Abstract
Microbiological culture of milk samples has been used as a standard diagnosis for Mycoplasma mastitis. This technique is effective in isolating mollicutes that are Mycoplasma-like; however, isolates may be misinterpreted as Acholeplasma species, which are indistinguishable from Mycoplasma species by culture. A study to contrast the abilities of 2 culture-based tests, digitonin and nisin disc diffusion assays and a conventional polymerase chain reaction (PCR) technique, to discriminate between Mycoplasma and Acholeplasma was performed using 16S ribosomal RNA gene partial sequencing as the gold standard of comparison. A total of 288 bovine mollicute field isolates (248 from milk and 40 from other organ sites) and 13 reference strains were tested. Results obtained from the digitonin disc diffusion assay when it was performed with all field isolates were 92.7% and 99.0% in agreement with the gold standard using 5 mm and 3 mm of zone of growth inhibition as thresholds, respectively. Considering only milk isolates, agreements between the digitonin disc diffusion assay with the gold standard were 97.2% and 100% using 5 mm and 3 mm of zone of growth inhibition as thresholds, respectively. Culture identification using the nisin disc diffusion assay and the PCR was in a 100% agreement with the gold standard. Comparable results using culture-based nisin and digitonin disc diffusion assays, and PCR, to distinguish Mycoplasma and Acholeplasma species was found, especially for isolates from bovine milk.
Keywords
Introduction
Mycoplasma species have been reported to cause severe mastitis in dairy cattle worldwide. 7 Mycoplasma species that cause mastitis are categorized as contagious organisms that can be readily transmitted from cow to cow. Precise detection of infected animals at an early stage is a critical component of mycoplasma mastitis control strategy. 16
Mycoplasma species are fastidious pathogens and have special growth requirements, and isolation of these organisms requires a specific medium, such as modified Hayflick agar. 9 Acholeplasma laidlawii is a common nonpathogenic saprophytic contaminant in the dairy environment and is occasionally found in both tank-milk and cow-milk samples.4,13 Acholeplasma laidlawii is normally considered a nonpathogenic agent, 12 although some studies reported the isolation of this species from mastitic milk.5,20 Acholeplasma species cannot be distinguished from Mycoplasma species on modified Hayflick medium. 9 Therefore, additional discriminatory tests are needed to distinguish between Mycoplasma and Acholeplasma. A major difference between Mycoplasma and Acholeplasma is their sterol requirement. Mycoplasma species are unable to synthesize sterols or fatty acids and thus require exogenous sterol from media. 18 Acholeplasma species do not require exogenous cholesterol for growth. 23 Digitonin can form a complex with sterol, interrupting the uptake of exogenous sterol by Mycoplasma but not Acholeplasma. Thus, growth of Mycoplasma but not Acholeplasma is inhibited by digitonin.
Nisin, an antimicrobial peptide, can also be used to differentiate between Acholeplasma species and Mycoplasma species. 1 Nisin is produced by Lactococcus lactis and can inhibit growth of most Gram-positive and some Gram-negative bacteria. 1 It has been reported that nisin readily inhibited growth of Acholeplasma species whereas Mycoplasma species were substantially less affected by nisin, 1 and, therefore, it has been proposed that a nisin disc diffusion assay on agar plates could be used to distinguish between these 2 genera. 1
Mycoplasma and Acholeplasma species can also be distinguished based on their genotypic characteristics. A duplex nested polymerase chain reaction (PCR) method for detection has been reported. 25 The PCR assay was designed to target the 16S-23S ribosomal RNA (rRNA) intergenic spacer (IGS) region. Because more than one operon of transfer RNA genes in the targeted IGS region are usually presented in the genome of Acholeplasma but not in the genome of Mycoplasma, 19 more than one size of amplicon are usually detected with Acholeplasma species whereas only one size of amplicon is presented with Mycoplasma species. This enables the discrimination of Acholeplasma species from other Mycoplasma species using a PCR assay. 25
Discrimination of Acholeplasma species from Mycoplasma species has been practiced using the digitonin and/or the nisin disc diffusion assay, and by PCR methods, for several years.1,21,27 What is not known is how comparable are these tests of mollicute genera and how accurately these tests can discriminate between these genera. Several local and/or regional veterinary practices who manage their own diagnostic services have inquired on the utility of PCR or digitonin assays in Mycoplasma species identification. An extensive comparative study using a significantly large set of field isolates of various mollicute species has not been conducted. A tested 26 assay system for PCR analysis of mastitis pathogens in milk that is commercially available a has limitations in that the assay does not analyze Mycoplasma species, and the supplementary kit b only detects Mycoplasma bovis. Thus, at best, the tested commercial PCR system available will identify M. bovis but not other mollicutes. Therefore, the objective of the current study was to determine agreement at the genus identification level of Mycoplasma and Acholeplasma species obtained from the digitonin and nisin disc diffusion assays and PCR, using the 16S rRNA gene partial sequence analysis as a gold standard.
