Abstract
A total of 18 samples from 4 outbreaks of gizzard erosions in broiler chickens in Europe were used in the current study. Fowl adenoviruses were found in samples from all 4 outbreaks, and isolates were identified as Fowl adenovirus A (FAdV-A) serotype 1. As described earlier, polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) analysis of the long fiber gene was conducted. However, all 18 samples showed the same pattern as apathogenic FAdV-1 strains: Ote and chicken embryo lethal orphan (CELO) viruses. Nucleotide and amino acid sequences of the long and short fiber of several isolates from broiler chickens with gizzard erosions were analyzed, and 100% identity between the field isolates on the protein level was revealed. Only 1 nonsynonymous mutation (T→A) was present in the long fiber of studied isolates compared to the CELO strain. The same mutation was also present in the Ote strain. Four nonsynonymous mutations were present in the long fiber of studied isolates compared to Ote strain. In the short fiber, 6 nonsynonymous mutations were found in the studied isolates compared to the CELO strain. However, the short fiber of pathogenic isolates was 100% identical to apathogenic Ote strain. In conclusion, the usefulness of PCR-RFLP analysis of the long fiber gene of FAdV-1 isolates in distinguishing between those that induce gizzard erosions and those that do not remains questionable for the isolates obtained from European poultry flocks. The role of certain FAdV-1 strains with their long and short fiber in pathogenicity regarding gizzard erosions is still not clear.
Fowl adenoviruses (FAdVs) belong to the family Adenoviridae, genus Aviadenovirus. It is the general opinion that they are endemic worldwide and can be isolated from both healthy and sick birds. The FAdVs are mainly responsible for naturally acquired outbreaks of inclusion body hepatitis-hydropericardium syndrome, respiratory tract disease, and gizzard erosions. 2
The viruses are diagnosed routinely by virus isolation in embryonated eggs or cell culture followed by polymerase chain reaction (PCR). 17 The FAdVs have been grouped into 5 different species (A–E) based on their hexon gene sequences and are further subdivided into 12 serotypes (1–8a and 8b–11) based on cross-neutralization test. 3,5 Use of PCR followed by restriction enzyme digestion or sequencing of the products allows the differentiation of field isolates to species and presumptive serotypes. 9,10,16
Gizzard erosions associated with FAdV infection have been reported from slaughtered commercial broiler chickens in Japan, and Fowl adenovirus A serotype 1 (FAdV-1) has been isolated in most cases. 1,13,14 Gizzard erosions were also experimentally reproduced in chickens inoculated with isolated FAdV-1. 12 However strain Ote, the reference strain of FAdV-1 in Japan, failed to induce gizzard erosions. Therefore, the existence of strains of FAdV-1 of variable pathogenicity was postulated: those capable of causing gizzard erosions (“pathogenic”) and those that are incapable of causing these lesions (“apathogenic”). 11
Contrary to most adenoviruses, FAdV-1 has 2 fibers of different lengths. 6 The distal parts (or knobs) of these fibers are involved in cellular receptor(s) binding. In addition, fibers of FAdV have been shown to play an important role in the infectivity and pathogenicity of FAdV. 15 Moreover, it was suggested that PCR–restriction fragment length polymorphism (PCR-RFLP) analyses of long fiber genes could be used to distinguish between pathogenic and apathogenic FAdV-1 strains in regard to gizzard erosions. 11
Recently, gizzard erosions associated with FAdV-1 infection have been described in chickens in Italy and Poland. 7,8 It is believed that chicken embryo lethal orphan (CELO), a European reference FAdV-1 strain, does not induce gizzard erosions in chickens, similar to Ote (the FAdV-1 reference strain commonly used in Japan). The current report describes the detection and identification of FAdV-1 in samples from broiler chickens with gizzard erosions from several European countries. In the samples, PCR-RFLP analyses of the long fiber genes were conducted. Nucleotide and amino acid sequences of the long and short fibers of several isolates from broiler chickens with gizzard erosions were analyzed and compared to those of apathogenic FAdV-1 reference strains: Ote and CELO. The aim was to examine the usefulness of sequence analysis of fiber genes for characterizing field isolates from Europe.
