivity of the RT-LAMP assay, serial dilutions of plasmid DNA containing the N gene of BCoV-1 were examined. Briefly, the full length of the BCoV-1 N gene of was amplified and cloned into pGEM-T Easy vector n to generate recombinant plasmid pTN. The Escherichia coli harboring plasmid pTN were inoculated into 10 ml of lysogeny broth and incubated at 37°C overnight on a shaker. The pTN plasmid was extracted from the bacterial pellet and verified with PCR and DNA sequencing. The recombinant plasmid was quantified by spectrophotometer, e the copy number was calculated, and the plasmid DNA standards were prepared by 10-fold serial dilutions. The sensitivity of the RT-LAMP assay was examined by using serial dilutions of plasmid (108, 107, 106, and sequentially down to 100 copies per tube, respectively) with triplicate samples. The results indicated that the RT-LAMP can detect as few as 10 copies of plasmid DNA (Fig. 2), indicating a high sensitivity.
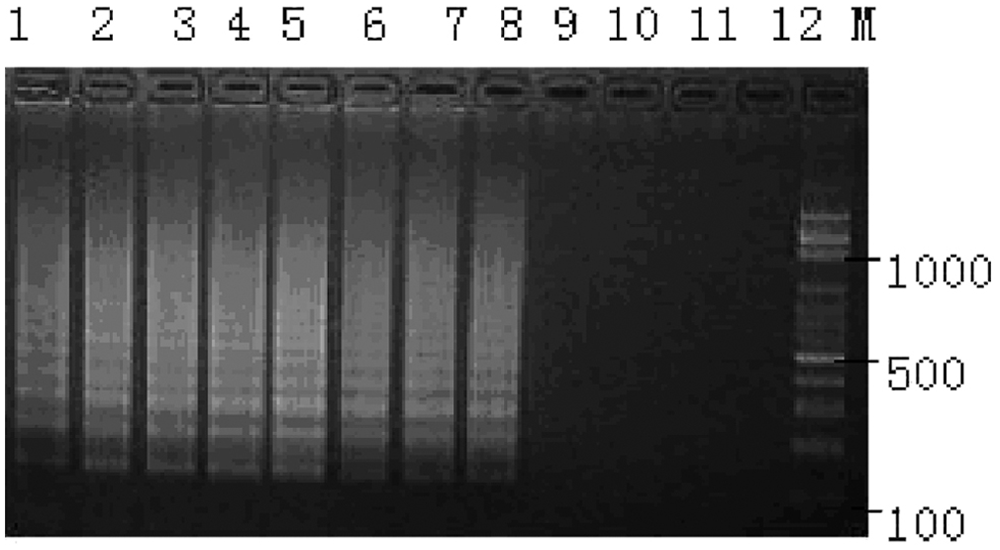
The sensitivity of reverse transcription loop-mediated isothermal amplification was determined using 10-fold serial dilutions of the plasmid DNA. Lane 1: 108 copies; lane 2: 107 copies; lane 3: 106 copies; lane 4: 105 copies; lane 5: 104 copies; lane 6: 103 copies; lane 7: 102 copies; lane 8: 101 copies; lane 9: 100 copies; lanes 10–12: negative control; lane M: 100-bp DNA ladder.
To mimic the real scenario for BCoV-1 testing from fecal samples, different copies (108, 107, 106, down to 101) of plasmid pTN were mixed with feces, and the RNA was re-extracted and detected by LAMP assay. The results showed 102 copies of pTN were detectable in bovine feces. The PCR inhibitors in the feces may be responsible for the lower sensitivity.
The semi-nested RT-PCR was carried out with the specific primers N1 (outer sense primer), N2 (antisense primer), and N3 (inner sense primer; Table 1). The N1 and N2 primers were used for the first amplification after reverse transcription reaction, while N3 and N2 were used for the second round of amplification (semi-nested). The reactions were performed as described previously. 15 Positive amplification was determined by the presence of a 201-bp band and confirmed by DNA sequencing.
To assess the LAMP assay, 12 BCoV-1–positive fecal specimens and 9 negative fecal specimens previously confirmed by semi-nested RT-PCR were examined. The results showed that 12 out of 12 were positive and 9 out of 9 were negative, indicating 100% agreement with the semi-nested RT-PCR.