Materials and methods
Organisms
A total of 288 isolates from bovine sources were collected. Isolates were obtained from milk samples (n = 248) and from polyester-tipped swabs (n = 40) rubbed over the surfaces of organ sites.3,22 Organ sites included the accessible mucosal surfaces of the nares, ears, eyes, and vagina, as well as the internal organs at necropsy including the respiratory and urogenital systems, the stifle and tarsal joints, lymph nodes, spleen, and pericardial sac. Isolates had been presumptively identified as Mycoplasma from isolation on modified Hayflick agar. c Additionally, 13 reference strains were chosen (Table 1). A filtration-cloning technique, performed as previously described, 28 was used on both reference strains and field isolates. All isolates and strains were cultured in pleuropneumonia-like organism (PPLO) broth c in a 10% CO2 environment for 4 days. Glycerol (30% v/v) was added to cultured broth before storage at −85°C until use.
Sources of reference strains of Mycoplasma and Acholeplasma species.*
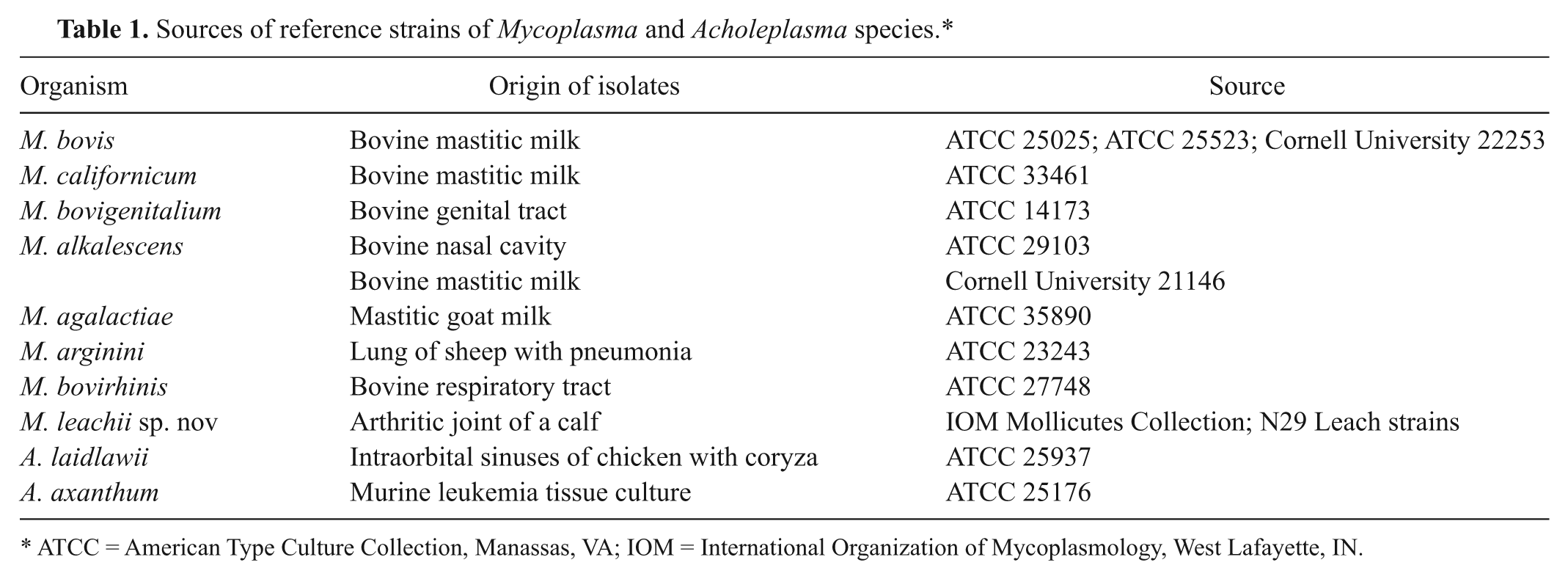
ATCC = American Type Culture Collection, Manassas, VA; IOM = International Organization of Mycoplasmology, West Lafayette, IN.
Digitonin disc diffusion assay
The digitonin disc diffusion assay was performed as previously described. 28 Digitonin d stock solution of 1.5% (w/v) in 95% ethanol was stored at 4°C. Digitonin discs were made by adding 25 µl of the stock solution to 6-mm blank paper discs. e All digitonin discs were dried overnight at room temperature and stored at 4°C until use.
Both unknown and reference mollicutes were cultured in PPLO broth and incubated in a 10% CO2 environment for 4 days. Then, 200 µl of each culture was spread on modified Hayflick agar plate. Digitonin discs were pressed gently to the surface of the agar after the inoculum had dried. Plates were incubated for 4–7 days at 37°C in a 10% CO2 incubator. After incubation, all plates were examined under the stereomicroscope, and the zones of inhibition were measured from the edge of the disc to the edge of clear zone of no growth (mm). Digitonin tests were considered positive when the clear zone was >5 mm, negative when the zone was <3 mm, and all else would normally be considered an ambiguous result. In the analysis of the digitonin test, a threshold of >3 mm and >5 mm were both tested.
