Abstract
The incidence and severity of disease associated with toxigenic Clostridium difficile have increased in hospitals in North America from the emergence of newer, more virulent strains. Toxigenic C. difficile has been isolated from food animals and retail meat with potential implications of transfer to human beings. The objective of the present study was to determine the prevalence of C. difficile in pork from sausage manufacturing plants and retail meat in Texas. Twenty-three C. difficile isolates were detected from 243 meat samples (9.5%) from 3 sausage-manufacturing plants and 5 retail meat outlets from 2004 to 2009. Twenty-two isolates were positive for toxins A, B, and binary toxin, and were characterized as toxinotype V, PFGE type-NAP7, or “NAP7-variant.” Susceptibilities to 11 antimicrobial agents in the current study were similar to those reported previously for toxinotype V isolates, although the results suggested somewhat reduced resistance than reported for other meat, animal, or human clinical toxinotype V isolates.
Since 2003, the incidence and severity of disease associated with toxigenic Clostridium difficile have increased in hospitals in North America. 15,16 Indications are that these increases may be due to the emergence of a new strain of toxigenic C. difficile (restriction endonuclease analysis type BI, North American pulsed-field gel electrophoresis pattern 1 [BI/NAP1], toxinotype III) that exhibits increased levels of resistance, virulence, and toxin production. Community-acquired, C. difficile–associated disease also is on the increase. 1 The origins of this new strain have yet to be determined. Various strains of C. difficile, including NAP1, toxinotype III, can be isolated from food animals and meat 17,24,26 ; however, the predominant strains from food animals are NAP7 and NAP8, toxinotype V. 6,8,19,26 Because food animals can be colonized by C. difficile, and the bacteria has been isolated from retail meats, 17,21,24-26 some researchers speculate that C. difficile is a food-associated organism and consumption of contaminated meat could be responsible for increased community-associated C. difficile infection. 6,8,14,22 The objective of the present study was to determine the prevalence of C. difficile in 3 sausage manufacturing plants and 5 retail meat outlets in Texas.
The authors collected meat samples and swabs from 3 sausage-manufacturing plants during 2004–2009 and tested the samples for C. difficile. Plant A was sampled 5 times per year during 2004, 2005, and 2006 for a total of 149 samples (10 g each) of pork trim. Plant B was sampled in November of 2009; 20 samples were collected from 9.1 kg of pork trim and 12 sponge swabs a were collected from equipment and facilities (total of 32 samples). Plant C was sampled in December of 2009; 12 samples were collected from twelve 454-g chubs of ground pork and 10 sponge swabs a were collected from holding pens, feces, hide, live animal, and carcasses (a total of 22 samples). Meat samples from plant C represented a 12-hr production shift of approximately 32,727 kg of ground pork. Sponge samples from plants B and C were collected according to the procedures described for testing of meat production facilities. 3-5 After collection, meat and sponge samples from plants were placed into sterile bags and transported on ice to the Food and Feed Safety Research Unit, Agricultural Research Service, U.S. Department of Agriculture, College Station, Texas.
On 4 separate dates during 2008–2009, 40 retail meat samples were collected from 5 different meat markets and grocery stores in Bryan and College Station, Texas. Samples included both store and national brands and consisted of ground pork, ground beef, ground chicken, ground turkey, pork sausage, pork and beef summer sausage, pork longaniza, pork chorizo, and pork beer bratwurst.
Alcohol shock, enhanced enrichment and/or concentration techniques, restrictive media, and anaerobic incubation were utilized to cultivate C. difficile according to procedures for retail meat as previously described. 21 Briefly, cycloserine cefoxitin fructose broth (CCFB) was prepared by adding 0.1% sodium taurocholate, D-cycloserine (250 mg/l), and cefoxitin (8 mg/l) to CM0601 base without agar b in an anaerobic chamber. Meat packages were opened aseptically in an anaerobic chamber, and 5 g was removed and inoculated into 20 ml of CCFB and incubated anaerobically at 37°C for 15 days. Alcohol shock was used to induce spore germination by mixing 2 ml of homogenized culture broth and 96% ethanol (1:1, v/v) for 50 min. After centrifugation (3,800 × g for 10 min), the supernatant fluid was removed, 600 µl of sterile deionized water was added to the sediment and the suspension was mixed thoroughly. The sediment suspension was streaked onto cycloserine cefoxitin fructose agar c and incubated anaerobically at 37°C for 5 days.
