Abstract
Avian bornavirus (ABV) is the causative agent of proventricular dilatation disease (PDD), a highly devastating and contagious disease of psittacines (parrots and parakeets), which has resulted in the death of many captive birds. Accurate diagnosis of bornavirus infection is therefore important for the identification and isolation of infected birds. The current study showed that nonvascular contour (chest) feather calami provide a ready and minimally invasive source of RNA for the detection of ABV by reverse transcription polymerase chain reaction (RT-PCR). Storage of the feathers at room temperature for at least a month did not affect the results. Serological analysis by enzyme-linked immunosorbent assay (ELISA) showed that identification of anti-bornaviral nucleoprotein P40 antibodies can identify many birds with a past or present infection. The presence of anti-avian bornaviral P24 phosphoprotein and P16 matrix protein antibodies was quite variable, rendering these antibodies less useful for diagnosis of ABV infection. The significance of the present findings is that the use of nonvascular feathers as a source of RNA allows sample collection under conditions where storage of other samples would be difficult. Serum detection by ELISA of anti-P40 antibodies allows the identification of infected birds when RT-PCR fails.
Keywords
Introduction
Proventricular dilatation disease (PDD) is a severe, often fatal disorder in a wide variety of psittacine birds, such as cockatoos, macaws, amazons, and African grey parrots. 12,15,19,21,33,34 Although a viral etiology had long been proposed for the disease, 14 only recently have advanced molecular techniques identified a bornavirus (Avian bornavirus [ABV]) as the causative agent of PDD. The disease affects the central nervous and the digestive systems. The classical diagnosis of PDD is based on crop biopsy 16 to detect lymphoplasmacytic infiltrates, the presence of undigested seeds in the feces of affected birds, as well as on other, mainly neurological signs, such as ataxia. The recent identification of ABV as the cause of PDD 19,21 has resulted in the need for diagnostic tests based on the presence of bornaviral RNA, protein, and anti-bornaviral antibodies for identifying birds infected with ABV. 9,23,29 Since the disease is considered to be a major threat to captive psittaciform populations, accurate diagnosis using non- or minimally invasive procedures is important in order to identify infected birds and isolate them from uninfected individuals.
In mammals, bornavirus is a non-cytolytic, enveloped, neurotropic virus. In birds, the virus can be detected in many tissues, including the central nervous system. 9,23,29,33
The genome of ABV consists of a nonsegmented negative single-strand RNA component of approximately 9,000 nt. Due to its recent discovery, ABV has not been fully characterized. However, nucleotide sequence comparison of the ABV genome and its mammalian counterpart 4,19,21 suggests that except for their difference in tropism, a high degree of similarity exists between the 2 viruses. In a previous study, 9 serum samples from birds diagnosed with PDD were shown to contain antibodies against the P-40 nucleoprotein of ABV, making serological testing 3 a method for identifying ABV-exposed birds. The current investigation aims to extend the previous findings to 2 additional bornaviral proteins, the P24 and P16 proteins, with a view to determining whether other bornaviral proteins might be better suited for serological diagnosis of bornaviral infection. Another objective of the study was to discover if plucked nonvascular feathers can be used as alternative source of viral RNA for reverse transcription polymerase reaction (RT-PCR) assays in place of fecal material or cloacal swabs, thus avoiding problems associated with the presence of nonspecific inhibitors usually present in fecal matter or the relatively invasive procedure of collecting cloacal swabs.
Materials and methods
Source of materials
Feathers, cloacal swabs, and blood were from birds maintained in the exotic bird collection of the senior author (de Kloet). Approximately 200 µl of blood was obtained by brachial vein puncture and collected in ethylenediamine tetra-acetic acid (EDTA)-containing serum collection tubes. a Forty-four birds were obtained in January 2010 as the survivors from a collection that had experienced serious infections of ABV in which more than half (105/198) of the 6–12-month-old birds had died with PDD-like clinical signs (undigested seeds in their fecal matter and neurological signs such as ataxia). The birds were housed in a 7 m (l) × 2.5 m (w) × 2 m (h) outdoor aviary with a soil floor. The 44 birds consisted of 11 green-cheeked conures (Pyrrhura molinae), 2 African grey parrots (Psittacus erithacus), 2 golden-capped conures (A. auricapilla), and 29 Aratinga species consisting of sun conures (Aratinga solstitialis) and jenday conures (A. jandaya) or their hybrids. The close similarity of the sun and the jenday conures 32 made accurate determination of their specific numbers impossible. The birds were full-winged and able to fly. They were all free of any outward clinical signs of PDD. The birds were maintained on a diet of equal parts of peanuts, sunflower seed, a wild birdseed mixture, b and a vitamin-enriched parakeet seed mix, c as well as apples, oranges, and bananas for a total of 3 kg per day for the entire group of birds. Fresh drinking water was continuously provided.