Nisin disc diffusion assay
Nisin solution at a concentration of 5.16 mg/ml f was used as a stock solution. A ten-fold dilution was made from the stock solution. A 20-µl aliquot from the stock solution (N0: 103.2 µg) and the dilution (N-1: 10.32 µg) was added to separate 6-mm blank paper discs. All nisin discs were dried overnight at room temperature and stored at 4°C until use. Test organisms were cultured and plated as described for the digitonin disc assay. Nisin discs, both N0 and N-1, were applied to the agar plates and were incubated for 4–7 days at 37°C in a 10% CO2 incubator. After incubation, all plates were examined under the stereomicroscope and were measured for the zone of inhibition (mm). According to a prior study, 1 a positive nisin test was indicated by the presence of a visible growth inhibition zone around the nisin disc. A negative nisin test was indicated by the lack of any inhibition zone.
Genomic DNA extraction for PCR
Mollicute isolates were re-cultured in PPLO broth and incubated at 37°C in a 10% CO2 incubator for 4 days. After incubation, the DNA extraction was performed on all cultures by following instructions for extraction of Gram-negative bacterial genomic DNA. g All genomic DNA extracts were frozen at −80°C until used for PCR.
Discrimination between Acholeplasma and Mycoplasma species using conventional PCR
A one-step PCR was performed by a modification of a previously described assay. 25 The PCR was performed in a total volume of 50 µl containing 1× PCR buffer (20 mM Tris–HCl, 2 mM MgCl2, and 50 mM KCl; pH 8.4 ), 50 µM of each deoxynucleoside triphosphate (dNTP), 20 pmol of each primer, and 1 U Taq DNA polymerase. g A set of primers including F2, R2, and R34 was used to amplify the 16S-23S rRNA IGS regions of Mycoplasma and Acholeplasma. The sequences of the F2 and R2 primers were 5’-GTG(C/G)GG(A/C)TGGATCACCTCCT-3’ and 5’-GCATCCACCA (A/T) A(A/T)AC(C/T)CTT-3’, respectively, with the 16S-23S rRNA IGS region of Mycoplasma species as the target. The sequence of the R34 primer was 5’-CCACTGTGTG CCCTTTGTTCCT-3’ with the 16S-23S rRNA IGS region of Acholeplasma species as the target. A 5-µl aliquot of the DNA extract was used as the template and added to 45 µl of reaction mixture. The thermal cycling protocol was initial denaturation at 94°C for 30 sec, followed by 35 cycles of denaturation at 94°C for 30 sec, primer annealing at 55°C for 2 min, and extension at 72°C for 2 min. A final extension was performed at 72°C for 5 min. The final PCR products were electrophoresed on a 2% agarose gel, and DNA bands were visualized by ultraviolet fluorescence. A single band of PCR product indicated the presence of a Mycoplasma species. The presence of more than 1 band indicated an Acholeplasma species.
16S ribosomal RNA gene partial sequencing
Identification of all 301 isolates was confirmed using 16S rRNA gene partial sequencing as a gold standard. A region of approximately 1,500 base pairs of the 16S rRNA gene was amplified using a universal primer pair, 24 pH and pA. The pA and pH primers had sequences of 5’-AGAGTTTGAT CCTGGCTCAG-3’ and 5’-AAGGAGGTGATCCAGCCGC A-3’, respectively. The PCR reaction mixture, a total volume of 50 µl, included 3 U of Taq DNA polymerase, 1× PCR buffer (20 mM Tris–HCl, 1.5 mM MgCl2, and 50 mM KCl; pH 8.4), 50 µM of each dNTP, 10 pmol of each primer, and 1 µl of the genomic DNA as template. The thermal cycling protocol involved initial denaturation at 95°C for 3 min followed by 35 cycles of denaturation at 94°C for 15 sec, primer annealing at 57°C for 15 sec, and extension at 72°C for 2 min. A final extension was performed at 72°C for 5 min. The PCR products were purified according to manufacturer’s directions and submitted for partial sequencing, h single strand, single direction using primer pA. Nucleotide sequences were compared with GenBank database using the Basic Local Alignment Search Tool (BLAST; http://www.ncbi.nlm.nih.gov/blast/Blast.cgi). 2 A nucleotide sequence was identified as a species giving the highest hit of at least 98% identity from BLAST search.
Statistical analysis
Nisin and digitonin disc assays were run in duplicate. If results of duplicate assays were not in accordance then a third assay was performed. The definitive identification of an isolate was then given when results of 2 out of 3 replicates agreed. Genera classification by digitonin and nisin disc diffusion assays, and PCR, were compared with results from 16S rRNA gene partial sequencing and presented as percentage agreement. The percentage agreement with 16S rRNA gene partial sequencing, the gold standard, was calculated for each technique where the denominator was the number of isolates identified as either Mycoplasma or Acholeplasma by the gold standard and the numerator was the number of isolates identified as either Mycoplasma or Acholeplasma by each technique.