List of samples used in the current study. *
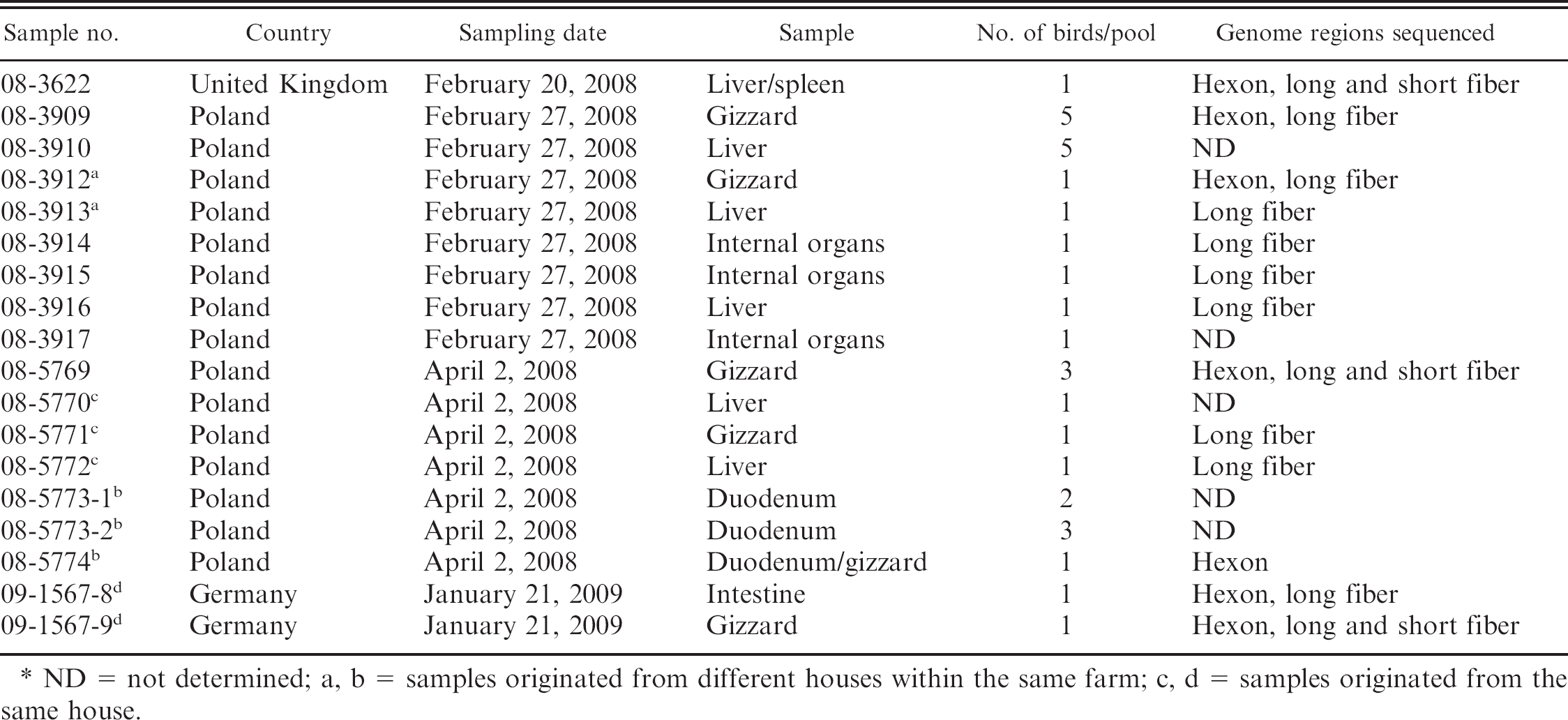
ND = not determined; a, b = samples originated from different houses within the same farm; c, d = samples originated from the same house.
