Abstract
Highly pathogenic avian influenza (HPAI) A(H5N1) clade 2.3.4.4b virus was identified in 4 cull dairy cows condemned by the U.S. Department of Agriculture because of pneumonia with accompanying systemic changes. Histologic findings were bronchopneumonia in 3 cows and embolic pneumonia and nephritis in 1 cow. In addition to detection of HPAI A(H5N1) virus by reverse-transcription real-time PCR in various formalin-fixed, paraffin-embedded tissues, influenza A virus was detected by immunohistochemistry and in situ hybridization in the pulmonary respiratory epithelium of 2 of the cows with bronchopneumonia and in renal medullary tubules of the cow with nephritis. A PCR panel screening for common bovine respiratory pathogens in the cows with bronchopneumonia revealed variable coinfections with Histophilus somni, Pasteurella multocida, Mannheimia haemolytica, Mycoplasmopsis bovis, and bovine coronavirus. We describe the distribution of HPAI A(H5N1) virus in naturally infected cows while highlighting the need for research on the roles of coinfection and immune response in influenza viral replication.
Since the first report in March 2024 of highly pathogenic avian influenza (
Sialic acids (
We focused on 4 cows brought into federally inspected slaughter establishments with pneumonia accompanied by systemic changes and identified as naturally infected with HPAI A(H5N1) virus via PCR testing. All 4 cows in our study were condemned on postmortem evaluation and thus did not enter the food supply. Of the PCR-positive cows, replication of the influenza virus within respiratory epithelium of 2 cows and within renal tubules of a third cow was demonstrated via in situ hybridization (ISH). Interestingly, concurrent infections with other microorganisms were confirmed via PCR testing and histochemical staining within the affected lungs and kidney, respectively. This raises questions regarding interactions between comorbidities and the host immune response on the ability of HPAI A(H5N1) virus to replicate in bovine tissues.
In April 2024, 3 cull dairy cows originating from a known HPAI A(H5N1) virus-positive herd were condemned by the USDA’s Food Safety and Inspection Service (FSIS) by in-plant personnel because of severe pneumonia and lesions consistent with infectious systemic disease. FSIS, the Animal and Plant Health Inspection Service’s National Veterinary Services Laboratories (NVSL; Ames, IA, USA), and the Iowa State University Veterinary Diagnostic Laboratory (ISU-VDL; Ames, IA, USA) performed ancillary testing to investigate infection with the HPAI A(H5N1) virus. Preliminary HPAI A(H5N1) virus PCR results from these 3 cows led to the implementation of an exploratory study of the distribution of HPAI A(H5N1) virus in tissues of cull dairy cows condemned during inspection. This exploratory study targeted dairy cows with an unknown HPAI A(H5N1) virus herd status that were condemned because of pneumonia, septicemia, or both by Public Health Veterinarians (PHVs) during routine inspection at U.S. slaughter facilities. From 2024 Apr 3 to 2024 May 13, PHVs sampled at least 10 condemned dairy cows at each of 11 targeted high-throughput cull cow slaughter facilities, operating in 8 states (California [2], Idaho, Michigan, Minnesota, Pennsylvania [2], South Dakota, Texas [2], Wisconsin) for a total of 111 cows. Tissue samples from these cows and the 3 preliminary cows mentioned above were submitted in 10% neutral-buffered formalin to the USDA FSIS Eastern Laboratory Pathology Branch. Tissues collected by PHVs included lung, mediastinal lymph node, kidney, liver, spleen, heart, and diaphragm. Samples were processed to generate formalin-fixed, paraffin-embedded (FFPE) tissue blocks for histologic examination at FSIS, PCR testing at NVSL, and immunohistochemistry and ISH at ISU-VDL.