Results
Mycoplasma isolates identified by 16S rRNA gene partial sequencing
A total of 267 isolates (92.7%) were identified as Mycoplasma species based on the results of 16S rRNA gene partial sequencing. When including the 13 reference strains, the following Mycoplasma species were represented in this pool of identified isolates with the total number of identified in parentheses: M. bovis (231), M. californicum (23), M. bovigenitalium (6), M. alkalescens (4), M. arginini (4), M. yeatsii (2), M. cottewii (1), M. bovirhinis (1), M. felis (1), M. leachii (1), and M. agalactiae (4). The 16S rRNA gene partial sequencing identified 21 A. laidlawii and 2 A. axanthum isolates. All reference strains were correctly identified by 16S rRNA gene partial sequencing.
Discrimination between Mycoplasma and Acholeplasma using digitonin disc diffusion assay
All reference strains of Mycoplasma species showed zones of growth inhibition around the digitonin discs as indicated in Table 2. Sizes of zones of growth inhibition ranged from 3 to 13 mm. No zones of growth inhibition were observed with Acholeplasma species (Table 2).
Zone of growth inhibition (mm) observed with the digitonin and nisin disc diffusion assays for reference strains (Mycoplasma and Acholeplasma species) isolates.*
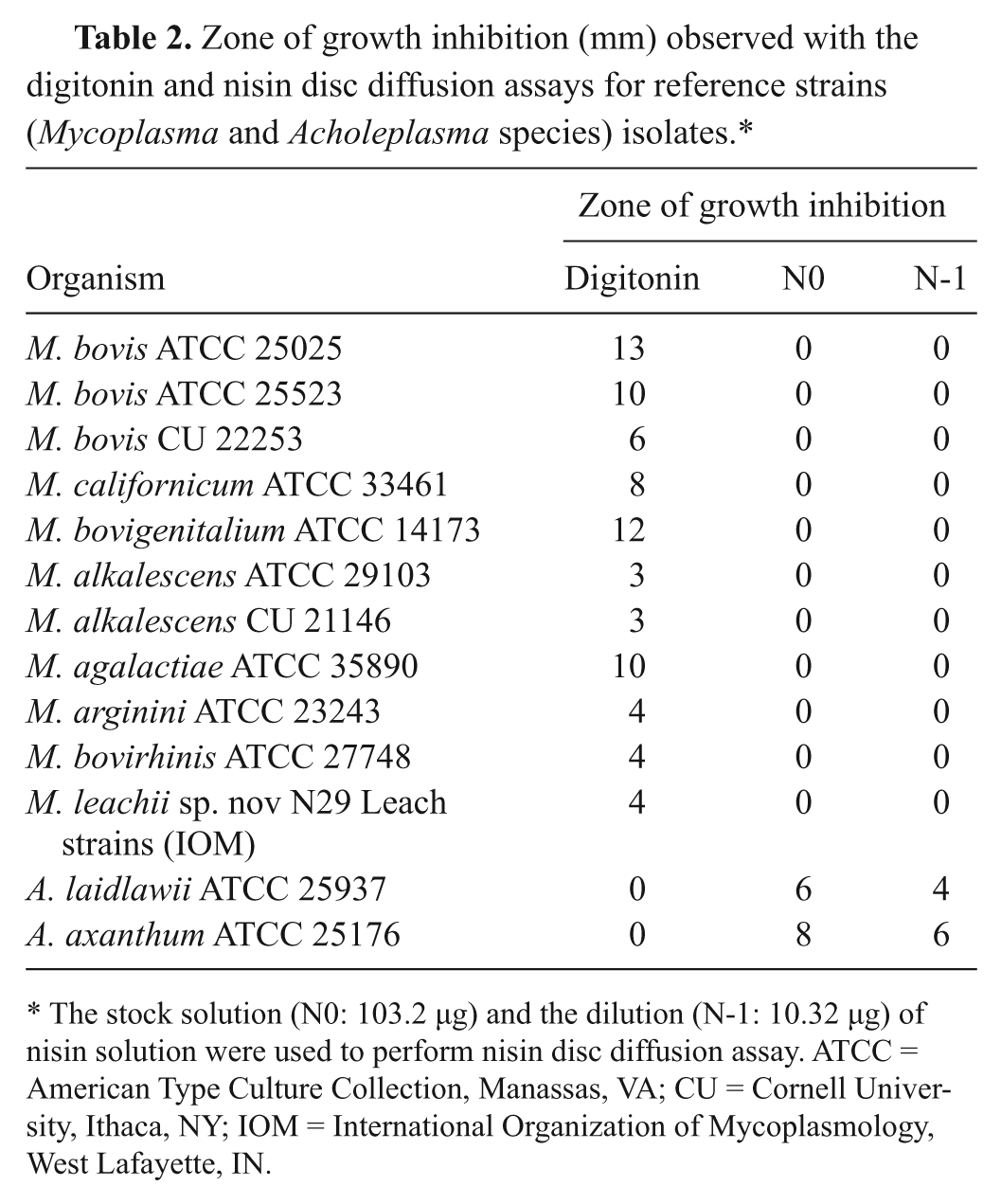
The stock solution (N0: 103.2 µg) and the dilution (N-1: 10.32 µg) of nisin solution were used to perform nisin disc diffusion assay. ATCC = American Type Culture Collection, Manassas, VA; CU = Cornell University, Ithaca, NY; IOM = International Organization of Mycoplasmology, West Lafayette, IN.