In order to analyze the genetic diversity among FAdV-1 isolates, a total of 18 virus samples from broiler chickens with gizzard erosions from 3 European countries sampled in 2008–2009 were used. Some characteristics of the samples are listed in Table 1. Samples 08-3622, 08-3909, 08-3910, 08-3914, 08-3915, 08-3916, 08-3917, and 08-5769 consisted of different organs from 1–5 individual birds from different farms. Samples from different houses within farms and from single houses were also analyzed (Table 1). Pathomorphological lesions were similar as those described previously, 14 with blood in the gizzard and necrosis below the keratinoid layer. Bacteriologic investigations from carcasses were not very conclusive, and Escherichia coli was dominant in various organs. Avian reovirus was isolated from samples 08-3917, 08-5771, 08-5773-1, and 08-5773-2, whereas all others were negative. For comparison, apathogenic FAdV-1 strains Ote and CELO were used.
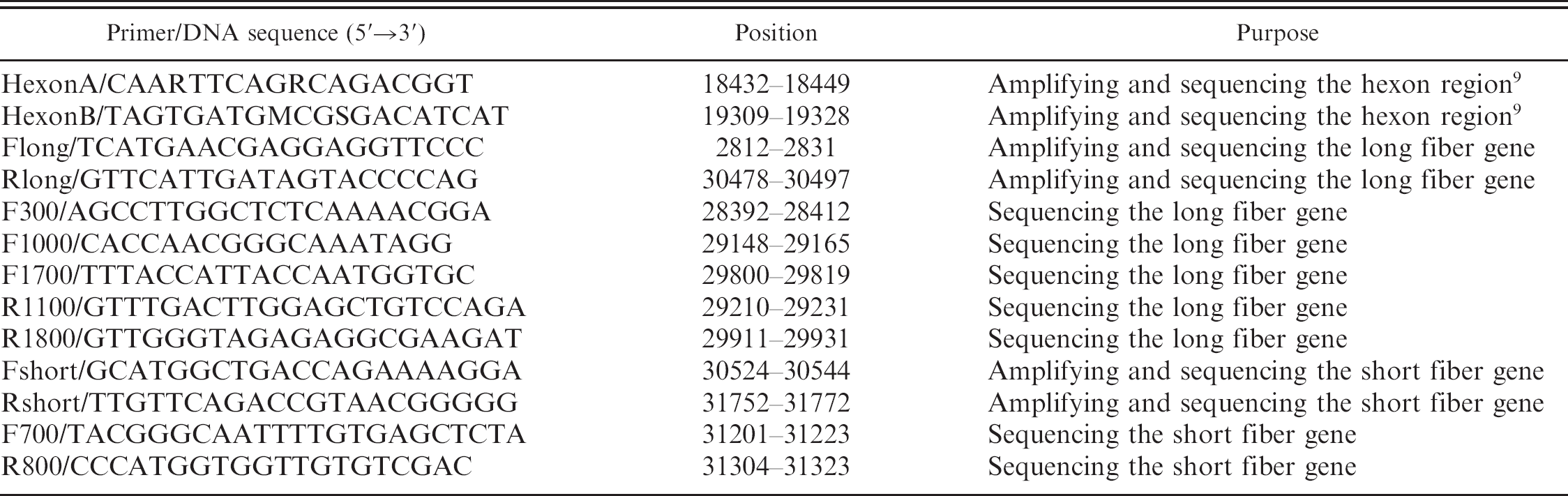
Oligonucleotides used in the current study for amplifying and sequencing the Fowl adenovirus A serotype 1 genome regions. Genomic positions are given for the GenBank sequence of the chicken embryo lethal orphan strain U46933.