Mammary gland tissue was excluded from our study because the udders were removed upstream in the slaughter process before inspection, as udders are not considered suitable for human consumption and to prevent contamination of the carcass. Each tissue type from the exploratory study was processed in separate cassettes; tissues collected from the initial 3 cull cows were comingled in tissue cassettes ( Table 1 ; cases 1–3). Scrolls (10-µm thick) of FFPE tissues from each animal were submitted to the NVSL for reverse-transcription real-time PCR (RT-rtPCR) for influenza A virus (IAV), as well as primer-probe sets specific for H5, N1, and H5 clade 2.3.4.4b subtyping by RT-rtPCR. All reactions consisted of 40 cycles, with the IAV RT-rtPCR being semiquantitative and utilizing an exogenous internal control to ensure that the PCR performed within an accepted level of efficiency. The other PCR reactions (H5, N1, H5 clade 2.3.4.4b) were run in parallel or serially on the same extraction, and the qualitative results were used to characterize the virus. Of the FFPE tissue from 114 cows (initial 3 cull cows from the Idaho slaughter plant and 111 cows from the follow-up exploratory study) tested by RT-rtPCR for HPAI A(H5N1) virus, nucleic acid was detected in 4 cows for each of the primer-probe sets specific for IAV, H5, N1, and H5 clade 2.3.4.4b (Table 1). HPAI A(H5N1) virus nucleic acid was detected in each of the lung samples. Other tissues tested from the 4 infected cows tended to have higher Ct values compared with those in the lung. One exception was case 2 in which the kidney Ct value for IAV was lower than that of the lung.