Of 288 field isolates, 267 isolates showed zones of growth inhibition around the discs, ranging in size from 1 to 20 mm, mean ± standard error (SE) of 11.07 ± 0.12 mm. Descriptive statistics of zones of growth inhibition were indicated in Table 3. Zones of inhibition observed from M. bovigenitalium and M. bovis were the largest with 14.67 ± 0.70 and 11.85 ± 0.08, respectively. Twenty-four isolates were categorized as negative, which included 20 isolates of A. laidlawii, 1 isolate of A. axanthum, 2 isolates of M. yeatsii, and 1 isolate of M. cottewii (Table 3).
Descriptive statistics of zone of growth inhibition (mm) observed with the digitonin and nisin disc diffusion assays for discrimination of Mycoplasma and Acholeplasma species.*
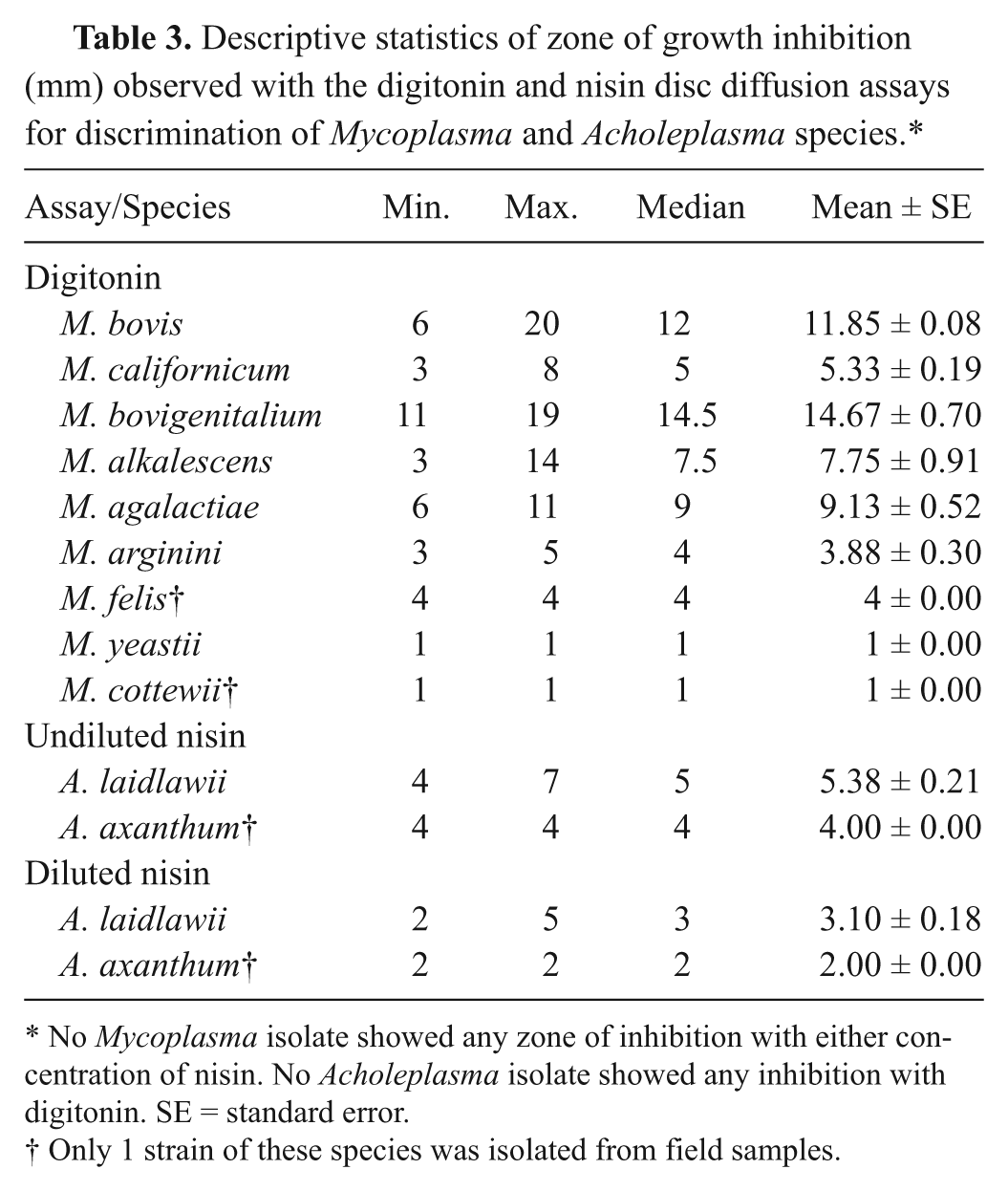
No Mycoplasma isolate showed any zone of inhibition with either concentration of nisin. No Acholeplasma isolate showed any inhibition with digitonin. SE = standard error.
