Abstract
In the course of an epidemiologic surveillance program for swine diseases carried out in Spain, 206 cytopathic viruses were isolated from 600 porcine fecal samples between 2004 and 2005. The virus isolates were examined using reverse transcription polymerase chain reaction (RT-PCR) methods specific for different types of porcine picornaviruses, including members of the Teschovirus, Enterovirus, and Sapelovirus genera, and PCR for porcine adenoviruses. Of the 206 isolates, 97 (47%) were identified as teschoviruses, 18 (9%) as sapeloviruses, and 7 (3%) as porcine adenoviruses. Neither Porcine enterovirus B nor Swine vesicular disease virus was found among the isolates. The present study confirms that teschoviruses are highly prevalent in porcine fecal samples, at least in Spain. It also reveals that these viruses commonly circulate among apparently healthy pigs.
Enteric viruses are pathogens infecting the gastrointestinal tract of vertebrates. They replicate in the gastrointestinal epithelia, are shed in feces in large amounts, and often persist in the environment for long periods, making the fecal–oral route (through fecally contaminated food, water, or fomites) their main route of transmission. The presence of enteric viruses in water and environmental samples is considered a good correlate of fecal contamination. 3,6
In swine, important enteric viruses belong to the Picornaviridae family, particularly the Enterovirus, Teschovirus, and Sapelovirus (formerly Porcine enterovirus A or Porcine enterovirus serotype 8) 12 genera. Porcine adenoviruses (family Adenoviridae) have also been frequently found in pig fecal and environmental samples. 8 Among the Picornaviridae family, Swine vesicular disease virus (SVDV), a subtype of Human enterovirus B and belonging to the coxsackievirus B group, causes a disease of high relevance for trade and is included in the World Organization for Animal Health (OIE) list of notifiable animal diseases. 13 In Spain, a specific program for surveillance of swine diseases is being implemented. This surveillance program, which includes SVDV, involves feces collection at farms where serologic data raise suspicion of the disease, according to the Manual for Diagnosis of SVD in force in the European Union. 1 In 2004 and 2005, a total of 600 porcine fecal samples were examined under this surveillance program. The samples were collected from asymptomatic, healthy pigs from 10 autonomous communities within Spain, representing 79% of Spanish territory and 68% of Spain's total porcine census in 2005.
All specimens were subjected to virus isolation by inoculation of fecal homogenates in pig kidney cell lines (IB-RS2 a or PK-15 b ) following standard methods for SVDV diagnosis. The isolation was considered negative when no cytopathic effect could be observed after 3 blind cell-culture passages. Using this method, 206 cytopathic viral isolates (34% of the samples) were obtained. All of them were negative for SVDV using SVDV-specific real-time reverse transcription polymerase chain reaction (RT-PCR) 1,11 ; therefore, it was concluded that all cytopathic viruses isolated were different from SVDV. These isolates were generically designated as “nonvesicular porcine enteric viruses” (NV-PEVs). Their identity was thus investigated in order to recognize and characterize the different types of cytopathic enteric viruses, particularly those belonging to the Picornaviridae and Adenoviridae families, that can be found in swine fecal samples in Spain, and to evaluate their relative prevalence.
Assessment of the performance of the polymerase chain reaction (PCR), reverse transcription (RT)-PCR, and realtime RT-PCR methods for Porcine teschovirus (PTV), Porcine sapelovirus (PSV), Porcine enterovirus B (PEV-B), and Porcine adenovirus (PAdV).