Psittaciformes that were hatched in the collection and had never been outside the collection consisted of 11 African grey parrots (P. erithacus), a chattering lory (Lorius garrulous), and a sulphur-crested cockatoo (Cacatua sulphurea) as well as 52 peach-faced lovebirds (LB, Agapornis roseicollis). These birds were housed in aviaries at least 50 m away from the aviary with the conures, and were fed the same diet and were not infected with bornavirus. 9
RNA isolation from feathers, cloacal swabs, and blood
For the present investigation, feathers, cloacal swabs, and blood were collected 2 months after the acquisition of the birds, in order to ensure that they had fully acclimatized to their new environment. Nonvascular contour (ventral) feathers were collected by plucking and either processed directly or air-dried at 37°C for 3 hr, and stored for up to 4 weeks at room temperature in plastic bags. For the extraction of RNA, the approximately 3-mm long calami of 5 plucked feathers were cut off, incubated overnight at 25°C with shaking in 500 µl of 5 M guanidinium isothiocyanate, 0.5% sarcosyl, 25 mM sodium citrate, and 0.1 M mercaptoethanol (pH 7, solution D 5 ) and subsequently homogenized in a 1.5-ml Eppendorf tube with a Teflon pestle as described previously for brain tissue. 9 After the consecutive addition and mixing of 100 µl of 2 M sodium acetate buffer (pH 4.1), 500 µl of 90% phenol, and 100 µl of chloroform, the mixture was subjected to 15 min of centrifugation at 10,000 × g in a microfuge, as previously described. 5 The aqueous supernatant was mixed with an equal volume of ethanol, and the RNA was purified by centrifugal spin column chromatography. d After washing twice with 500 µl of 70% ethanol containing 0.15 M potassium acetate and 0.1 mM EDTA (pH 8.0), the columns were dried by an extra centrifugation step of 5 min at 10,000 × g and the RNA eluted with 20 µl of sterile water. For RNA isolation from feather vanes, the same extraction procedure was applied as described for the feather calami: a 4 mm × 4 mm fragment, cut from the center of the vane, was used. Blood and cloacal swab samples were processed within 3 hr after collection. White blood cells were isolated from 200 µl of whole blood by a scaled-down version of a method that utilizes centrifugation through 1% methylcellulose. 35 RNA was isolated from the partially purified white blood cells by acid guanidinium thiocyanate-phenol extraction 5,9 and spin column chromatography. RNA was isolated from cloacal swab and whole blood by a modified acid guanidinium isothiocyanate procedure 5 as described previously. 5,9 RNA samples were stored at −80°C.
Reverse transcription polymerase chain reaction
Amplification of bornaviral RNA by RT-PCR using the previously described M and N primers 21 was carried out in a water-cooled thermocycler e as described previously. 9 The M primers amplify a 360-bp segment of the bornaviral P16 matrix protein gene. The N primers amplify a 395-bp segment of the bornavirus P40 nucleoprotein gene. For routine RT-PCR, 1 µl of RNA extract was incubated in 20 µl of 20 mM Tris-HCl buffer (pH 8.8), 10 mM KCl, 10 mM (NH4)2SO4, 2 mM MgCl2, 0.1% Triton-X100, 5% dimethyl sulfoxide, 5 mM of dithiothreitol, 15 µg of bovine serum albumin, 10 0.01% bromocresol red as tracking dye, 0.5 µM of primers and 0.5 U Taq DNA polymerase, f 2.5 U of AMV reverse transcriptase, f and 4 U of RNase inhibitor. f Thermocycling conditions were as follows: after 30 min of incubation at 45°C, the samples were heated for 2 min at 94°C followed by 30 cycles 20 of 20 sec at 92°C, 30 sec at 55°C, and 80 sec at 68°C, followed by a final extension of 10 min at 72°C. The amplification products were analyzed by agarose gel electrophoresis using 1.5% gels in Tris–borate–phosphate buffer (pH 8.4), containing 0.1 µg of ethidium bromide per ml. 24 Control RT-PCR for successful extraction of RNA was carried out with the G primers GAPDHF and GAPDHR, which amplify a segment of the glyceraldehyde 3′-phosphate dehydrogenase gene 21 and by the R primers Rib18F and Rib18R, which amplify a highly conserved 37 204-bp segment of avian ribosomal 18S RNA. 13,42 The sequences of the primers are given in Table 1.