Only 1 strain of these species was isolated from field samples.
All the M. bovis isolates tested (n = 228) had zones of growth inhibition of >5-mm threshold. Thus, at this threshold, M. bovis had 100% agreement with the 16S rRNA gene partial sequencing technique (Table 4). Equivalent agreements with M. bovigenitalium (5/5), M. alkalescens (2/2), and M. agalactiae (3/3) isolates were also observed (Table 4). Percentages of agreement for M. californicum was only 31.8% (7/22) using the 5-mm threshold (Table 4). The overall Mycoplasma species agreement of digitonin disc diffusion assay to 16S rRNA gene partial sequencing technique was 92.7% (267/288) using the ≥5-mm threshold. Considering only milk isolates, agreements between the digitonin disc diffusion assay with the gold standard were 97.2% (241/248) and 100% using 5 mm and 3 mm of zone of growth inhibition as thresholds, respectively.
Agreement of genera determination of Mycoplasma and Acholeplasma species by digitonin disc diffusion assay with 16S ribosomal RNA gene partial sequencing.*
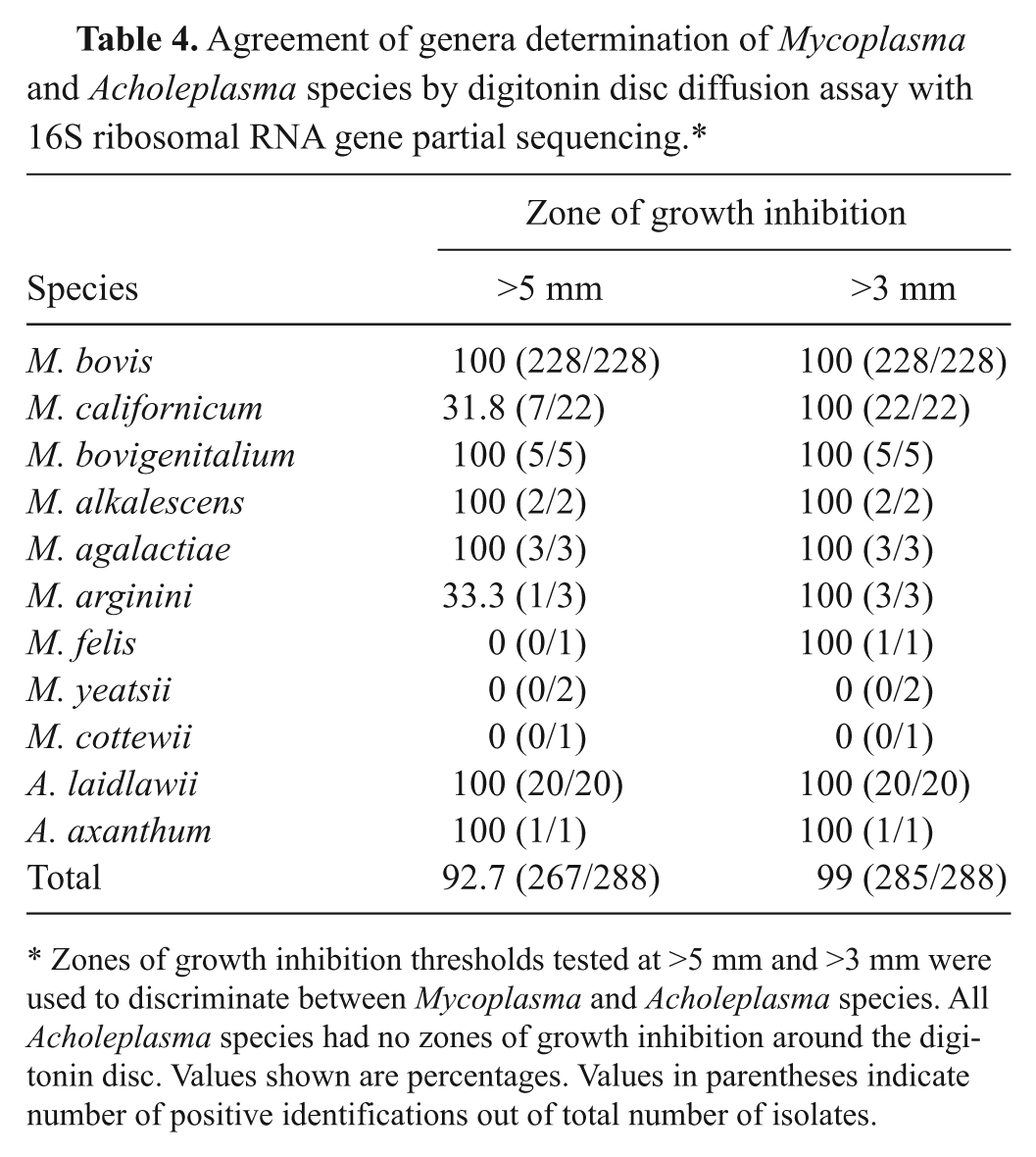
Zones of growth inhibition thresholds tested at >5 mm and >3 mm were used to discriminate between Mycoplasma and Acholeplasma species. All Acholeplasma species had no zones of growth inhibition around the digitonin disc. Values shown are percentages. Values in parentheses indicate number of positive identifications out of total number of isolates.