RT-rtPCR results (Ct) for influenza A(H5N1) virus (IAV, EA H5 HPAI, IAV N1, H5 2.3.4.4) in formalin-fixed, paraffin-embedded (FFPE) tissue samples from cull dairy cows.
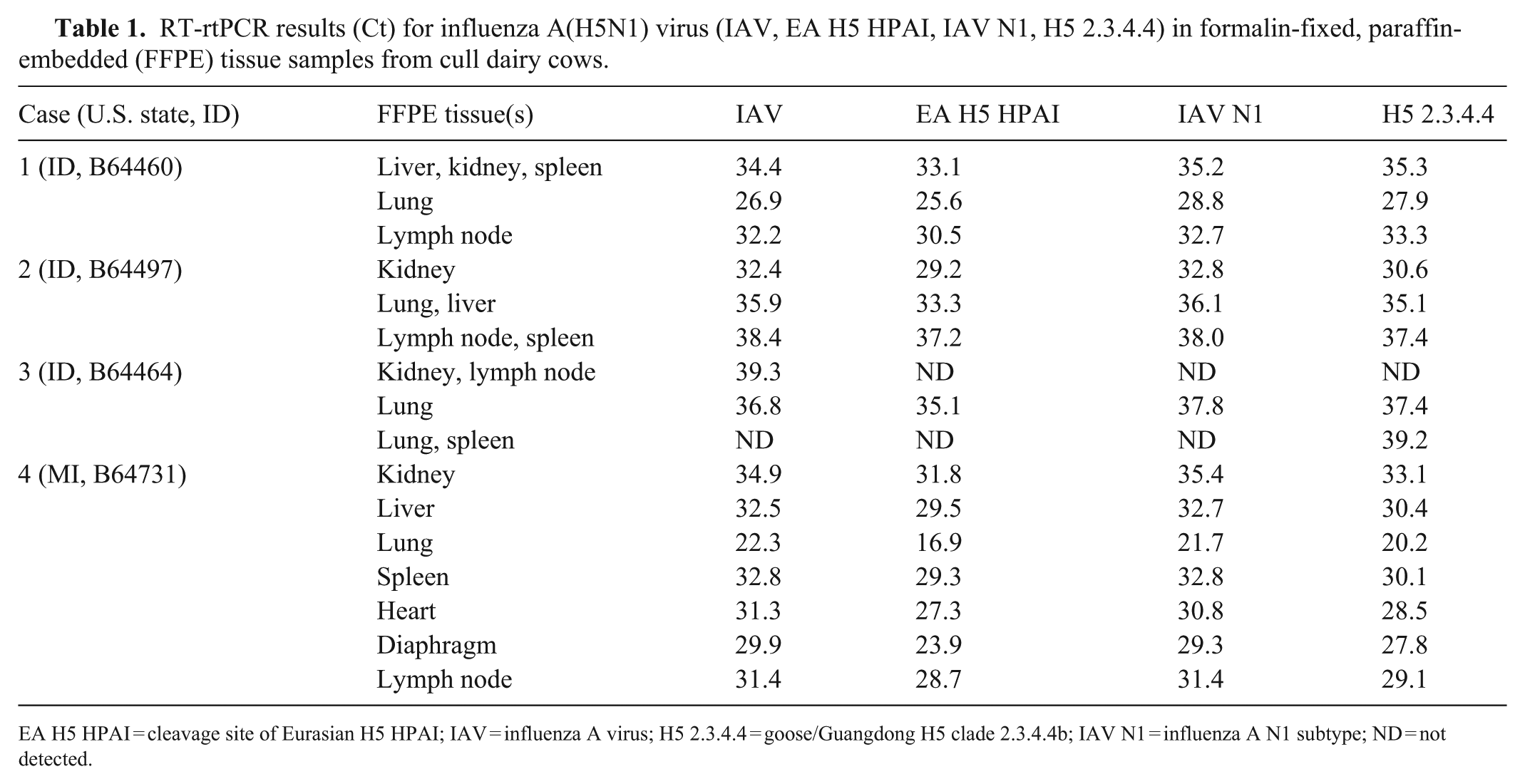
EA H5 HPAI = cleavage site of Eurasian H5 HPAI; IAV = influenza A virus; H5 2.3.4.4 = goose/Guangdong H5 clade 2.3.4.4b; IAV N1 = influenza A N1 subtype; ND = not detected.
Unstained slides from 4 HPAI A(H5N1) virus PCR-positive cows were submitted to the ISU-VDL for IAV nucleoprotein immunohistochemistry (IHC), as described previously.
2
ISH targeting the RNA sequence encoding IAV matrix protein was also performed to co-localize genomic material in respiratory tissues. The RNAscope procedure was performed (Formalin-fixed paraffin-embedded (FFPE) sample preparation and pretreatment for the RNAscope 2.5 HD assay, RNAscope 2.5 HD detection reagent–red; Advanced Cell Diagnostics [ACD]) following the manufacturer’s recommendations. ACD probes used included V-influenza-H1N1-H5N1-M (catalog 436211), RNAscope negative control probe–DapB (310043), RNAscope positive control probe–Bt-PPIB (319451). Positive control tissue for IAV IHC and ISH consisted of IAV-positive porcine lung and bovine mammary gland (
Influenza A virus detection by immunohistochemistry (IHC) and in situ hybridization (ISH) in tissue samples from cull dairy cows.
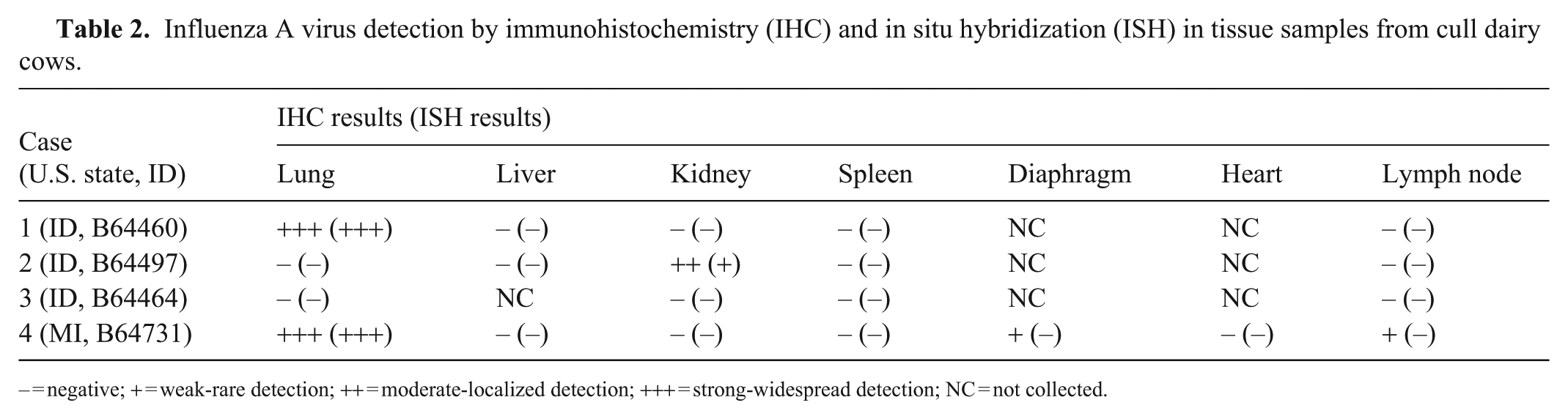

– = negative; + = weak-rare detection; ++ = moderate-localized detection; +++ = strong-widespread detection; NC = not collected.