Target genes, recombinant proteins synthesized, and names and sequences of the primers used.*
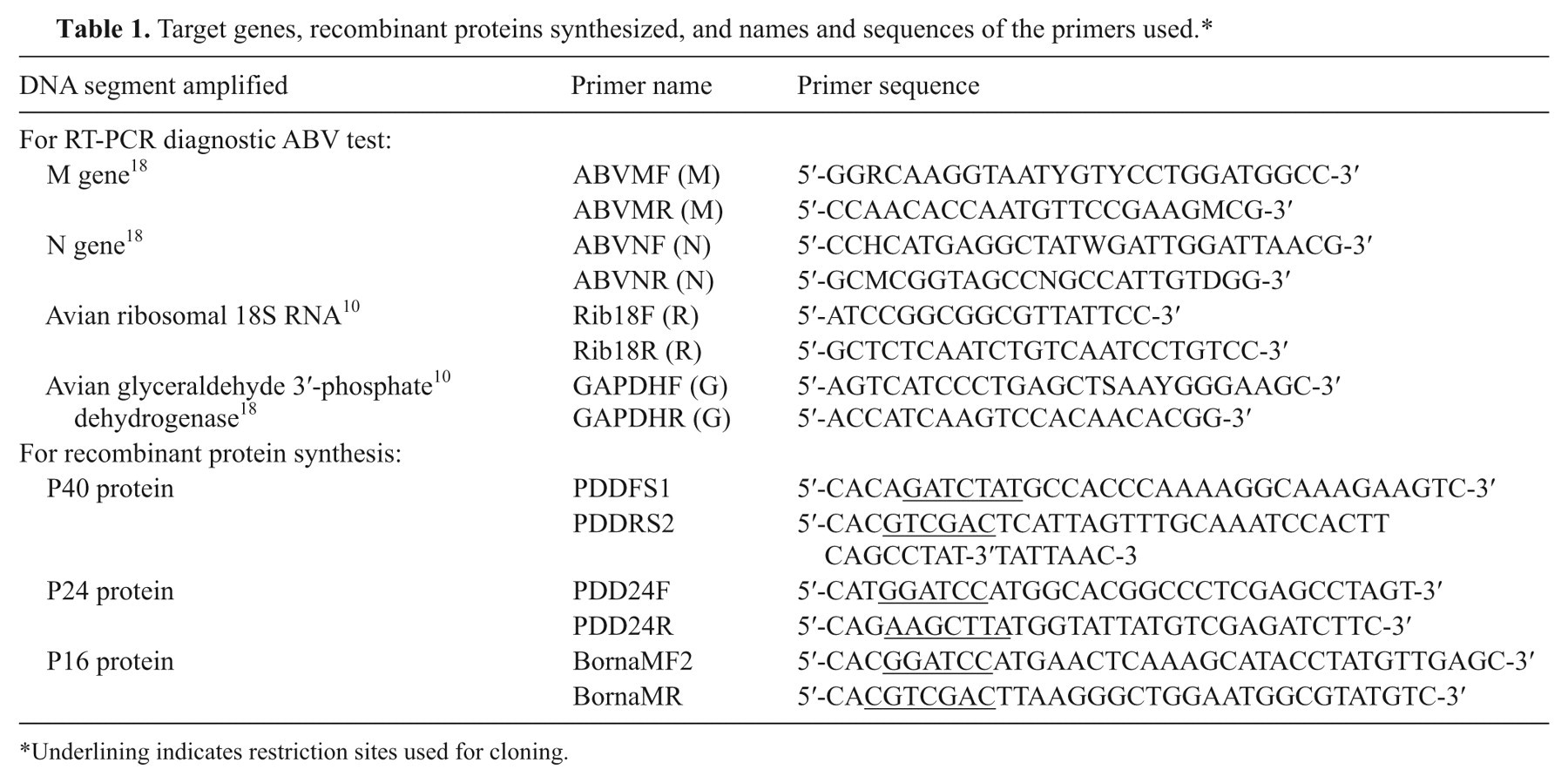

Underlining indicates restriction sites used for cloning.
Plasma samples
Twenty-five µl of blood, obtained by brachial venipuncture, was collected in EDTA-coated capillaries g and diluted 10-fold in 250 µl of buffer A (0.15 M NaCl, 0.025 M EDTA [pH 8.0]). After low-speed centrifugation (5 min at 500 × g), the supernatant was used as the 20-fold diluted plasma sample, assuming that plasma amounts to 50% of the blood volume. Diluted plasma samples were stored at −20°C until analysis (1–30 days after collection).
Enzyme-linked immunosorbent assay for antibodies against Avian bornavirus–specific P40, P24, and P16 proteins
Avian bornavirus recombinant P40, P24, and P16 proteins were produced in Escherichia coli as a fusion product with the maltose-binding protein (MBP) using a modified 22 pMAL system. 9,f The modification consisted of the introduction of a thrombin site for cost-effective separation of the MBP fusion partner from the recombinant protein. The pMal system was chosen because of the superior solubilization properties of MBP on its fusion partner. f Amplification of DNA fragments coding for the P40, P24, and P16 proteins was carried out with the primers PDDFS1 x PDDRS2, PDD24F x PDD24R, and BORNAMF2 x BORNAMR, respectively (Table 1), which were derived from the strain with GenBank accession no. EU781967. Expression and purification of the recombinant proteins was carried out by amylose affinity chromatography using the protocols provided by the manufacturer. f The sequences of the genes of the proteins have been entered into the database with GenBank accession nos. FJ792853 (P40), HM369502 (P24), and HM369503 (P16).
Enzyme-linked immunosorbent assay (ELISA) was performed as described previously. 9 One µg of recombinant fusion protein in 50 µl of 0.05 M sodium carbonate (pH 9.5) was applied to the wells of microtiter plates, g incubated for 1 hr at room temperature, washed with phosphate buffered saline (PBS) containing 0.05% Tween 20 (PBST), and blocked overnight at 4°C in 5% fat-free dried milk h in PBST, and washed again with PBST. One µg MBP was applied to wells as a control for nonspecific binding of antibodies. For routine ELISA, 50 µl of a 1:800 dilution of serum in 5% milk in PBST was added to the wells, incubated with shaking for 1 hr, washed again with PBST, followed by incubation for 1 hr with 50 µl of a 4,000-fold dilution of goat-anti-bird antibody conjugated with horseradish peroxidase i in 5% milk in PBST. After washing with PBST, 50 µl of 0.05% 3,3′,5,5′-tetramethylbenzidine (TMB1) j was added, and color development was allowed for 2 min at room temperature (25°C) and the reaction stopped with 50 µl of 0.1 N HCl. Absorbance at 450 nm was read in a microplate reader. k Antibody titers were determined from 2-fold serial dilutions of the plasma samples and represent the reciprocal of the highest dilution that still gave a color change as compared to serum of noninfected birds. 30 As cut-off value for identifying positive sera a value of 3 standard deviations above the mean absorbance value of 8 sera from the group of 8 PDD-negative African grey parrots was taken. 6 Western blotting was performed as described previously 9 except that the TMB1-stabilized horseradish peroxidase procedure was used for detection instead of alkaline phosphatase–based chemiluminescence.
Results
Serological analysis
Table 2 shows the results obtained by serological analysis by ELISA of serum samples for the presence of anti-P40, -P24, and -P16 antibodies of the birds tested in comparison with the results obtained by RT-PCR analysis of feather calami. Figure 1 shows the ELISA profiles of the sera of a number of birds used in the current study. Separate studies by Western blotting (data not shown) using thrombin-digested fusion proteins demonstrated that the sera reacted with proteins with a molecular size of 40, 23, and/or 17 kb, the approximate molecular sizes of the P40, P24, and P16 proteins, respectively, confirming the identity of the proteins used in the ELISA.