In contrast, when the presence of a zone of inhibition of ≥3 mm was considered to be the threshold for a positive test, the accuracy of the digitonin disc diffusion assay, in terms of agreement with the 16S rRNA gene partial sequencing technique, was 100% for all Mycoplasma species except for 2 isolates of M. yeatsii and 1 of M. cottewii as shown in Table 4. Using the 3-mm threshold, the overall agreement of using digitonin disc diffusion assay to distinguish between mycoplasmas and acholeplasmas compared to 16S rRNA gene partial sequencing technique was 99% (285/288). All M. californicum isolates were correctly identified with the ≥3-mm zone of growth inhibition threshold.
Discrimination between Mycoplasma and Acholeplasma using nisin disc diffusion assay
All reference strains of Acholeplasma species showed zones of growth inhibition around the nisin discs as indicated in Table 2. Sizes of zones of growth inhibition ranged from 6 to 8 mm and 4 to 6 mm for N0 and N-1, respectively. No zones of growth inhibition were observed with reference strains of Mycoplasma species (Table 2).
No zones of nisin-inhibited growth were seen with any Mycoplasma tested. Descriptive statistics of zones of growth inhibition are indicated in Table 3. All isolates of Acholeplasma species had zones of growth inhibition with the nisin disc diffusion assay for both N0 and N-1 discs. Ranges of zones of inhibition were 4–7 mm and 2–5 mm for N0 and N-1, respectively. For A. laidlawii, mean ± SE of zones of inhibition for nisin disc diffusion assay were 5.38 ± 0.21 mm and 3.10 ± 0.18 mm for N0 and N-1, respectively. All Mycoplasma and Acholeplasma isolates were correctly identified by the nisin assay with a percentage agreement of 100% (288/288) with 16S rRNA gene partial sequencing.
Discrimination between Mycoplasma and Acholeplasma using PCR technique
The amplicons of all Mycoplasma and Acholeplasma isolates correctly identified each genus; Mycoplasma always yielded a single amplicon and Acholeplasma yielded 2 amplicons. The PCR test resulted in a 100% agreement (288/288) with 16S rRNA gene partial sequencing.
Discussion
The main focus of the current study was to contrast the accuracy of Mycoplasma and Acholeplasma species identification using the digitonin and nisin disc diffusion assays and PCR, as compared to the gold standard 16S rRNA gene partial sequencing technique. The nisin and PCR tests were in perfect agreement (100%) with the gold standard. Results obtained from the digitonin disc diffusion assay agreed more than 90% of the time with the gold standard. The use of a threshold of >5 mm in the digitonin assay yielded 92.7% agreement, while a threshold of >3 mm yielded 99% agreement with the gold standard. Moreover, the Acholeplasma species tested all had zones of growth inhibition of <3 mm. Therefore, the results obtained from the digitonin disc diffusion assay can distinguish between the 2 genera originally isolated from mastitis cases. In aggregate, these findings indicate that all 3 techniques performed in the present study can be used to distinguish between Mycoplasma and Acholeplasma species isolated from bovine origin, and from milk samples specifically, with a high degree of certainty. Of culture-based tests, the nisin disc assay appeared to be superior to the digitonin.