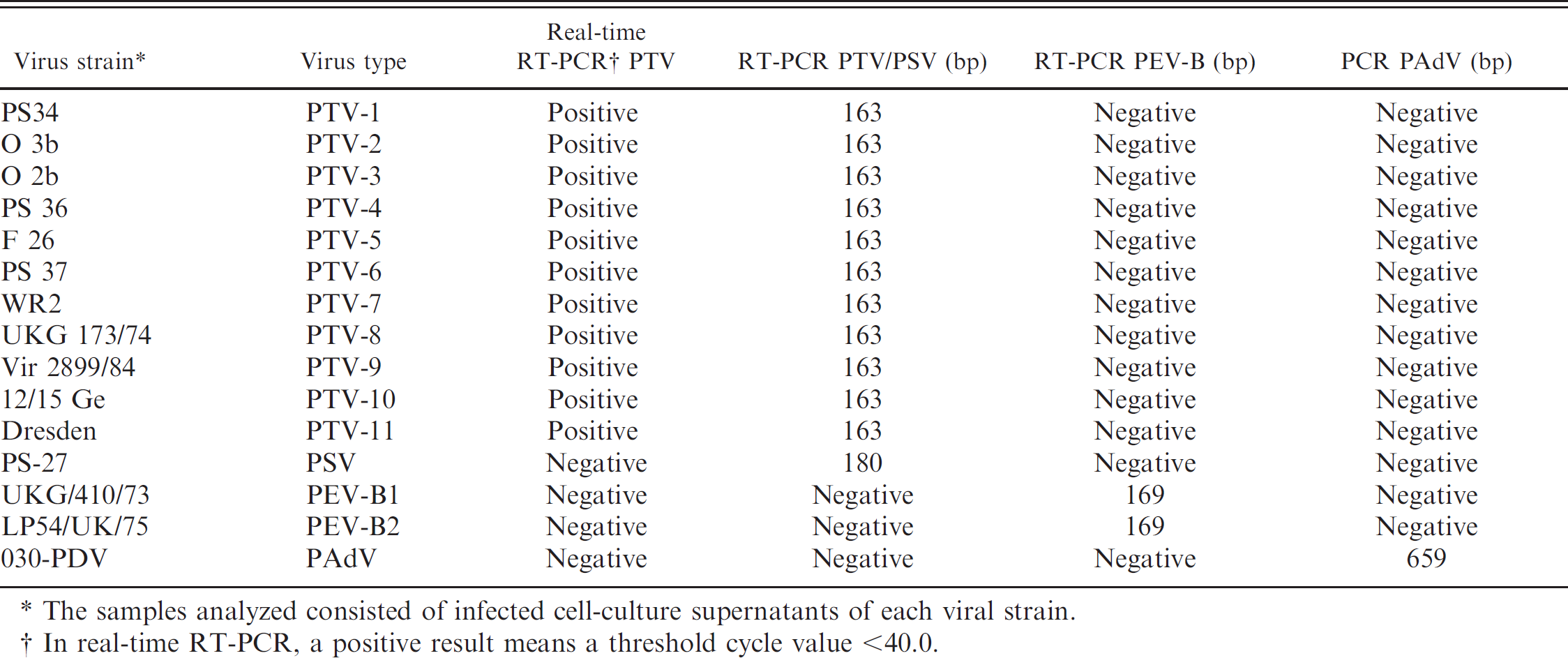
The samples analyzed consisted of infected cell-culture supernatants of each viral strain.
In real-time RT-PCR, a positive result means a threshold cycle value <40.0.
The molecular identity of the 206 NV-PEVs obtained was investigated by applying a series of analyses for different groups of viruses through specific PCR and RT-PCR methods. The first test applied was a screening for Porcine teschovirus (PTV) using a real-time RT-PCR, 4 validated with the type strains for the 11 known PTV serotypes c , d (Table 1). Using this PTV-specific assay, 97 of the 206 isolates yielded positive results. Second, a previously described RT-PCR method for the differential diagnosis of Porcine sapelovirus (PSV) and PTV 10 was used. This conventional RT-PCR is able to distinguish PTV and PSV by the size of the band amplified. To control the specificity of the technique, samples corresponding to PSV and PTV reference strains (1–11) c , d were assessed, giving bands of the expected sizes (Table 1). When the assay was applied to all 206 virus isolates, 18 PTV-negative isolates were identified as PSV, whereas only 48 of 97 PTV-positive isolates were found positive for PTV in this second test; this latter result possibly reflects the higher sensitivity of the real-time RT-PCR protocol applied in the first analysis for the detection of PTV. Next, 2 more tests were applied for 2 groups of cytopathic porcine enteric viruses likely to be found in swine fecal samples: an RT-PCR assay for enteroviruses that detects Porcine enterovirus B (PEV-B) 5,9 and a specific assay for the detection of Porcine adenovirus A–C (PAdV-A–C; protease gene). 8 Both tests were assessed using specific viral material as a performance control: in the first method, 2 serotypes of PEV-B (1 and 2 reference strains) c were used, whereas in the second method, a PAdV isolate e was used. Each technique was able to detect its homologous virus and did not cross-react with other viruses analyzed in the present study (Table 1). The screening for PAdVs identified 7 isolates as belonging to this group, whereas the screening for PEV-B yielded negative results in all samples analyzed. A summary of the results obtained is shown in Table 2. One of the PAdV-positive samples was also positive for PTV. Because no cross-reaction was observed between both tests for each type of virus (Table 1), the result was interpreted as a double infection in the animal from which this sample originated.
