Abstract
Food-borne Salmonella infections can produce symptoms from mild gastroenteritis to severe systemic disease and death, representing an important public health issue in U.S. livestock and livestock products, which have been implicated as frequent sources of Salmonella contamination. Concerns have been raised about the spread of antibiotic resistance in Salmonella strains, particularly those that originate from food animal sources, as a result of prophylactic and therapeutic antimicrobial use in these species. Longitudinal comparisons of Salmonella serovars isolated from porcine tissues submitted to the Iowa State University Veterinary Diagnostic Laboratory in 2003 and 2008 were conducted to evaluate changes in serovar dynamics and antimicrobial resistance. Incidence of recovered group C Salmonella enterica serovar Choleraesuis var. Kunzendorf decreased between 2003 and 2008, while recovery of group B strains Salmonella Typhimurium var. 5— (formerly, Copenhagen), Salmonella Agona, Salmonella Derby, Salmonella Heidelberg, and Salmonella Typhimurium increased. Significant changes in resistance interpretation were seen in Salmonella Derby with regard to spectinomycin and sulfadimethoxine; in Salmonella Heidelberg with regard to florfenicol, spectinomycin, and sulfadimethoxine; and in Salmonella Choleraesuis var. Kunzendorf, Salmonella Typhimurium, Salmonella Typhimurium var. 5—, and Salmonella Agona with regard to spectinomycin. Only 2 of 293 isolates in 2003 and 5 of 395 isolates in 2008 were resistant to enrofloxacin. Utilizing antibiotics approved for use in food animals to evaluate antimicrobial resistance provides more specific information on the selection pressure exerted on Salmonella populations through the use of these drugs.
Keywords
Salmonella is one of the most common food-borne pathogens in the United States, with 40,000 confirmed cases reported annually and subclinical infections estimated at 1.2 million per year (http://www.cdc.gov/salmonella/). 1,16 According to surveillance conducted by the Centers for Disease Control and Prevention (CDC) on food-borne illnesses confirmed in 2006, Salmonella infections accounted for 25.6% of hospitalizations and 30.6% of deaths due to food-associated intoxications; thus, these infections represent a significant public health concern. 6,14,16,20,23 Of the Salmonella isolates from human cases of food poisoning in 2005 identified by the CDC's FoodNet, 6 serovars accounted for 61% of isolates recovered: Typhimurium (19%), Enteritidis (18%), Newport (10%), Heidelberg (6%), Javiana (5%), and Salmonella 4,5,12 (3%), all of which have been isolated from animals. 4,7,16 The overlap of serovars recovered from animals and humans highlights the risk of transmission between them. 3,6,16
Livestock species are frequently implicated as sources of Salmonella infections in humans, either through the ingestion of contaminated food products or via direct contact. 5,23 Infections with host-adapted strains such as Salmonella Choleraesuis var. Kunzendorf as well as host-nonspecific strains such as Salmonella Typhimurium produce a variety of disease syndromes in swine, including septicemia, enteritis, and respiratory disease. 3,6,9,22 Antimicrobial use in food-producing animals is aimed at decreasing the morbidity and mortality associated with clinical disease as well as increasing growth 9 and is often implicated as the cause for the development and spread of strains demonstrating antibiotic resistance. 4,12,23 Authors of many studies that evaluate swine Salmonella isolates for antimicrobial resistance patterns perform testing on fecal isolates from swine operations, environmental samples, or slaughter specimens rather than on clinical isolates. 2,15,18,21 In addition, these isolates are often evaluated for susceptibility to antimicrobials used in humans and not for those labeled for use in food animals. 3,16
The objective of the current study was to evaluate serovar distribution and antimicrobial susceptibility interpretations of swine Salmonella isolates from clinically ill pigs in diagnostic samples submitted to the Iowa State University Veterinary Diagnostic Laboratory (ISU VDL; Ames, Iowa) during 2003 compared to those submitted during 2008 and to determine if changes in prevalence or resistance could be identified. The hypothesis was that there was no difference in antimicrobial resistance in the serovars from 2003 to 2008.