Comparison of indirect enzyme-linked immunosorbent assay (ELISA) for detection of antibodies against the P40, P24, and P16 proteins in sera versus detection of M- and N-genes using reverse transcription polymerase chain reaction (RT-PCR) from feather RNA samples of the psittaciformes used in the present study.*
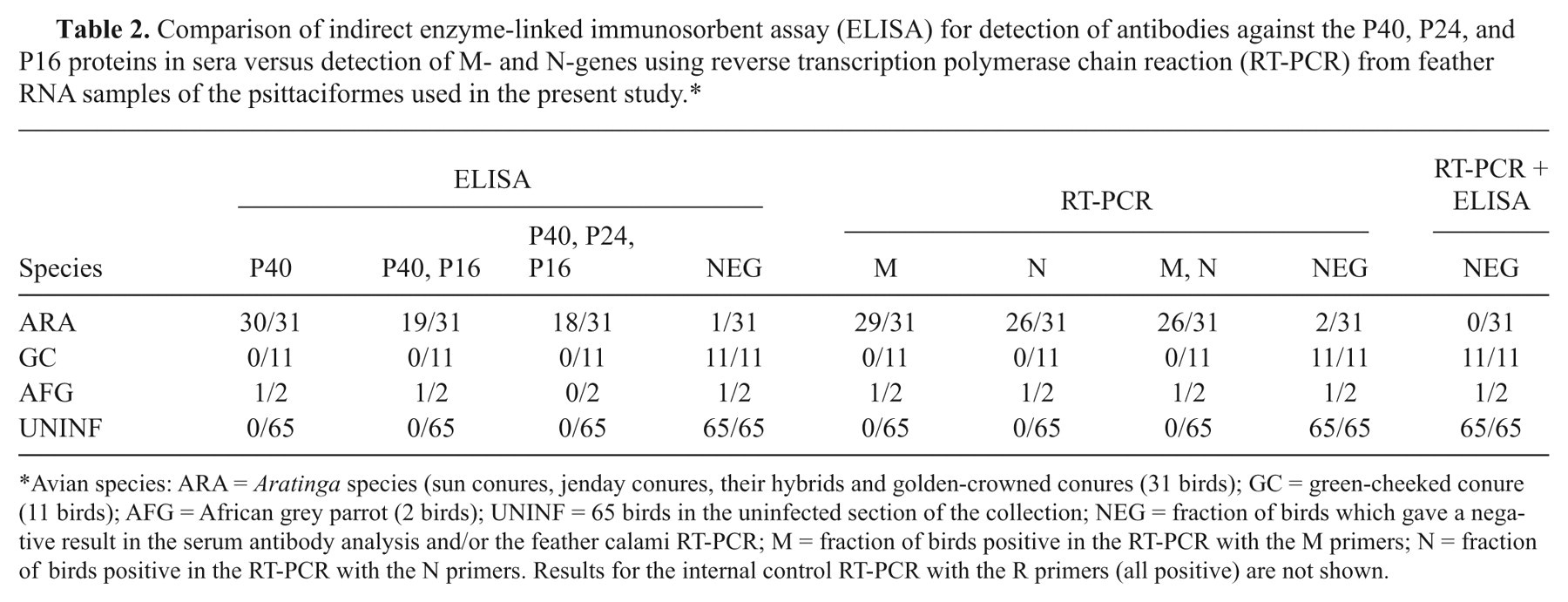
Avian species: ARA = Aratinga species (sun conures, jenday conures, their hybrids and golden-crowned conures (31 birds); GC = green-cheeked conure (11 birds); AFG = African grey parrot (2 birds); UNINF = 65 birds in the uninfected section of the collection; NEG = fraction of birds which gave a negative result in the serum antibody analysis and/or the feather calami RT-PCR; M = fraction of birds positive in the RT-PCR with the M primers; N = fraction of birds positive in the RT-PCR with the N primers. Results for the internal control RT-PCR with the R primers (all positive) are not shown.