Sponge samples were cultivated in an identical fashion to meat except for the initial step when the whole sponge (wet weight, 13.4 g) was aseptically removed from the bag and, with sterile scissors, cut into 4 pieces, and inoculated into 20 ml of CCFB in an anaerobic chamber. Presumptive diagnosis consisted of the presence of colonies morphologically similar to C. difficile, L-proline aminopeptidase activity, d biochemical characterization, e and by presence of the tcdC regulatory gene.
Polymerase chain reaction (PCR) was used to detect toxin A and B genes, tcdC gene deletion, toxinotyping, and cdtB binary toxin gene. DNA extraction was accomplished by the use of a commercial kit. f The PCR procedures for toxin A gene (tcdA), toxin B gene (tcdB), cdtB binary toxin gene, and regulatory gene (tcdC) detection followed the techniques according to PCR protocols as utilized by the Centers for Disease Control and Prevention (CDC; Atlanta, Georgia). 10,11,13 Toxinotyping was performed by amplification of the tcdA and tcdB genes followed by restriction. The toxin A gene was restricted with EcoRI, g and the toxin B gene was restricted with HinC II and AccI. g Banding patterns for toxinotypes were compared to a toxinotype V control and toxinotype III control. Patterns differing from the 2 controls were compared to images in published articles. 23
Each C. difficile isolate was tested for susceptibility to 11 antimicrobial agents (ampicillin, chloramphenicol, tetracycline, amoxicillin–clavulanic acid, imipenem, cefoxitin, metronidazole, ciprofloxacin, clindamycin, piperacillin–tazobactam, and vancomycin) by use of a commercially available test h according to the manufacturer’s recommendations. Results were interpreted according to Clinical and Laboratory Standards Institute standard criteria, 2 except minimum inhibitory concentrations (MIC) for ciprofloxacin were based on values for trovafloxacin, and vancomycin interpretation was based on MICs reported for Gram-positive aerobes. Quality control strains, Bacteroides fragilis (American Type Culture Collection [ATCC] 25285), Bacteroides thetaiotaomicron (ATCC 29741), and Eggerthella lenta (ATCC 43055) were tested using recommended breakpoints for MICs. 2
Pulsed field gel electrophoresis (PFGE) was used to compare C. difficile isolates. The PFGE procedures followed techniques of a modified 7-day protocol utilized by the CDC (A. Thompson, personal communication, 2008). 13 Gels were made, restricted with SmaI, g run for 18 hr on a commercial PFGE system, i stained, and imaged. j Salmonella choleraesuis subsp. choleraesuis (Smith) Weldin serotype Braenderup (ATCC BAA-664 [H9812]) was used as a quality control organism and was run in every fifth lane of gels.
Plant A had 14 (11/51 in 2004, 1/53 in 2005, and 2/45 in 2006) C. difficile isolates detected from 149 pork trim samples (9.4%; Table 1). Thirteen isolates were determined to be toxinotype V and 1 was toxinotype XI; 13 were positive for toxins A and B, and binary toxin, and 1 was negative for toxins A and B, and positive for binary toxin; 5 were PFGE type-NAP7 (including the isolate belonging to the toxinotype XI), and 9 were NAP7-variant (91% similar). Plant B had 1 C. difficile isolate detected from 20 pork trim samples (5.0%) and none detected from 12 equipment or carcass samples. Plant C had 2 C. difficile isolates detected from 12 ground pork samples (16.7%), and 3 C. difficile isolates detected from 10 animal sponge samples (30%; carcass hide, post excision-hide, and ears). All 6 isolates from plants B and C were positive for toxins A and B, and binary toxin, had 39-bp deletion in the tcdC regulatory gene, and were toxinotype V. Results of PFGE indicated that 2 isolates were NAP7, 1 of the isolates was 90.2% similar to NAP7, and the other 3 isolates were 88.6% similar to NAP7.