The organ pools were homogenized, and homogenates were used directly for DNA isolation and for inoculation of confluent monolayers of chicken embryo liver cells. The supernatants of obviously infected cell monolayers showing cytopathic effect were also used for DNA isolation. Total DNA was isolated from 180 μl of tissue homogenates or cell culture supernatants using a commercial kit according to the manufacturer's instructions. a
Diagnosis of FAdV was based on pathomorphological signs, virus isolation, and detection of FAdV DNA in tissues or infected cells using PCR for the loop 1 region of hexon gene 9 (data not shown). Oligonucleotide primers for the amplification of DNA, corresponding to the entire long and short fiber genes (Table 2), were designed on the basis of the published sequence data of the CELO strain. 4 The PCR products were approximately 2,382 base pair (bp) and 1,240 bp for long and short fiber genes, respectively (data not shown).
After gel electrophoresis, PCR products were extracted from the gel. The PCR products corresponding to the entire long fiber gene of 18 samples were analyzed by RFLP analysis using HinfI. 11 The PCR products were also directly sequenced in both directions using primers listed in Table 2. Sequences were submitted to the GenBank database under accession nos. FN557181–FN557186.
Assembly and analysis of sequences, as well as the nucleotide alignments, were performed with the Accelrys Gene version 2.5 b and Lasergene c software. For sequence analysis, primer-binding sites were excluded from analysis, and 750, 2,291, and 1,197 bp fragments of the hexon, the long fiber genes, and the short fiber genes, respectively, were analyzed. Nucleotide sequences of the hexon gene of several isolates from broiler chickens with gizzard erosions and of apathogenic FAdV-1 reference strain Ote were analyzed. The PCR products obtained from 12 samples and the Ote strain through amplification of the long fiber gene were sequenced. In addition, the sequence of the short fiber gene from 3 field isolates and Ote was established (Table 1). Hexon, long fiber gene, and short fiber gene sequences of completely sequenced CELO strain were also used in sequence analysis. 4 In addition, predicted amino acid sequences of long and short fiber of isolates from birds with gizzard erosions were compared to those of reference strains.
Phylogenetic analysis was performed using Accelrys Gene version 2.5, and the tree was generated using the neighbor-joining method on uncorrected p-distances. For phylogenetic analysis, a 687-bp region (corresponding to 1–687 bp of the GenBank sequence AF339914) of the hexon gene was used. Other sequences used were previously published sequences of FAdV reference strains. 9,16
It was possible to amplify fragments of hexon genes from representative samples of all 4 outbreaks of gizzard erosions used in the present study (data not shown). In phylogenetic analysis, 6 subgroups were identified within FAdVs based on hexon gene sequences (data not shown). In general, each subgroup corresponded to different FAdV species (A–E), and the strain TR22 formed a separate cluster. All tested isolates segregated into the same cluster together with the CELO virus. The samples positive for presence of FAdV-1 DNA originated from broiler chickens with gizzard erosions from the United Kingdom, Poland, and Germany (Table 1), which suggests that the virus is possibly widespread within Europe. However, until now, very limited data about the clinical significance of FAdV-1 infection in Europe were available. More epidemiologic studies are needed to reveal more insights into the clinical importance of FAdV-1 infections and to confirm the role of FAdV-1 in this disease condition, as indicated from reproduction studies. 12
The patterns from the PCR-RFLP analysis of 7 samples are shown in Figure 1. Surprisingly, the RFLP patterns of all samples were identical, and they were the same as that of apathogenic FAdV-1 strains Ote and CELO. No isolate had the pathogenic-type restriction pattern or mixed restriction pattern as reported previously. 11 The RFLP patterns of all other 11 samples were the same as that of apathogenic FAdV-1 strains (data not shown). Sequence analyses of the long fiber gene of FAdV-1 isolates and prediction of HinfI recognition sites confirmed the results obtained by PCR-RFLP analysis (data not shown). In conclusion, the usefulness of PCR-RFLP analysis of the long fiber gene of FAdV-1 isolates in distinguishing between those that induce gizzard erosions and those that do not remains questionable for the isolates obtained from European poultry flocks.