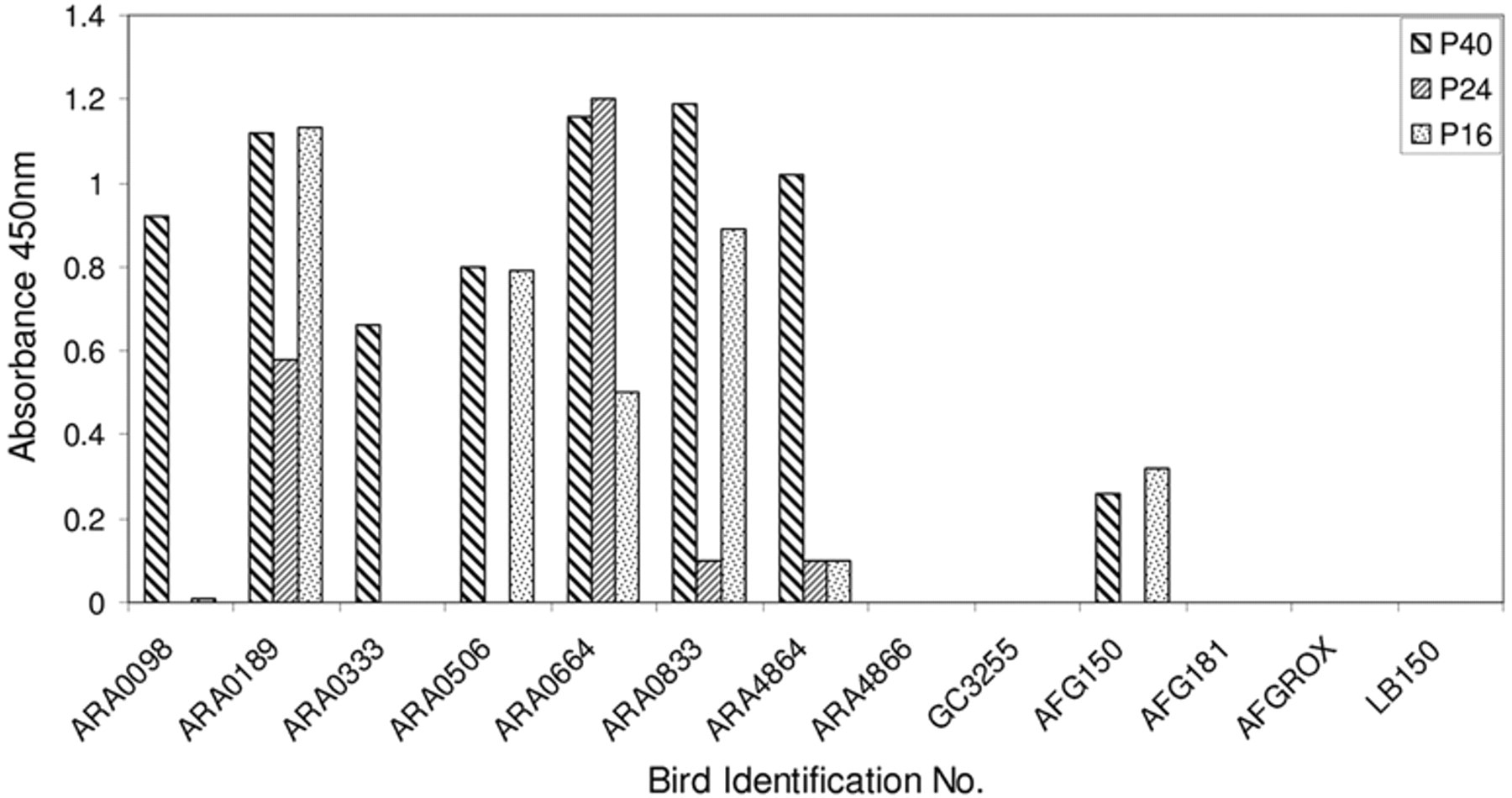
Variability of the anti-bornaviral P40, P24, and P16 protein antibody content measured by enzyme-linked immunosorbent assay in sera of bornavirus-exposed psittaciformes. Values on the vertical axis are absorbance readings at 450 nm in the 3,3′,5,5′-tetramethylbenzidine analysis, corrected for nonspecific binding to maltose-binding protein protein. AFGRox and LB150 are control birds from the unexposed section of the collection.
The results show that almost all serum samples of the Aratinga species (30/31) obtained from the collection with PDD contained measurable amounts of antibodies against the P40 protein. The sera of 11 of these birds contained only antibodies against the P40 protein (shown in Fig. 1 for ARA0098 and ARA0333), 19 birds also contained variable amounts of antibodies against the phosphoprotein P24 and the matrix protein P16 (shown in Fig. 1 for ARA0189, ARA0664, ARA0833, and ARA4864), with the exception of 1 bird (Fig. 1, ARA0506), which had no measurable antibodies against P24. Only 1 bird (Fig. 1, ARA4866) was seronegative for avian bornaviral P40, P24, and P16 protein. The serum samples of the 11 green-cheeked conures, although from the same infected facility and housed in the same aviary as the bornavirus-infected Aratinga spp. did not show the presence of antibodies against the P40, P24, or P16 proteins (shown in Fig. 1 for GC3255). Serum from 1 of the exposed African grey parrots (Fig. 1, AFG150) contained antibodies against the P40 and P16 proteins, whereas the other (Fig. 1, AFG181) did not. None of the sera from the 65 control birds kept isolated from the Aratinga spp contained antibodies against the P40, P24, or P16 proteins (shown in Fig. 1 for the African grey parrot AFGROX and the peach-faced lovebird LB150).
Titer measurements for some of the birds showed that the antibody titer could be as high as 80,000 for antibodies to the P40 protein, 10,000 for antibodies to the P24 protein, and 6,000 for the antibodies to the P16 protein and that the relative titers of the antibodies against the P40, P24 and P16 proteins showed a wide variation among individual sera, some had anti-P40 titers of about 2,400, whereas there were no measurable antibodies to P24 or P16.