Two hundred forty-eight isolates tested in the current study (86.1%) were from milk samples from cows with clinical mastitis. Of these mastitis isolates, 85.5% (212/248), 4.4% (11/248), and 0.8% (2/248) were M. bovis, M. californicum, and M. bovigenitalium, and 9.3% (23/248) were identified to be other species. This finding is consistent with other studies done in California identifying M. bovis as the most prevalent Mycoplasma species, followed by M. californicum and M. bovigenitalium, isolated from samples of clinical mastitis cases 11 and bulk-tank milk samples. 15
The digitonin disc diffusion assay has been used as a standard method to discriminate Acholeplasma species from Mycoplasma species in many laboratories capable of culturing Mollicutes as suggested by the Subcommittee on the Taxonomy of Mollicutes. 10 In the current study, all isolates identified to be M. bovis, M. bovigenitalium, and M. agalactiae were accurately identified as Mycoplasma based on the presence of zone of growth inhibition using either >5 mm or >3 mm as a threshold. Using the zone of growth inhibition threshold of >5 mm as suggested previously, 28 68.2% (15/22) of M. californicum isolates were misclassified. Using the zone of growth inhibition threshold of >3 mm, all mycoplasma isolates originally isolated from milk samples were correctly categorized as Mycoplasma species. There were only 3 isolates originally isolated from bovine lung tissues (2 M. yeatsii isolates and 1 M. cottewii isolate) that were slightly sensitive to digitonin (1 mm of zone of growth inhibition) and thus would be misclassified regardless of the threshold of zone of inhibition used. Results from the current study for these 3 isolates differ from another report 6 where the sensitivity to digitonin was greater as measured by a zone of growth inhibition of 6–11 mm for M. yeatsii and M. cottewii.
It appears that zones of growth inhibition by the digitonin disc diffusion assay vary by species. Larger zones of growth inhibition (mean ± SE) were observed for M. bovigenitalium (15.30 ± 0.67 mm), M. alkalescens (12.75 ± 0.48 mm), and M. bovis (11.89 ± 0.07 mm) compared to M. californicum (5.23 ± 0.19 mm) and M. arginini (5.20 ± 0.92 mm). An earlier study 27 reported larger zones for M. arginini (12 mm) and M. californicum (9.74 mm) whereas small zones were observed for M. canadense (8.35 mm) and M. alkalescens (7.87 mm). In the current study, the digitonin disc diffusion assay correctly identified Acholeplasma isolates. Several studies8,14 demonstrated some variation of zones of growth inhibition of Acholeplasma species isolated from bovine origin. The studies reported zones of inhibition of Acholeplasma species to digitonin ranging from 0.5 to 1.5 mm. Another study 27 reported zones of inhibition among 20 isolates of A. laidlawii ranging from 1 to 4 mm. However, all Acholeplasma isolates included in the current study showed no zone of growth inhibition around the digitonin discs.
It has been reported that the nisin disc diffusion assay could be used to discriminate Acholeplasma from Mycoplasma. 1 All 5 Acholeplasma species had zones of inhibition whereas all 11 Mycoplasma species did not show any zone of inhibition. 1 The findings of the current study were similar in that growth of all Acholeplasma and no Mycoplasma were inhibited by the nisin discs at 2 concentration levels. There was no advantage to using the more concentrated solution (N0) of nisin as opposed to a ten-fold diluted concentration (N-1).
The PCR also had 100% agreement with the gold standard. Although PCR-based platforms for the detection of mastitis pathogens directly in milk have been evaluated,17,21 an extensive study of the ability of PCR assays to distinguish between Acholeplasma and Mycoplasma species using field-based milk samples has not been made. The results reported herein are consistent with previous findings that identified 2 PCR products for Acholeplasma species and 1 PCR product for all Mycoplasma species as described. 25
In conclusion, the present study suggests a high and comparable efficiency of using nisin and digitonin disc diffusion assays and PCR to distinguish Mycoplasma and Acholeplasma species. Thus, the study confirms those of others that all the assays can distinguish between Acholeplasma and Mycoplasma species. Choosing a technique to use to discriminate between Mycoplasma and Acholeplasma species depends upon the instrumentation requirements, labor, cost, and toxicity of chemicals. Smaller veterinary service diagnostic laboratories may choose to use the less sophisticated tests of the digitonin or nisin disc diffusion–based assays. The PCR assay requires more instrumentation but will generate a more rapid result. Results of the current study indicate that the nisin and digitonin disc diffusion assays performed well and yielded accurate results for standard culture of isolates of veterinary origin. The genotypic test, using PCR assay, was very effective in discriminating mollicutes as a Mycoplasma or Acholeplasma species.
Footnotes
Acknowledgements
The authors would like to thank ImmuCell Corp. (Portland, ME), especially Dr. Joseph Crabb, for the kind supply of nisin. The authors are grateful to Dorothy Newkirk and Veerasak Punyapornwithaya for their excellent assistance.
a.
PathoProof™ Mastitis Complete-12, Finnzymes Oy, Espoo, Finland.
b.
PathoProof™ Mastitis Major-3, Finnzymes Oy, Espoo, Finland.
c.
BBL, Franklin Lakes, NJ.
d.
Sigma-Aldrich, St. Louis, MO.
e.
BD, Sparks, MD.
f.
ImmuCell Corp., Portland, ME.
g.
Invitrogen Corp., Carlsbad, CA.
h.
Macrogen Corp., Rockville, MD.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) received no financial support for the research, authorship, and/or publication of this article.