Records of Salmonella serovars isolated from various clinical samples, including tissues and intestinal contents from ill pigs submitted to the ISU VDL in 2003 and 2008, were examined for serovar identification and antimicrobial susceptibility testing results. Samples from nonclinical animals were excluded from analysis. Bacterial identification was performed using biochemical testing, Gramnegative identification 96-well biochemical plates, a and serological tests. Serovar determination and confirmation was performed at the National Veterinary Services Laboratory (Ames, IA).
Susceptibility determinations were performed by microbroth dilution methods using an automated system. a Minimum inhibitory concentration (MIC) determinations were performed in accordance with criteria provided in M31 [A2] b (2003 isolates) and M31 [A3] c (2008 isolates). The MIC interpretations of “intermediate” were considered “resistant” to more stringently identified changes in resistance patterns between the 2 time periods. Isolates identified in 2003 were tested against 17 antimicrobials, including ampicillin, ceftiofur, chlortetracycline, clindamycin, enrofloxacin, erythromycin, florfenicol, gentamicin, neomycin, oxytetracycline, penicillin, spectinomycin, sulfadimethoxine, tiamulin, tilmicosin, trimethoprim—sulfameth-oxazole, and tylosin. Isolates identified in 2008 were tested against all of the above antimicrobials with the exception of erythromycin; additionally, danofloxacin and tulathromycin were included in the panel. Escherichia coli American Type Culture Collection (ATTC) 25922, Pseudomonas aeruginosa ATTC 27853, Enterococcus faecalis ATTC 29212, and Staphylococcus aureus ATTC 29213 were used as quality control organisms. “Percent resistant” was calculated as the number of isolates classified as “intermediate” or “resistant” divided by the total number of isolates tested and expressed as a percentage. Statistical comparisons between levels of resistance to each listed antimicrobial between the 2 time points were performed using Fisher's exact test at the P < 0.05 level of significance using SAS statistical software. d
Salmonella was identified from clinical samples from 293 pigs in 2003 and from 395 pigs in 2008. In 2003, recovered serovars included 100 group B (34.1%), 181 group C (61.8%), 2 group D (0.7%), 4 group E (1.4%), and 6 untypeable (2.0%). Isolates identified in 2008 consisted of 274 group B (69.4%), 92 group C (23.3%), 1 group D (0.3%), 22 group E (5.5%), and 6 untypeable (1.5%). Serovar distributions from 2003 and 2008 as well as source of isolate (gastrointestinal [GI] and non-GI) are listed in Table 1.
The most frequently identified serovar recovered from swine samples in 2003 was Salmonella Choleraesuis var. Kunzendorf (168/293, 57.3%), distantly followed by Salmonella Typhimurium var. 5— (formerly, Copenhagen; 39/293, 13.3%), Salmonella Derby (18/293, 6.1%), Salmonella Heidelberg (17/293, 5.8%), Salmonella Typhimurium (10/293, 3.4%), and Salmonella Agona (7/293, 2.4%). In contrast, Salmonella Typhimurium var. 5— was the most commonly identified isolate from clinical submissions in 2008 (120/395, 30.4%), while Salmonella Choleraesuis var. Kunzendorf cases fell to just 59 of 395 (14.9%). Other commonly identified isolates included Salmonella Derby (37/395, 9.4%), Salmonella Typhimurium (35/395, 8.9%), Salmonella Heidelberg (34/395, 8.6%), and Salmonella Agona (30/395, 7.6%). Group E serovars represented a small number of serovars isolated in both 2003 (4/293, 1.4%) and 2008 (22/395, 5.6%), although the numbers did increase in 2008.