RT-PCR analysis of feather RNA
Under the RT-PCR conditions described herein, the maximum sensitivity for the detection of ABV with the M and N primers was approximately 3,000 molecules of viral RNA as judged by the visibility of the amplification product in the electrophoretic analysis and staining with ethidium bromide and using RNA transcribed in vitro from a cloned M or N fragment as a standard (data not shown). Figure 2 shows a comparison of the results of an RT-PCR analysis with the N, M, G, and R primers of RNA prepared from cloacal swab, feather, and blood samples of 2 sun conures. The results show that whereas the feather calami (Fig. 2a) as well as swab samples (Fig. 2b) from bird ARA0664 produced amplification products with the 4 primer pairs; with bird ARA4864, the feather calamus RNA sample (Fig. 2d) produced amplification products with all 4 primer pairs, but the swab sample (Fig. 2e) did not, suggesting that the presence of RT-PCR inhibitors or the absence of RNA in the swab samples was responsible for the negative results. Similar results were found with the other Aratinga spp. (A. solstitialis, A. jandaya, and A. auricapilla): the feather calami always produced RT-PCR products, whereas the results obtained with the swab samples were quite variable. As was described previously, 9 RNA isolated from blood (Fig. 2c, 2f) or partially purified white blood cells (data not shown) did not produce amplification products with the M or N primers, although the G and R primers did, demonstrating that failure of the RT-PCR of the blood sample with the M and N primers was not caused by the presence of RT-PCR inhibitors or the absence of RNA. Separate experiments (data not shown) demonstrated that the vane of the feathers was negative in the M and N-primer RT-PCR of all birds. Further experiments showed that storage of the dried feathers of eight of the sun conures for at least 4 weeks at ambient temperature (20–25°C) did not affect the results, in agreement with observations 4 that dried feathers form a very stable source of RNA (shown for bird ARA0833, Fig. 2g, 2i). No amplification products are formed with the M and N primers when feather extracted RNA of bird GC3255, one of the green-cheeked conures, was used, whereas the G and M primers did, demonstrating that RNA isolation was successful (Fig. 2h). In the current investigations it was also found that the G primers gave variable amounts of product when feather calamus RNA was used. Therefore, a ribosomal RNA fragment has been used as an additional internal control for a successful RT-PCR. 13
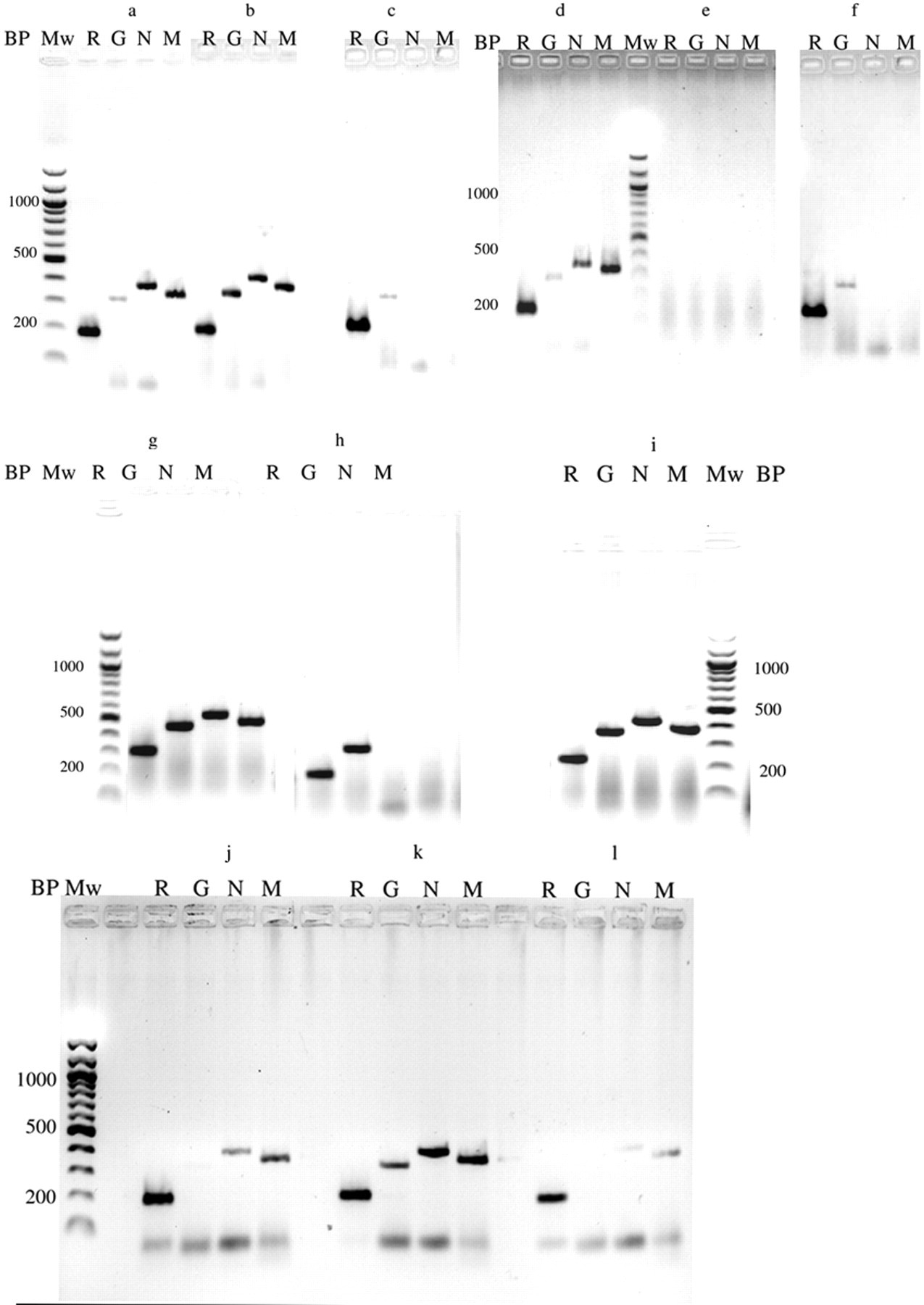
Gel-electrophoretic analysis of the amplification of Avian bornavirus–specific RNA from feather calami, cloacal swabs, and blood of selected conures. Lane M: reverse transcription polymerase chain reaction (RT-PCR) with the bornaviral M primers; lane N: RT-PCR with the N primers; lane R: control RT-PCR with the ribosomal RNA primers; lane G: control RT-PCR with the G primers; Mw = 100-bp ladder; BP = size of the amplified fragment in base pairs.