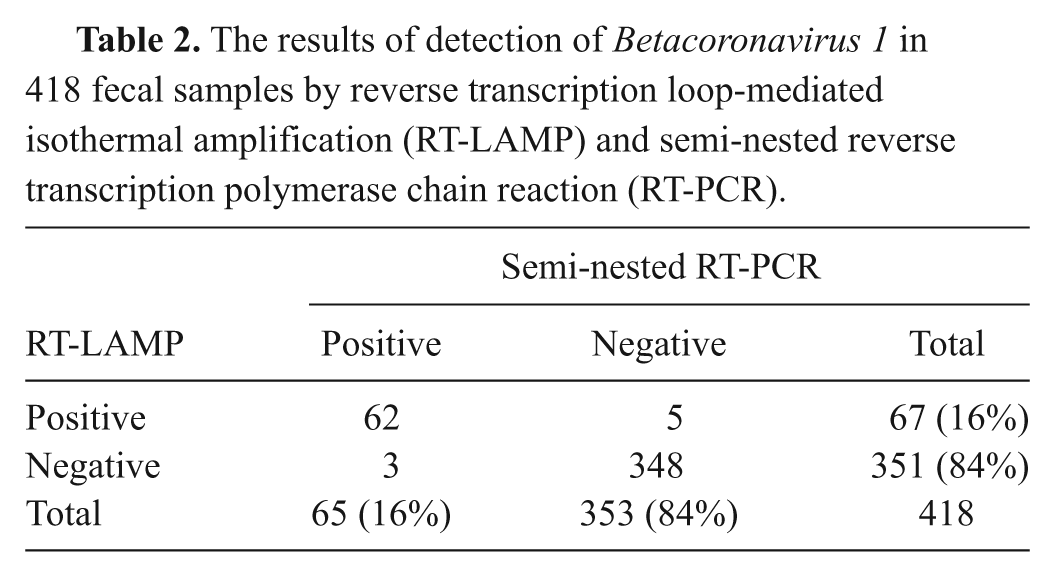
A total of 418 fecal samples from diarrheic calves were collected from cattle farms located in 6 geographical areas (Urumqi, Shihezi, Altay, Qoqek, Ili, and Korla) of Xinjiang province, China during October 2009 through February 2011. Fecal samples were collected from the rectum directly into sterile bottles and kept cold while in transit. Within 24 hr, all fecal samples were re-suspended in 0.01 M phosphate buffered saline (pH 7.2) at 10–20% (w/v), and then centrifuged at 5,000 × g for 10 min at 4°C. Supernatants were collected and subjected to total RNA extraction. The RT-LAMP assay was used to detect the presence of BCoV-1. In addition, semi-nested RT-PCR was employed to examine the presence of BCoV-1 and for testing the agreement between the 2 assays (RT-LAMP and semi-nested RT-PCR). As shown in Table 2, RT-LAMP exhibited high concordance with the semi-nested RT-PCR for BCoV-1 detection from fecal specimens. Sixty-two out of the 418 samples showed BCoV-1 N gene positive from both RT-LAMP and semi-nested RT-PCR assays, while 5 negative samples from semi-nested RT-PCR were BCoV-1 N gene positive by RT-LAMP assay, and 3 negative samples from RT-LAMP assay showed BCoV-1 N gene positive by semi-nested RT-PCR. Three hundred forty-eight samples were proved to be BCoV-1 N gene negative from both assays (Table 2). Accordingly, the percentage of agreement for these 2 assays was 98.2 %.
The results of detection of Betacoronavirus 1 in 418 fecal samples by reverse transcription loop-mediated isothermal amplification (RT-LAMP) and semi-nested reverse transcription polymerase chain reaction (RT-PCR).
The current study reports the development of a novel detection method for the rapid detection of BCoV-1. The RT-LAMP assay allows the user to perform specific and sensitive RNA amplification using simple instruments such as a water bath or a heating block. After specific amplification, the operator can easily visualize the amplification results by adding SYBR Green I into the reaction tube. From the color change of reaction mixtures, the positive results were easily determined by the naked eye. Therefore, this assay is suitable for the diagnosis of BCoV-1 infection in clinical practice, especially in developing countries or regions.
Due to the high efficiency of the LAMP assay, which is higher than conventional PCR, cross-contamination would be a potential issue when using this assay. From the authors’ experiences, the key points to avoid cross contamination are as follows: 1) sterilization of all work areas and laboratory consumables before and after performing the assay (e.g., ultraviolet illumination for at least 30 min); 2) physical separation of the key steps and use of different staff (if possible) to perform those steps; 3) ensuring that the cap of the tubes is tightly closed during RT-LAMP reaction; 4) use of filter tips on pipettes; and 5) change of personal protective equipment (laboratory coat, mask, and gloves) when handling different samples.
It is very important to select a conserved fragment for the design of specific RT-LAMP primers because BCoV-1 strains/isolates exhibit heterogeneity, to a certain extent, at the nucleotide level. Recent studies showed genetic differences between enteric and respiratory BCoV-1, 12 and among isolates from different geographical areas. 9 In the present study, 2 pairs of primers for RT-LAMP were designed based on the highly conserved fragment of N genes derived from both enteric and respiratory BCoV-1 strains and analyzed by BLASTn c for specificity.
The results from 418 clinical samples illustrated that the RT-LAMP assay had a high sensitivity and accuracy, a level that was the same as the semi-nested RT-PCR assay. The results also demonstrated that BCoV-1 was prevalent in calves for various geographical areas of Xinjiang province. In conclusion, a RT-LAMP assay was successfully developed for the detection of BCoV-1 genomic RNA in fecal specimens with the assay showing a high sensitivity and specificity. This rapid, easy-to-operate, and sensitive assay can be used for epidemiological surveillance of BCoV-1 and also for the diagnosis of BCoV-1 infection.
Footnotes
Acknowledgements
The authors thank Dr. Hongbin He for providing bovine Rotavirus A and Bovine viral diarrhea virus as well as for his continuous support, the staff from State Key Laboratory of Veterinary Etiological Biology for technical support, and the field staff who provided the clinical samples for this study.
a.
DNASTAR Inc., Madison, WI.
d.
QIAamp®, Qiagen GmBH, Hilden, Germany.
e.
NanoDrop™ ND-1000, Unicom UV 300; Thermo Fisher Scientific Inc., Waltham, MA.
f.
New England Biolabs Inc., Ipswich, MA.
g.
Takara Bio Inc., Dalian, China.
h.
Invitrogen Corp., Carlsbad, CA.
i.
University of Bari, Bari, Italy.
j.
Molecular Genetics Inc., Minnetonka, MN.
k.
University of Cambridge, Cambridge, United Kingdom.
l.
The Institute for Genomic Research, J. Craig Venter Institute, Rockville, MD.
m.
University of Zurich, Zurich, Switzerland.
n.
Promega Corp., Madison, WI.
The authors declare that they had no conflict of interests with respect to their authorship or the publication of this manuscript.
This study was supported by Natural Science Foundation of Xinjiang Uygur Autonomous Region (no. 2008JC08), China.