Numbers and percentages of the most frequently isolated serovars demonstrating low and high levels of antimicrobial resistance are presented in Figures 1 and 2, respectively. Statistically significant differences (at P < 0.05) in antimicrobial resistance between 2003 and 2008 were present in Salmonella Typhimurium, Salmonella Typhimurium var. 5—, Salmonella Choleraesuis var. Kunzendorf, and Salmonella Agona with regard to spectinomycin; in Salmonella Heidelberg with regard to florfenicol, sulfadimethoxine, and spectinomycin; and in Salmonella Derby with regard to sulfadimethoxine and spectinomycin.
The results of the present study indicate that clinical cases associated with Salmonella Choleraesuis var. Kunzendorf have decreased in frequency over the time period under study. The vast majority of these isolates (163/168 [97.0%] in 2003 and 52/59 [88.1%] in 2008) were detected in tissues outside of the intestinal tract, and 44.0% (74/168) were recovered from both GI and non-GI sites in the same animal. Although this serovar is considered host-adapted for swine, it has been associated with severe disease in humans. 6,13,16 The decreased incidence of this serovar from clinical swine isolates may be due to the increased efficacy of Salmonella Choleraesuis var. Kunzendorf vaccines used in swine production systems. 10
During this same time period, total isolates and cases of several group B serovars, including Salmonella Typhimurium, Salmonella Derby, Salmonella Agona, and Salmonella Heidelberg, and group C serovar Salmonella Infantis have increased. Since these serovars are included in the 20 most frequently identified Salmonella serovars in human infections, 6 further investigations into the epidemiology of swine Salmonella isolates is warranted. 16 The dramatic increase in Salmonella Typhimurium var. 5— may represent an emerging risk to both swine and human populations, as this serovar appears to be filling a niche created by the decline in Salmonella Choleraesuis var. Kunzendorf cases. This trend may be due to changing dynamics of concurrent swine diseases or it could indicate increased virulence associated with specific serovars. Although recovery of Salmonella Typhimurium var. 5— from non-GI tissues decreased from 2003 (13/39, 33.3%) to 2008 (7/120, 5.8%), the incidence of systemic disease associated with this serovar increased over these same time periods. In 2003, just over one-half (20/39, 51.2%) of the cases in which Salmonella Typhimurium var. 5— was isolated presented with signs of systemic disease. In 2008, 68.3% (82/120) of these submissions had systemic illness, indicating that this serovar may be increasing as either a primary or secondary infectious agent.
The 2006 USDA National Animal Health Monitoring System (NAHMS) survey on swine health and management evaluated samples from a random collection of swine operations in 17 states. The NAHMS study found that Salmonella Derby was the most frequently isolated serovar (29.3%) when it came to fecal samples, followed by Salmonella Typhimurium var. 5— (22.6%), Salmonella Agona (10.8%), and Salmonella Anatum (7.5%). Examination of clinical samples demonstrated that while Salmonella Derby was identified much less frequently in the current study than in the NAHMS report, the incidence of this serovar increased between 2003 (6.1%) and 2008 (9.4%). Similar increases were noted with Salmonella Agona (2.4% in 2003, 7.6% in 2008) and Salmonella Anatum (0.3% in 2003, 2.0% in 2008), signifying that Salmonella serovars shed in the feces of nonclinical animals may present an increasing risk for producing clinical disease in swine.