Histologically, severe suppurative bronchopneumonia was evident in 3 of the 4 positive cows (
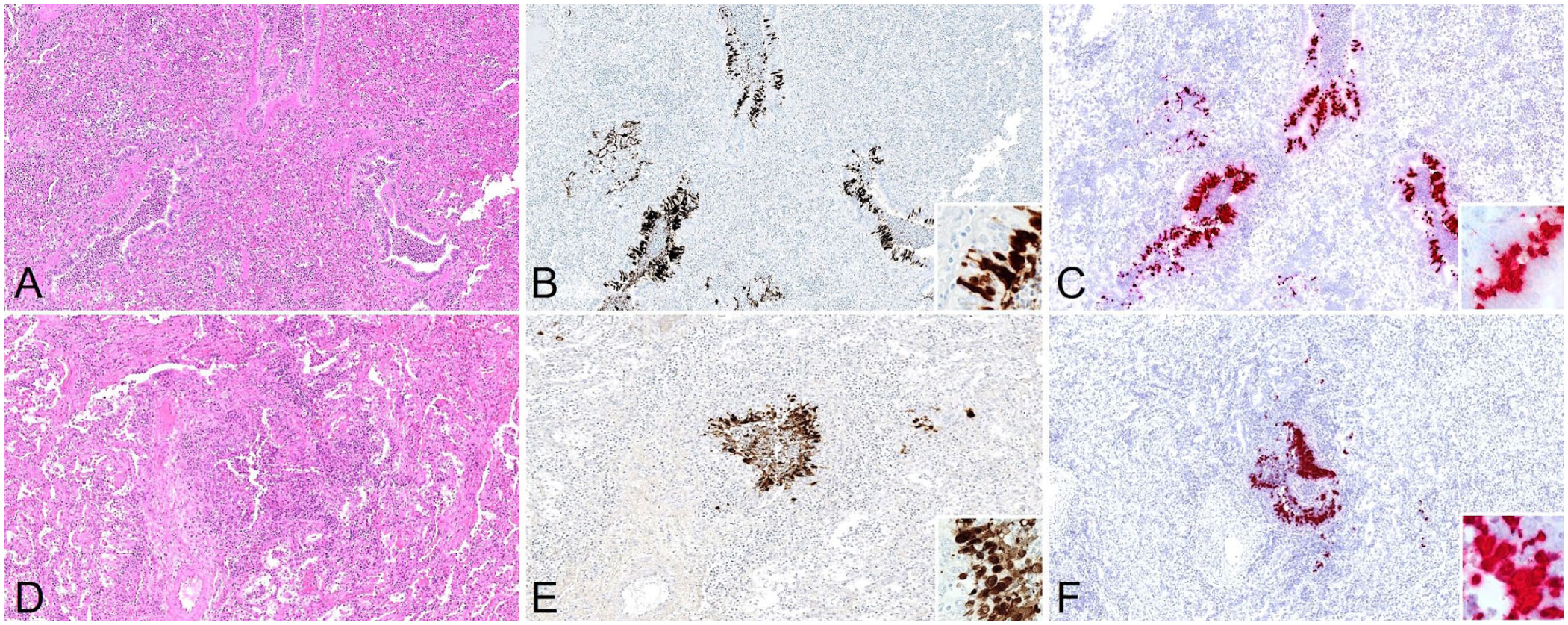
Detection of highly pathogenic avian influenza A(H5N1) clade 2.3.4.4b virus in the lung of cull dairy cows with bronchopneumonia.