The testing conducted resulted in the identification of 122 virus isolates out of the 206 viral isolates obtained from swine fecal samples from a wide territory within Spain. No disease symptoms were associated with the presence of these cytopathic viruses in the samples, so the infected individuals were considered to be asymptomatic carriers. Among the NV-PEVs analyzed, PTVs were the most abundant enteric viruses in the population studied. However, the study has important limitations: first, it focuses on enteric viruses that cause cytopathic effects in the cells used for viral isolation. This means that viruses that were either noncytopathic and/or unable to grow in the 2 cell lines used in the current study were missed. Important enteric viruses that are difficult or unable to grow in vitro comprise representatives of the astrovirus, norovirus, and calicivirus groups. Also, cell culture propagation of some PAdVs is difficult, so it is likely that this group could have been underestimated or biased in the present study, also bearing in mind that previous studies have identified PAdVs as highly prevalent in noncultured swine fecal and environmental samples in Catalonia (northeast Spain). 2,8
Previous studies conducted in Italy 5 using essentially the same approach (i.e., analyzing viral isolates derived from porcine fecal samples from the SVD surveillance program in Italy) obtained results that differ from the present results: whereas 15 of 33 isolates belonged to the PEV-B group, 14 were PSV, and the remaining 4 samples corresponded to PTVs. These differences could reflect distinct epidemiologic situations in Spain and Italy with regard to these NV-PEVs. Nevertheless, when comparing both studies, one should bear in mind that they differ in 2 important aspects: 1) the number of isolates analyzed (206 vs. 33) and 2) the sampling period, comprising 10 years in the Italian study (1992–2002) and only 2 years in the present study (2004 and 2005).
Summary of the results of the identification of viral isolates from swine feces performed in the current study.
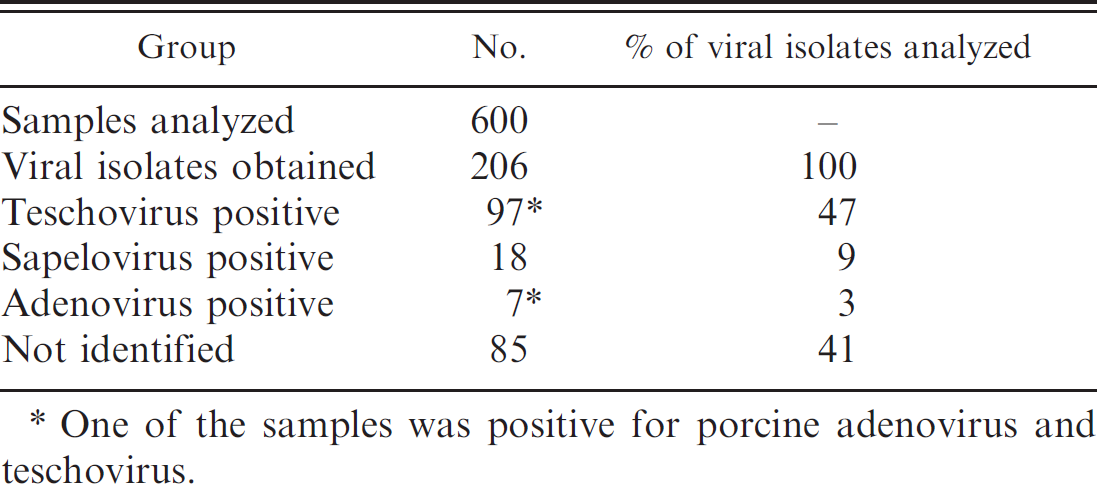
One of the samples was positive for porcine adenovirus and teschovirus.