Serovar distributions (number and percent), rank, and source of clinical swine samples recovered at the Iowa State University Veterinary Diagnostic Laboratory in 2003 and 2008.*
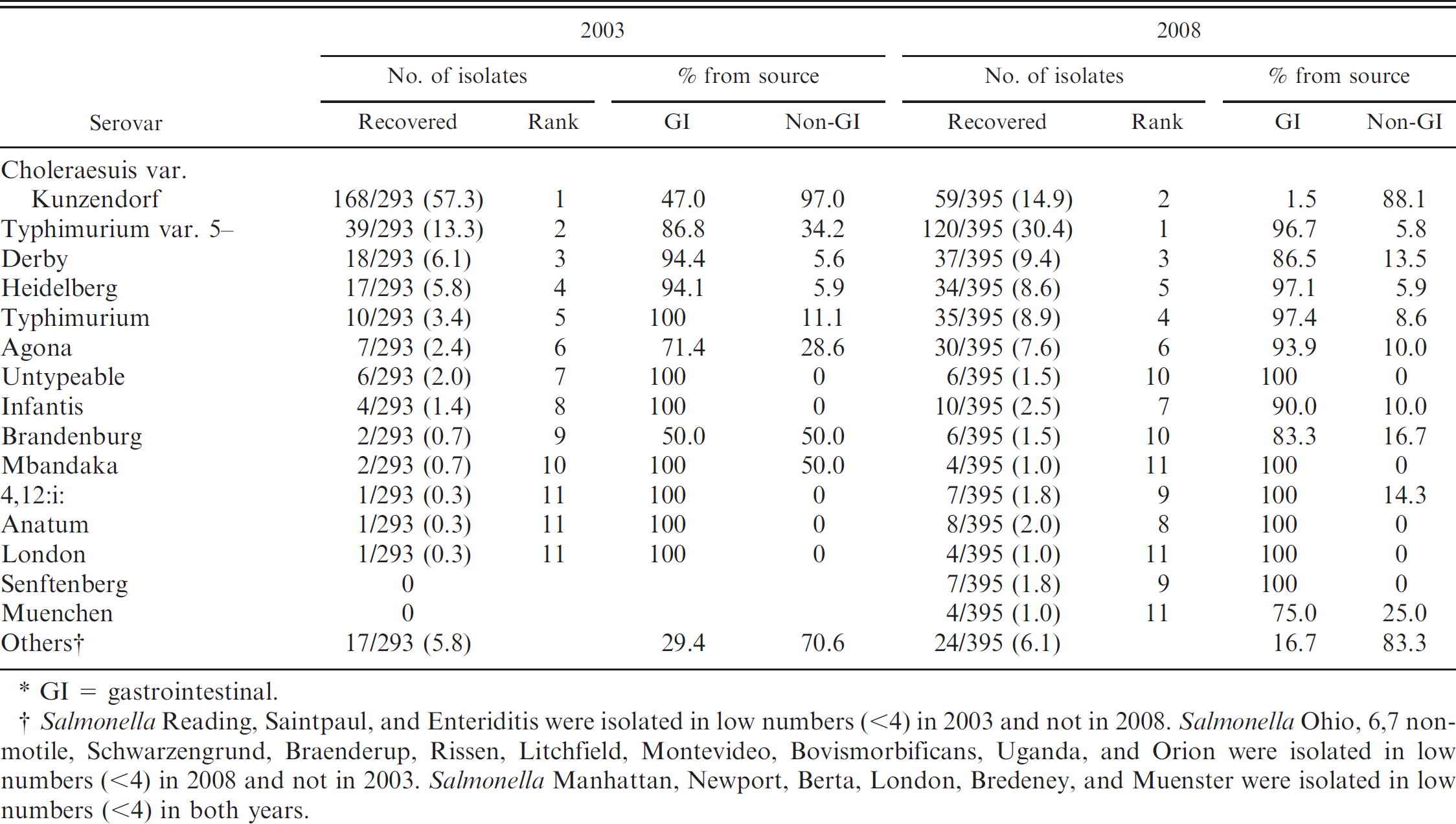
GI = gastrointestinal.
Salmonella Reading, Saintpaul, and Enteriditis were isolated in low numbers (<4) in 2003 and not in 2008. Salmonella Ohio, 6,7 non-motile, Schwarzengrund, Braenderup, Rissen, Litchfield, Montevideo, Bovismorbificans, Uganda, and Orion were isolated in low numbers (<4) in 2008 and not in 2003. Salmonella Manhattan, Newport, Berta, London, Bredeney, and Muenster were isolated in low numbers (<4) in both years.