Toxinotype; possession of tcdA gene, tcdB gene, and cdtB binary toxin gene; tcdC gene deletion; and pulsed-field gel electrophoresis (PFGE)-type of 23 Clostridium difficile isolates from retail meat outlets and sausage-manufacturing plants in Texas.*
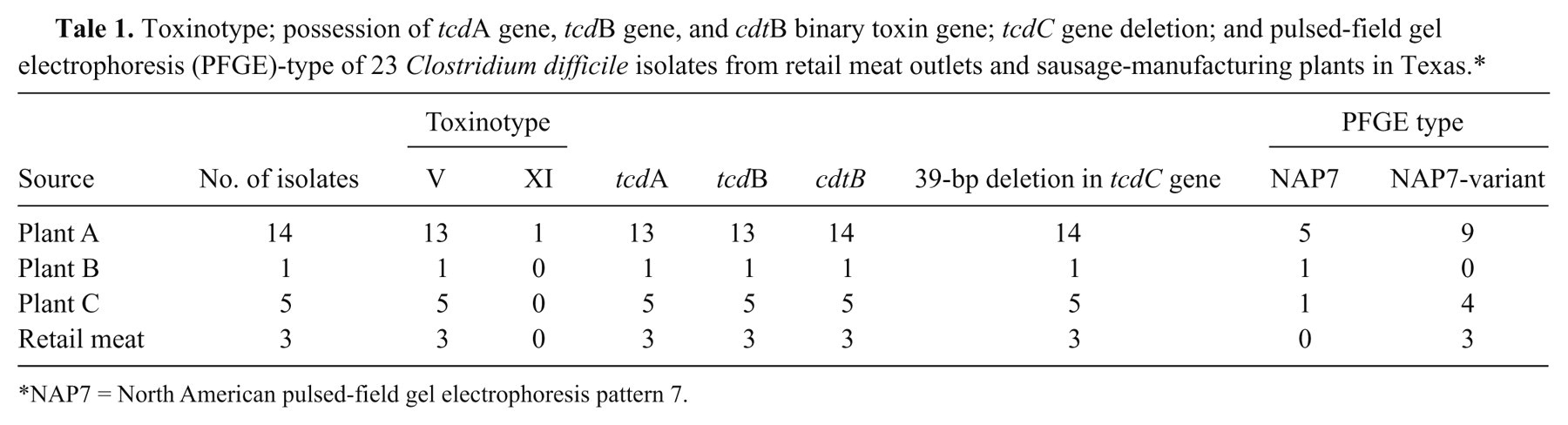
NAP7 = North American pulsed-field gel electrophoresis pattern 7.
Of the 40 samples of meat from retail facilities, C. difficile was detected in 3 samples (7.5%). The C. difficile isolates occurred in ground pork, pork chorizo, and ground turkey, and came from the same store on the same sample date. The pork chorizo and ground turkey were national brands whereas the ground pork was a store brand. The isolates were toxinotype V and 91% related to the other NAP7-variant isolates.
In the current study, 22 out of the 23 C. difficile isolates were toxinotype V and PFGE type NAP7 or NAP7-variant. This agrees with the majority of studies that have reported toxinotype V, NAP7 or NAP8, ribotype 078 as the predominant strain of C. difficile in pigs and pork 8,12,18 ; although toxinotype III (NAP1 and NAP1-related) strains have been recovered from pork, veal, and ground beef. 21,24,26 In agreement with the studies mentioned above, 2 C. difficile were detected from meat and 3 from the hide and ears of pork carcasses from plant C; PFGE results indicated that all 5 were toxinotype V and 88–90% related to NAP7. The authors are unsure of the clinical relevance of these findings as pertains to potential transfer of C. difficile from animals to meat to human beings.
The overall C. difficile prevalence for meat and sausage manufacturing plants in the present study was 9.5% (23/243), and with the exception of a single study that had a 42% prevalence rate, 24 prevalence in the current study compares favorably with the 6.7% in ground beef and 4.6% in veal chops, 20 12% in ground beef and ground pork, 26 12.8% in retail chicken, 27 and 20% in ground beef 21 from studies conducted in North America. These rates are in sharp contrast to C. difficile recovery rates from meat in Europe. In Sweden, only 2 (ground beef) of 82 (2.4%) meat samples were positive for C. difficile 25 In the Swedish study, samples consisted of diverse types of meat including beef, pork, mixed beef and pork, sheep, moose, reindeer, calf, poultry, and cooked meats and sausages. The isolates were not classified as to toxinotype or ribotype, but were positive for both toxins A and B. In a study in Austria, zero out of 84 meat samples (51 beef, 27 pork, 6 chicken) were positive for C. difficile 7 ; whereas, in another Austrian study, 3 out of 100 (3%) were positive for C. difficile. 9 All 3 C. difficile isolates from the Austrian study were from mixed beef and pork; 2 were ribotype AI-57 (Austrian isolate), and the third was ribotype 053 (positive for toxin A and B genes). French researchers detected 3 C. difficile isolates from 65 (4.6%) ground beef samples and zero out of 50 from pork sausage samples (Bouttier S, Barc MC, Felix B, et al.: 2008, Screening for Clostridium difficile in meat from French retailers. In: Proceedings of the 18th Meeting of the European Society of Clinical Microbiology and Infectious Diseases, Barcelona, Spain). Two belonged to toxinotype 0 (binary toxin negative, moxifloxacin susceptible) and 1 to toxinotype II (binary toxin positive, an 18-bp deletion in tcdC, moxifloxacin resistant).