Initial investigations also found that the RT-PCR with M primers sometimes produced positive results, when the N primers produced no or very low amounts of amplification product. A further investigation into the optimization of the RT-PCR conditions showed that reducing the amount of feather calamus extract often led to an increase in the amount of product formed in the N primer–specific RT-PCR, suggesting the presence of RT-PCR inhibitors or that excess template, giving rise to linear instead of exponential amplification was responsible for this result. 20 This is demonstrated in Figure 3a for bird ARA0664. The results show clearly that a 4-fold reduction (1 µl instead of 4 µl of feather extract in a 20-µl PCR) in the amount of template RNA leads to a considerable increase in the amount of product formed when the N primers are used, whereas no difference is observed with the M primers. Further reduction of the amount of template RNA (Fig. 3b) shows that in bird ARA0833, a 64-fold reduction in the amount of feather calamus extract still yields a visible amount of PCR product by agarose gel electrophoresis. Since the RNA in these experiments is routinely extracted from 5 feather calami, these data suggest that in such cases the RNA extracted from a single feather calamus may be sufficient to detect the presence of bornaviral RNA in infected birds. This is shown in Figure 2j, 2k, and 2l where single feathers of bird ARA0333 were used in the analysis.
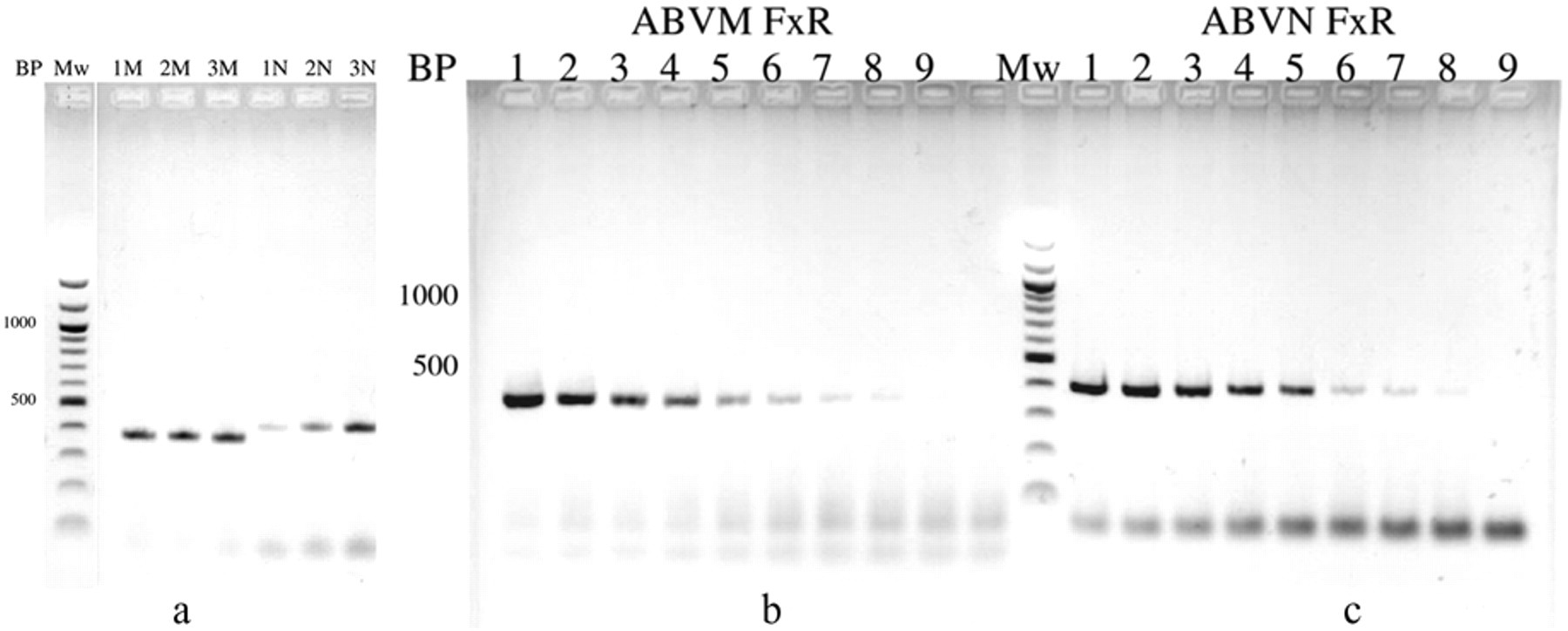
Effect of sample dilution on the formation of amplification products of the reverse transcription polymerase chain reaction (RT-PCR) with the M and the N primers. Mw = 100-bp ladder; BP = size of the amplified fragment in base pairs; lane M: RT-PCR with the M primers; lane N: RT-PCR with the N primers.
Table 2 shows that 29 of the 31 Aratinga sp. werebornavirus-positive by M and/or N primer RT-PCR of the feather calami. All samples were positive for the control ribosomal RNA RT-PCR, demonstrating that in all samples, RNA extraction and amplification was successful. One bird (ARA4866) that was negative in the serological test (Fig. 1) was positive in the feather RNA RT-PCR with the M and N primers. One African grey parrot (AFG150) was weakly positive in the M and N primer–specific RT-PCR, whereas the other (AFG181) was negative. The 11 green-cheeked conures were all negative in the feather RT-PCR with the M or N primers. All control birds from the noninfected section of the collection were negative in the M or N primer RT-PCR.
Discussion
The results of the current study show that for diagnosis of a current infection with ABV in psittacines, RT-PCR of plucked, contour feathers allows for relatively noninvasive sample collection in live birds. The results add therefore to the list of RNA and DNA viruses which can be detected in bird feathers. 1,8,17,18,26,40 In addition to being relatively noninvasive an additional advantage of using feathers is that avian bornaviral RNA remains stable for at least 4 weeks when stored at room temperature, whereas most other tissues require storage at temperatures below freezing. Although single feathers may be sufficient for the detection of bornaviral RNA, positive results are not consistently obtained from single feathers. Whether this is the result of differences in efficiency of RNA extraction of individual feathers, different amounts of tissue attached to the feather calami or age of the feathers remains to be investigated. The use of multiple feathers or substitution of the relatively insensitive single-step RT-PCR by more advanced real-time procedures may help in avoiding this difficulty.