The current study was focused on members of the Picornaviridae and Adenoviridae families, because viruses belonging to these groups have been detected previously in fecal and environmental samples from pig populations in the absence of disease, 2,4,5,8 However, a still substantial percentage (40%) of the isolates could not be identified. These isolates are thus cytopathic viruses not belonging to the aforementioned viral groups, and their identification would merit further investigation. Rotaviruses and coronaviruses are important cytopathic viruses, usually causing enteric diseases characterized by moderate to severe diarrhea. Although not likely to occur in healthy pig populations, these viruses cannot be excluded from being at least part of the group of isolates that were not identified in this study.
Porcine teschoviruses are specific for swine, persist for long periods in environmental conditions, 7 and have been shown to be a good marker for environmental fecal contamination of swine origin. 4 Three conclusions can be drawn from the fact that a high percentage (16%) of the healthy pig population examined in the present study was excreting PTVs in their feces: 1) although certain PTVs have historically caused serious diseases in pigs, PTVs currently circulating in Spain are apparently not causing overt symptoms of disease; whether this is due to a lack of intrinsic pathogenicity of the circulating viruses or, alternatively, to acquired immunity in these pig populations, remains to be studied; indeed, further research is needed to assess their pathogenicity in controlled trials; 2) PTVs may be a good target for microbial source tracking studies focusing on swine fecal contamination in Spain; whether this is also the case in other countries warrants further investigation; and 3) the swine enteric viruses found in the present study, which grow easily in cell culture, could mask the growth of other relevant viruses in the virus isolation method; this is of particular importance because this method is commonly used for confirmatory diagnosis of SVDV, which represents an important health threat for pig populations. To avoid this problem, studies should focus on identifying cell cultures that support the growth of specific types of swine enteric viruses. In particular, cell lines that are permissive to the growth of SVDV but nonpermissive to the growth of PTVs and/or other swine enteric viruses would be especially useful.
Acknowledgements
The authors are grateful to Diana Condes, Elena Martínez, and María García Casado for their skillful technical work. They also thank E. Brocchi, A. Moreno, and P. Cordioli for providing reference strains for Porcine teschovirus 1–7, PEV-B (serotypes PEV-9 and PEV-10), and Porcine sapelovirus (formerly, Porcine enterovirus A serotype PEV-8), and Roland Zell for providing Porcine teschovirus 8–11 strains. This study was funded by grant AGL2007-61859 from Ministerio de Ciencia e Innovación (MCINN). CCG has been supported by a scholarship from Instituto Nacional de Investigación y Tecnología Agraria y Alimentaria. The authors are indebted to the Spain's Ministry of Environment, Rural and Marine Affairs for supporting this work (agreement INIA.MARM CC08-020).
Footnotes
a.
IBRSs cells, Centro Pan-Americano de Febre Aftosa (PANAF-TOSA-OPAS/OMS), Rio de Janeiro, Brazil.
b.
PK-15 cells (ATCC-CCL-33), American Type Culture Collection, Manassas, VA.
c.
Porcine teschovirus 1–7, Porcine enterovirus B serotypes 1–2, and Porcine sapelovirus reference strains were obtained from the Istituto Zooprofilattico Sperimentale della Lombardia e dell'Emilia Romagna, Brescia, Italy.
d.
Porcine teschovirus 8–11 reference strains were obtained from the Institut für Virologie und Antivirale Therapie, Jena, Germany.
e.
Porcine adenovirus strain (ref 030-PDV) was obtained through the National Veterinary Service Laboratories, Ames, IA.