The sensitivities to 11 antimicrobial agents for C. difficile isolates for pork trim from plants A and B, ground pork and carcasses from plant C, and from the retail meat sources are listed in Table 2. The majority of sensitivity patterns were similar regardless of the source. However, 3 exceptions occurred: the 2 isolates resistant to clindamycin came from retail meat; all 4 isolates resistant to tetracycline came from plants B and C; and the 12 isolates intermediate to ciprofloxacin came from plant A.
Antimicrobial interpretive categories and interpretive results of 23 Clostridium difficile isolates from retail meat outlets and sausage-manufacturing plants in Texas.*
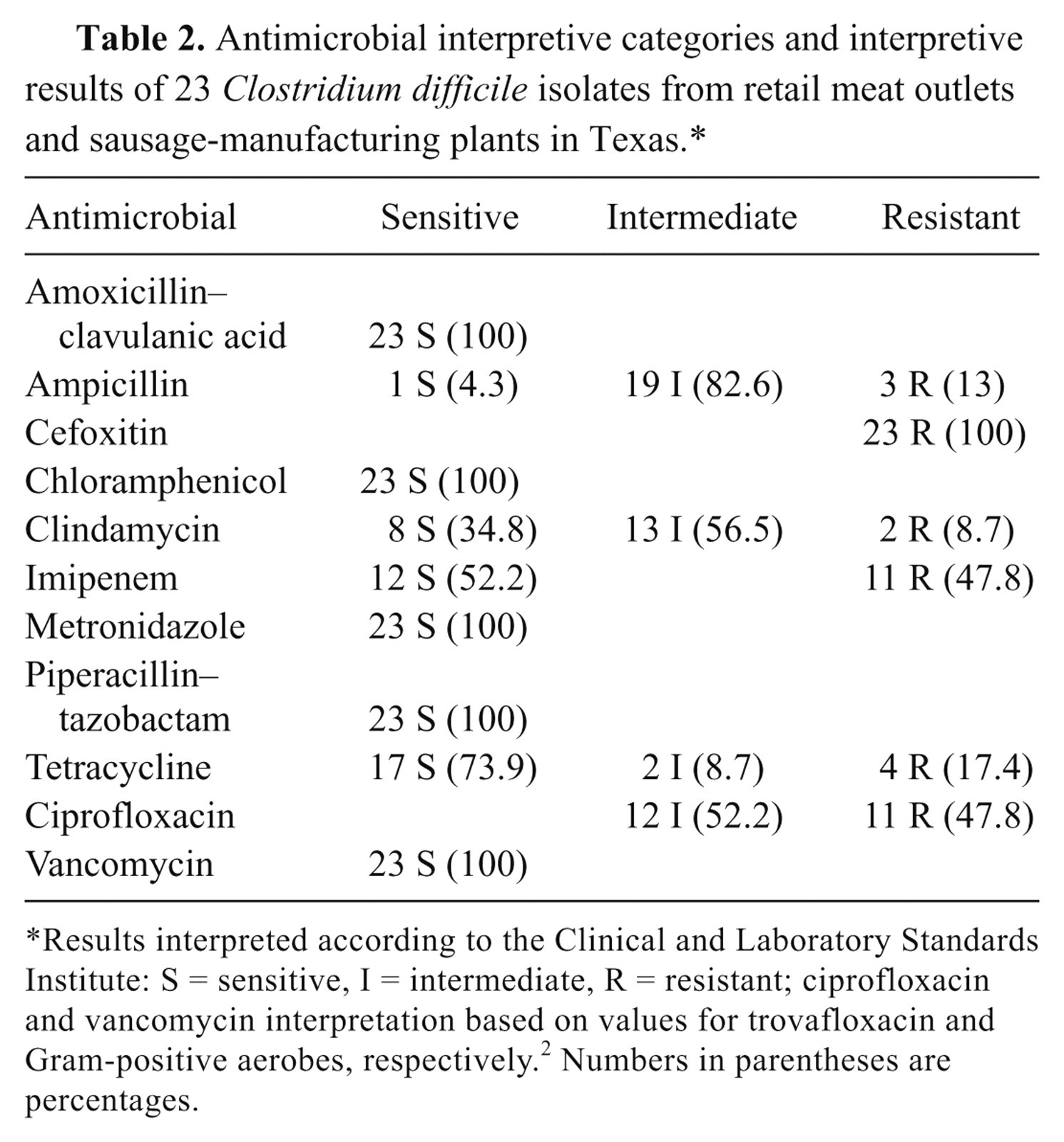
Results interpreted according to the Clinical and Laboratory Standards Institute: S = sensitive, I = intermediate, R = resistant; ciprofloxacin and vancomycin interpretation based on values for trovafloxacin and Gram-positive aerobes, respectively. 2 Numbers in parentheses are percentages.