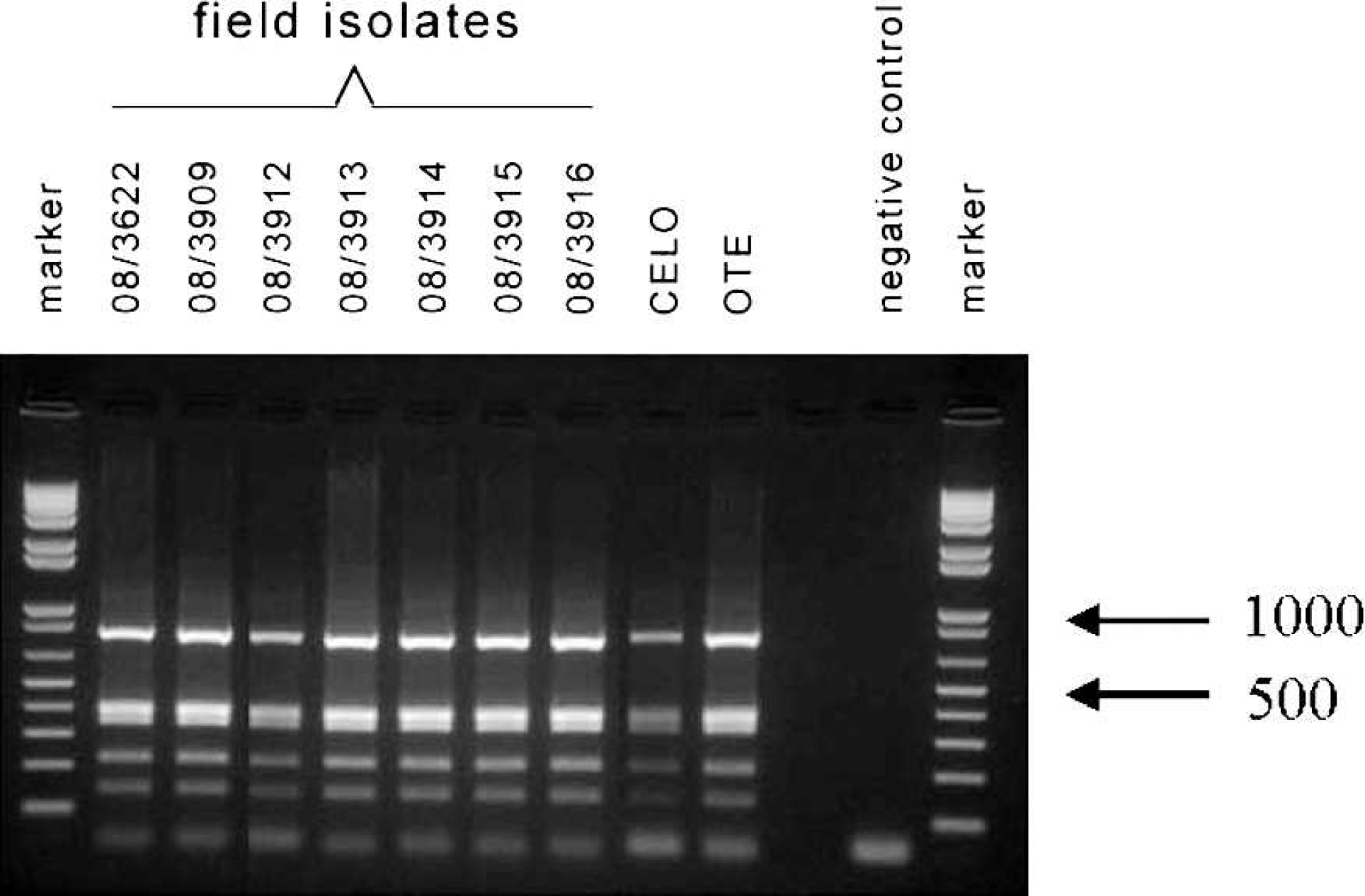
Polymerase chain reaction (PCR)–restriction fragment length polymorphism (RFLP) patterns of several Fowl adenovirus A serotype 1 (FAdV-1) samples from birds with gizzard erosions and the chicken embryo lethal orphan (CELO) and Ote strains. The PCR products corresponding to the entire long fiber gene were analyzed by RFLP analysis using HinfI.