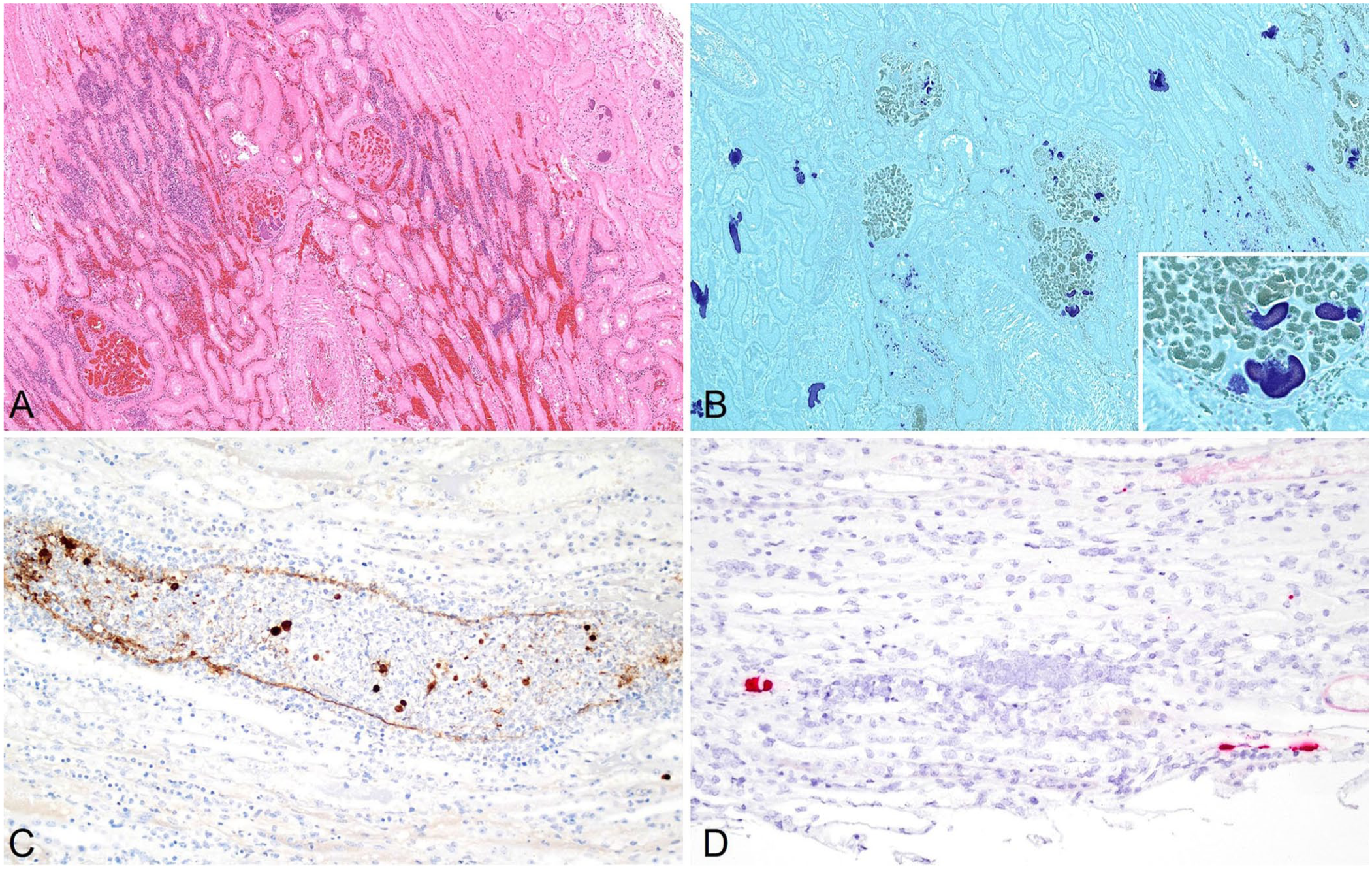
Detection of highly pathogenic avian influenza A(H5N1) clade 2.3.4.4b virus in the kidney of a cull dairy cow (case 2) with embolic nephritis.