The susceptibility patterns of the C. difficile isolates to 11 antimicrobial agents showed 100% sensitivity to amoxicillin–clavulanic acid, chloramphenicol, metronidazole, piperacillin–tazobactam, and vancomycin, whereas 100% were resistant to cefoxitin and chloramphenicol, 83% were intermediate to ampicillin, and 56% were intermediate to clindamycin. The greatest variability in the sensitivity data were for imipenem, ciprofloxacin, and tetracycline. These results are somewhat in contrast to a study in which C. difficile isolates from ground beef were 100% sensitive to vancomycin and metronidazole, but 100% resistant to levofloxacin and clindamycin. 21 However, in that study, isolates were primarily toxinotype III compared to toxinotype V in the current study. In other studies, toxinotype V isolates from meat 24 were sensitive to levofloxacin, moxifloxacin, and gatifloxacin, and 56% resistant and 41% intermediate to clindamycin compared to low-level (8.7%) resistance to clindamycin and moderate (47.8%) resistance to ciprofloxacin in the present study. Antibiotic susceptibility patterns of C. difficile toxinotype V isolates have been reported as 88% sensitive to clindamycin for bovine isolates, 0% sensitive for porcine isolates, and 9% sensitive for human isolates, but 100% of toxinotype V isolates, regardless of origin, were sensitive to gatifloxacin and moxifloxacin. 8 Antimicrobial susceptibility data were not included in the above-mentioned European meat studies (Bouttier S, et al.: 2008, Screening for Clostridium difficile in meat from French retailers). 9,25
In the current study, the authors conclude that sausage-processing plants and retail meat outlets in Texas exhibit a level of contamination with C. difficile that is comparable to that reported for North America, that the predominant isolate recovered was toxinotype V, PFGE type NAP7, and the antimicrobial susceptibility patterns of these isolates suggested somewhat reduced resistance than reported for other meat, animal, or human toxinotype V isolates.
Footnotes
Acknowledgements
The authors wish to thank Ms. Angela Thompson and Dr. Duncan MacCannell, Centers for Disease Control and Prevention, Atlanta, Georgia, for their excellent technical assistance with PFGE protocols. Mandatory Disclaimer: “Proprietary or brand names are necessary to report factually on available data; however, the USDA neither guarantees nor warrants the standard of the product, and the use of the name by the USDA implies no approval of the product, and/or exclusion of others that may be suitable.”
a.
HydraSponge®, 3M, St. Paul, MN.
b.
Oxoid Ltd., Cambridge, United Kingdom.
c.
CCFA, Anaerobe Systems, Walnut Hill, CA.
d.
Pro Disc, Remel Inc., Lenexa, KS.
e.
Rapid ID 32A, bioMérieux Inc., Durham, NC.
f.
QIAamp®, Qiagen Inc., Valencia, CA.
g.
New England Biolabs Inc., Ipswich, MA.
h.
Etest, AB Biodisk North America Inc., Piscataway, NJ.
i.
CHEF Mapper®, Bio-Rad Laboratories, Hercules, CA.
j.
Bionumerics software, Applied Maths, Austin, TX.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was funded in part by The National Pork Board, Grant no. 06-156 and USDA-CSREES-NRICGP, no. 2003-35212-13298.