The analyzed portion of the hexon gene was 100% identical in all sequenced samples from birds with gizzard erosions from Europe, and they were all 99.88% identical to the CELO strain (data not shown). Therefore, the FAdV-1 detected in Europe in 2008–2009 is related to the CELO strain. The FAdV-1 isolates from birds with gizzard erosions had 100% nucleotide sequence identity to each other within the long fiber gene, 99.13% nucleotide sequence identity to the CELO strain, and 99.78% nucleotide sequence identity to the Ote strain (data not shown). Within the short fiber gene, the FAdV-1 isolates from birds with gizzard erosions had 99.83–100% nucleotide sequence identity to each other, 98.91–99.08% nucleotide sequence identity to the CELO strain, and 99.75% nucleotide sequence identity to the Ote strain. Results on the genetic heterogeneity of the hexon region of FAdV-1 isolates from birds with gizzard erosions compared to the CELO strain are in agreement with a previously published study on FAdV-1 isolates from Italy. 8 When based on the hexon region, the isolates were more closely related to the CELO strain; however, when based on the long and short fiber genes, the isolates were more closely related to the Ote strain. In the present study, specific mutations could not be linked to the existence of gizzard erosions.
Moreover, amino acid sequences of the long and short fiber genes were analyzed to determine the role of fibers in the pathogenicity of FAdV-1 isolates in regard to gizzard erosions. Only 1 nonsynonymous mutation (T→A) was present within the long fiber gene of FAdV-1 isolates from birds with gizzard erosions compared to the apathogenic CELO strain (Fig. 2A). Because the same mutation was also present in the apathogenic Ote strain, it is very difficult to speculate about the effect of this mutation on the function of the long fiber and the pathogenicity of these isolates. Within the long fiber, 4 nonsynonymous mutations were found in the studied isolates compared to the Ote strain. Again, because the same amino acids were present at these