Although it was not possible to detect ABV RNA in blood or partially purified white blood cells using the one-step RT-PCR procedure in the present study, it is again possible that a more sensitive test (nested RT-PCR or quantitative real-time RT-PCR) might well give positive results. Although cloacal swabs are a widely used source of RNA for pathogen detection in birds, the frequent and unpredictable presence of RT-PCR inhibitors in fecal material 7,11 and consequently in cloacal swabs suggests that cloacal swabs are an unreliable source of diagnostic RNA. Procedures for the removal of such inhibitors are known, but do not apparently remove the inhibitors from all cloacal swab samples. 7
The sensitivity of the RT-PCR is dependent on many sequence characteristics of the template RNA and can range from primer incompatibility to simple point mutations anywhere in the amplicon. 2 The presence of RT-PCR inhibitors such as melanin, 10 a common element in feather coloration of birds and their ancestors, 41 may also lead to false negative results. This is unlikely in the present investigation, because this inhibition can be overcome by use of bovine serum albumin, which was routinely added to the PCR 10 in the present study. However, the presence of other PCR inhibitors in feathers cannot be excluded. Finally, excess template RNA, which includes viral genomic RNA as well as mRNA, resulting in linear instead of exponential amplification 20 may also be responsible for the initially poor results with the N primers as compared to the M primers. The finding that sample dilution 20 resulted in relative better amplification with the N as compared to the M primers supports this conclusion. Variations in the amount of amplified product may also reflect differences in gene expression and consequently different amounts of messenger RNA. Relative differences in the expression of bornaviral genes has been observed in rats experimentally infected with bornavirus where the expression of the P57 gene was severely repressed as compared to the expression of the P40 nucleoprotein gene. 31,39 Whether a similar differential expression of avian bornaviral P40 and P16 occurs is unknown. No bornaviral RNA could be detected in the vane of the feathers, an observation that contrasts with the finding for West Nile virus in the feathers of crows. 26
The data in the present study show that, similar to what has been found with bornavirus infection in mammals, 25,27,28 there is considerable variation in the nature and titer of antibodies formed in psittacines naturally infected by bornavirus. The varying antibody titers for different bornaviral proteins may reflect different immunological responses of individual birds to the proteins, but different relative amounts of bornaviral proteins in the infected birds giving rise to different antibody titers cannot be excluded. Although no studies have been done on the differential expression of avian bornaviral genes, studies in mammals have shown 38 that the molecular ratio of the P40 and P24 proteins can differ considerably in persistently and acutely infected cells and tissues. No anti-bornavirus antibodies could be detected in ARA4866, whereas this bird contained bornaviral RNA in the feather calamus M- and N-primer–specific RT-PCR test. These findings may bear some relationship with findings in sheep and cats where variable levels of anti-P40 and -P24 antibodies as well as animals with Borna disease without the presence of anti-bornaviral antibodies have been described. 25,27,28 Finally, the presence of birds without evidence of bornavirus infection, such as the 11 green-cheeked conures in a collection of heavily infected psittaciformes, suggests that there may be variation in susceptibility to bornavirus infection among different, even closely related, psittacine species. However, little is currently known about the role of various environmental factors in ABV infection, such as effects of humidity, temperature etc. on the stability of the virus outside the host, and it may well be that under the conditions at which the birds are housed and maintained, no active, infectious virus is present in the environment. Finally, an additional finding in these studies was that the GAPDH gene, a housekeeping gene widely used as an internal control for RT-PCR investigations, may not be suitable for feather RNA analysis, consequently ribosomal RNA has been used as a second internal standard. 13,36,42
In conclusion, for the diagnosis by RT-PCR of a current ABV infection in parrots and parakeets feather calami form an excellent source of RNA. However, when the RT-PCR fails to detect ABV infection, serological detection of anti-P40 nucleoprotein (and other bornaviral protein) antibodies can also identify a past and/or present exposure. The converse is also true, if the ELISA fails to detect antibodies against bornaviral protein, the feather RT-PCR test may still detect an infection as was shown for one of the birds. Since avian bornaviral RNA can be detected in fecal material, 9 a fecal–oral mode of transmission of bornavirus between birds has been considered most likely. The present data suggest that feathers could also contribute to the spread of the virus, as has been found for Chicken anemia virus. 8
Footnotes
Acknowledgements
The authors are indebted to Dineke de Kloet for her help in collecting the feather, cloacal swab, and serum samples for this study. Dr. Steven Miller is thanked for his help in sequencing the P40, P24, and P16 genes.
a.
Becton Dickinson, Franklin Lakes, NJ.
b.
Johnson Seed Cy, Ashburn, GA.
c.
Provita Parakeet, OK Feeds, Miami, FL.
d.
Zymo-spin-TM1, Zymo Research, Orange, CA.
e.
Ericom, San Diego, CA.
f.
New England Biolabs, Ipswich, MA.
g.
Sarstedt, Newton, NC.
h.
Publix Super Markets Inc., Lakeland, FL.
i.
Cat# A140-110P, Bethyl Laboratories, Montgomery, TX.
j.
Promega, Madison, WI.
k.
Tecan, Mannedorf, Switzerland.
The authors declared that they had no conflicts of interest with respect to their authorship or the publication of this article.
The authors declared that they received no financial support for their research and/or authorship of this article.