Additionally, FFPE lung tissue from these same 4 dairy cows was evaluated via a bovine respiratory panel real-time PCR (rtPCR), which included bovine alphaherpesvirus 1 (BoAHV1), bovine coronavirus (BCoV), bovine viral diarrhea virus (BVDV), bovine respiratory syncytial virus (BRSV), Mycoplasmopsis bovis, Mannheimia haemolytica, Pasteurella multocida, and Histophilus somni. FFPE tissue was extracted (MagMAX FFPE DNA/RNA Ultra kit; ThermoFisher). The bovine respiratory panel rtPCR panel was conducted per the ISU-VDL protocol as a multiplex panel with primers and probes (available on request) targeting BCoV, BRSV, BVDV, BoAHV1, M. haemolytica, P. multocida, H. somni, and M. bovis, using a 25 µL reaction, as described previously. 11
In cases 1, 3, and 4 with bronchopneumonia, H. somni and P. multocida were detected by rtPCR testing ( Table 3 ). M. haemolytica nucleic acid was detected in cases 1 and 3. BCoV nucleic acid was detected in case 4 and M. bovis in case 3. Case 2 with embolic nephritis and pneumonia, but lacking histologic evidence of bronchopneumonia, was negative for all viral and bacterial agents in the bovine respiratory panel.
Bovine respiratory panel rtPCR Ct results for formalin-fixed, paraffin-embedded lung samples from cull dairy cows.
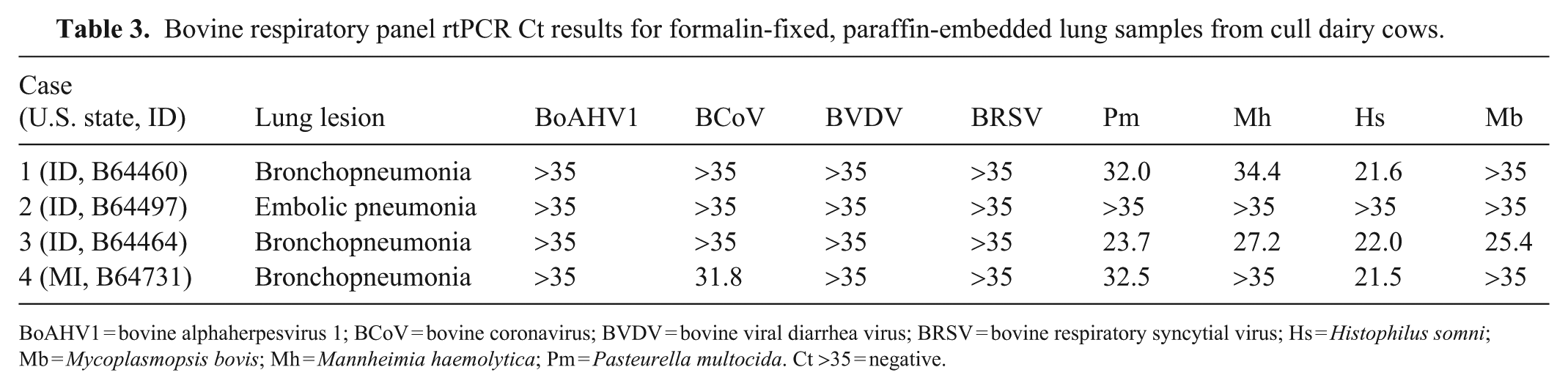
BoAHV1 = bovine alphaherpesvirus 1; BCoV = bovine coronavirus; BVDV = bovine viral diarrhea virus; BRSV = bovine respiratory syncytial virus; Hs = Histophilus somni; Mb = Mycoplasmopsis bovis; Mh = Mannheimia haemolytica; Pm = Pasteurella multocida. Ct >35 = negative.