positions in pathogenic isolates and the apathogenic CELO strain, further studies of the long fiber are needed to investigate its role in the pathogenicity of FAdV-1 isolates. Six nonsynonymous mutations were detected within the short fiber of FAdV-1 isolates from birds with gizzard erosions compared to the apathogenic CELO strain (Fig. 2B). Interestingly, all of them were within the knob domain. However, the same mutations were also present in the apathogenic Ote strain, and the short fiber of the studied isolates was 100% identical to the Ote strain at the amino acid level. Therefore, the short fiber does not seem to determine the pathogenicity of FAdV-1 isolates, at least in regard to gizzard erosions. This result is in agreement with previously published results on Japanese isolates. 11 To estimate the influence of viral genetics on pathogenicity, experimental infections with different FAdV-1 isolates would be necessary, keeping in mind that FAdV-1 is often vertically transmitted and can be isolated from healthy birds as well. Furthermore, exchange of fiber genes between individual isolates may be useful to determine their relevance for pathogenicity, as shown for FAdV-8 isolates. 15
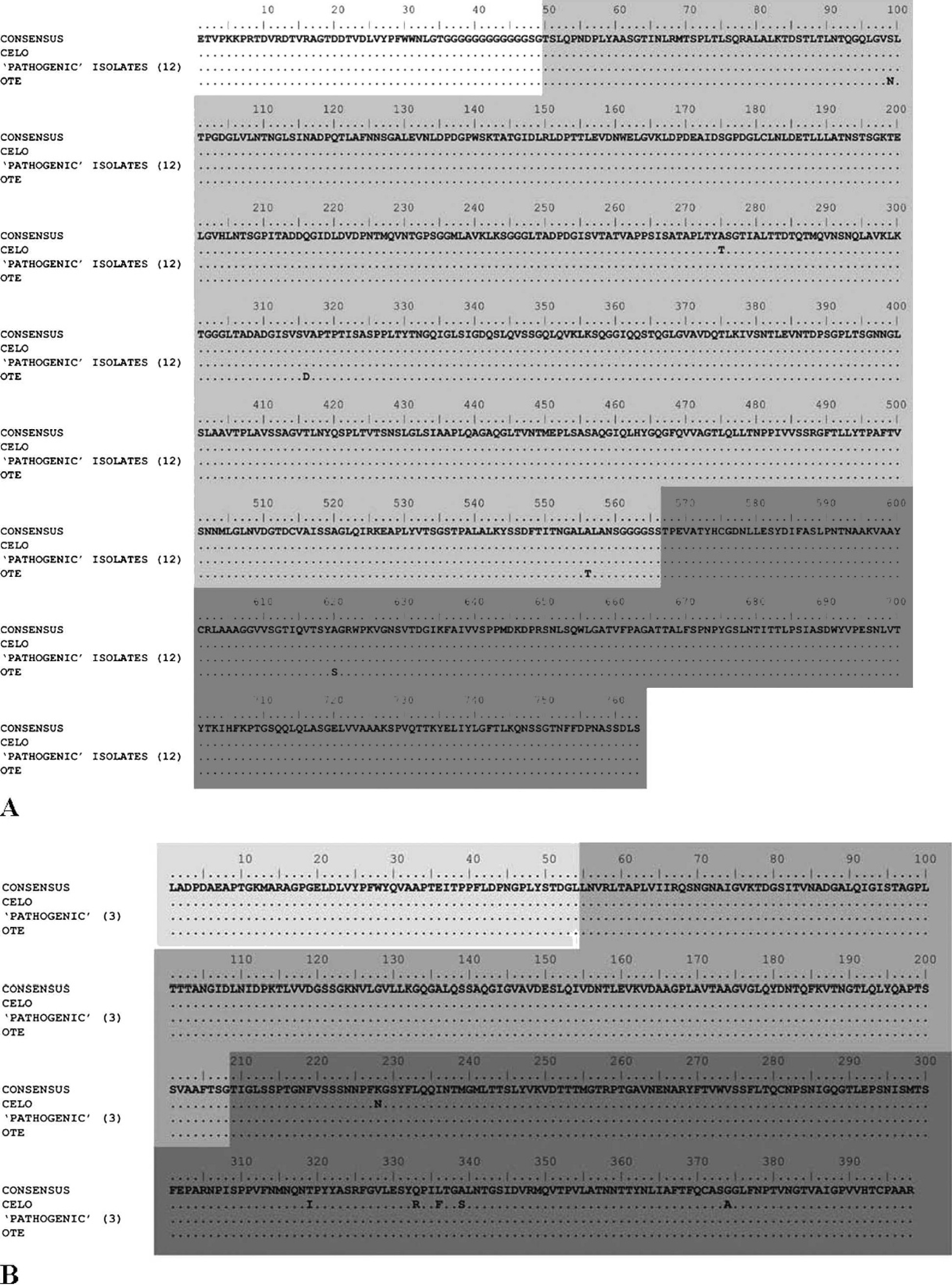
Comparison of the (
To the authors' knowledge, this is the first study comparing several isolates from outbreaks of gizzard erosions in broiler chickens based on nucleic acid sequences of long and short fiber genes. Further investigations of FAdV-1 isolates from birds with gizzard erosions and healthy chickens are necessary. In addition, it would be useful to analyze other genes and whole genomes of additional isolates. The evolution and further epidemiologic analysis of FAdV-1 isolates may also be important for any future vaccine developments to protect birds from gizzard erosions.
Acknowledgements. The authors are thankful to all the colleagues who supplied us with tissue samples and/or FAdV-1 isolates.
Footnotes
a.
DNeasy Blood and Tissue Kit, Qiagen, Vienna, Austria.
b.
Accelrys Inc., San Diego, CA.
c.
DNASTAR Inc., Madison, WI.