Our findings demonstrate viral replication within respiratory epithelium of the lung. The results further reveal co-infection of HPAI A(H5N1) virus with common bovine respiratory pathogens in cows with bronchopneumonia. Interestingly, the kidney of one cow with embolic bacterial nephritis had possible HPAI A(H5N1) viral replication within a few tubules of the renal medulla. Initial pathogenesis studies suggest that HPAI A(H5N1) virus is not a primary cause of respiratory disease in dairy cows.1,6 Unfortunately, we cannot confirm whether these animals had viral mastitis, because no mammary gland tissue or milk was sampled at slaughter. Furthermore, it is unclear if a bacterial bronchopneumonia developed secondary to an underlying HPAI A(H5N1) viral infection, or if damage to the lung tissue by bacterial pathogens created conditions that facilitated viral replication.
Although HPAI A(H5N1) virus was detected by PCR testing of the diaphragm and heart of case 4, virus was not detected in myofibers by IHC or ISH. The few cells within the diaphragm and lymph node with IHC cytoplasmic immunoreactivity may have been resident macrophages or satellite cells (muscle). The lack of nuclear immunoreactivity may indicate that virus was phagocytosed by these cells without nuclear invasion or viral replication. Due to the close proximity of the diaphragm to the lungs, it is also possible that these cells could have been circulating inflammatory cells from the lungs.
Our study is unique in that natural infection and replication of HPAI A(H5N1) virus were identified in the lungs and kidney of dairy cows brought to slaughter. Pneumonia in cows at postmortem inspection is common and often multifactorial in etiology (e.g., shipping fever). However, until recently, IAV was regarded as an unlikely contributing pathogen. Collectively, our findings raise questions about the circumstances surrounding HPAI A(H5N1) virus replication within the respiratory epithelium of the bovine lung. Can the virus cause primary pneumonia or does the virus take advantage of already diseased tissue? It also raises questions about the immune response and degree of influenza viral replication, given that most cows with shipping fever have at least some degree of immunosuppression. Finally, if replication of HPAI A(H5N1) virus is occurring at high rates in bronchial and bronchiolar epithelial cells, is horizontal transmission via respiratory droplets possible in these herds? Although the detection of IAV by ISH and IHC in the kidney was potentially an outlier, IAV RNA has been detected in urine from a cow on an affected farm. 8 Further studies on interactions between HPAI A(H5N1) virus and a host immune system that is under the stress of coinfection are warranted.
USDA FSIS Disclaimer: All HPAI A(H5N1) virus-infected cows were condemned at slaughter and did not enter the food supply. Cooking meat to the USDA FSIS recommended minimum internal temperature (71.1ºC for ground beef) has been shown to result in a reduction of avian IAV particles to nondetectable levels. 9
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387261417354 – Supplemental material for Detection of highly pathogenic avian influenza A(H5N1) clade 2.3.4.4b virus in cull dairy cows with underlying respiratory and systemic disease
Supplemental material, sj-pdf-1-vdi-10.1177_10406387261417354 for Detection of highly pathogenic avian influenza A(H5N1) clade 2.3.4.4b virus in cull dairy cows with underlying respiratory and systemic disease by Daniel J. Righter, Erin B. Howey, Chris L. Siepker, Eric R. Burrough, Drew R. Magstadt, Marta Mainenti, Asha Fears, Aaron D. Lehmkuhl, Gleeson Murphy, Kimberly Lehman, Mia Kim Torchetti, Suelee Robbe-Austerman and Carrie E. Schmidt in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Trish Archer, Karen Stockwell, and Shayla Belton at the USDA FSIS Eastern Laboratory Pathology Branch for their collaborative efforts, and PHVs Betina Proof, Leslie Luscher, and William Boyd for critical efforts in identifying and collecting samples from infected dairy cows. We further thank Jennifer Groeltz-Thrush at the Iowa State Veterinary Diagnostic Laboratory for performing IHC and ISH on tissues evaluated in this manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
ORCID iDs
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