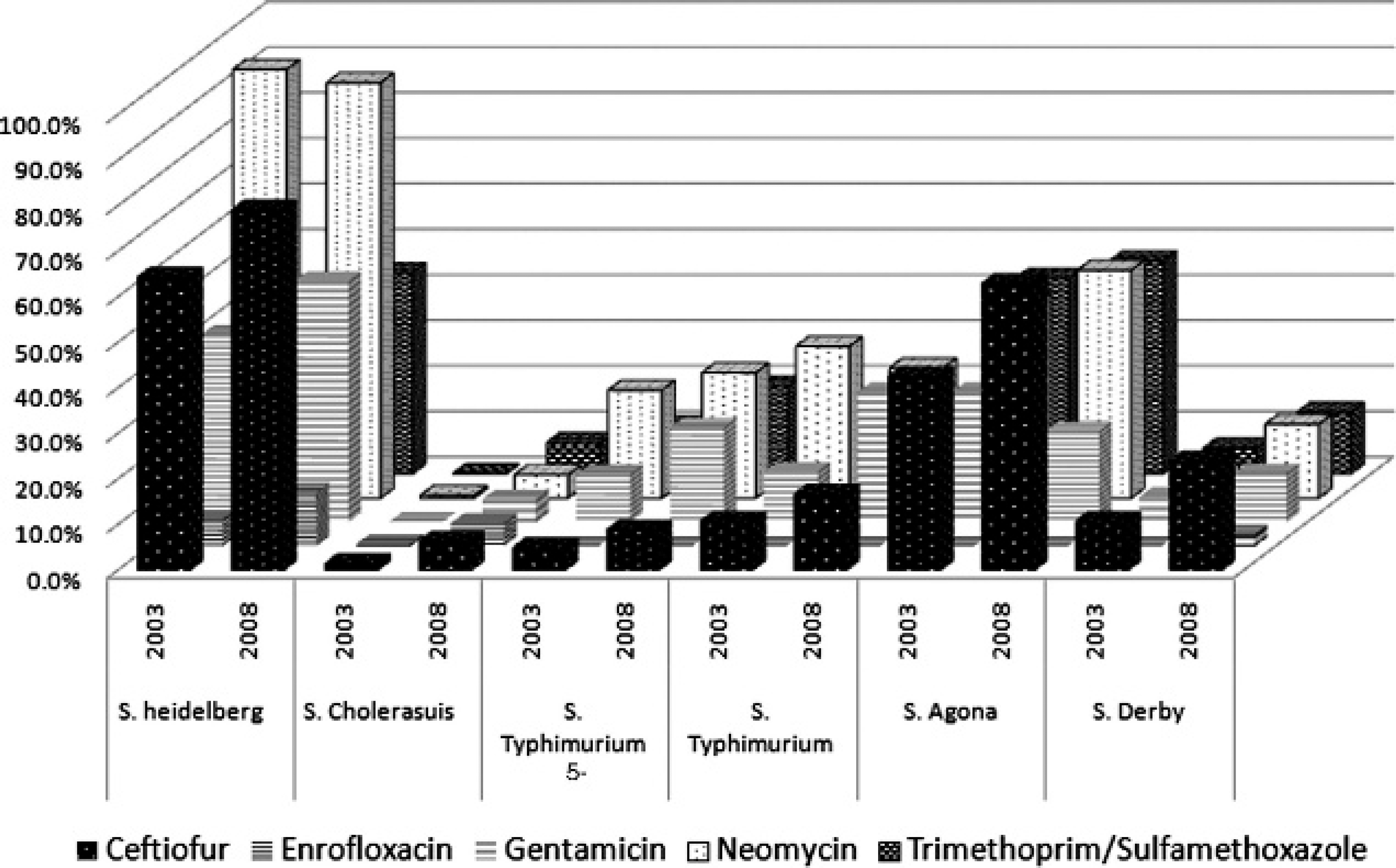
Antimicrobials with low-level resistance patterns to the most frequently isolated Salmonella serotypes recovered from swine in 2003 and 2008. Percent resistant was calculated as the number of isolates classified as “intermediate” or “resistant” divided by the total number of isolates tested and expressed as a percentage. No antimicrobials demonstrated significant difference (P < 0.05) in percent resistant between 2003 and 2008.
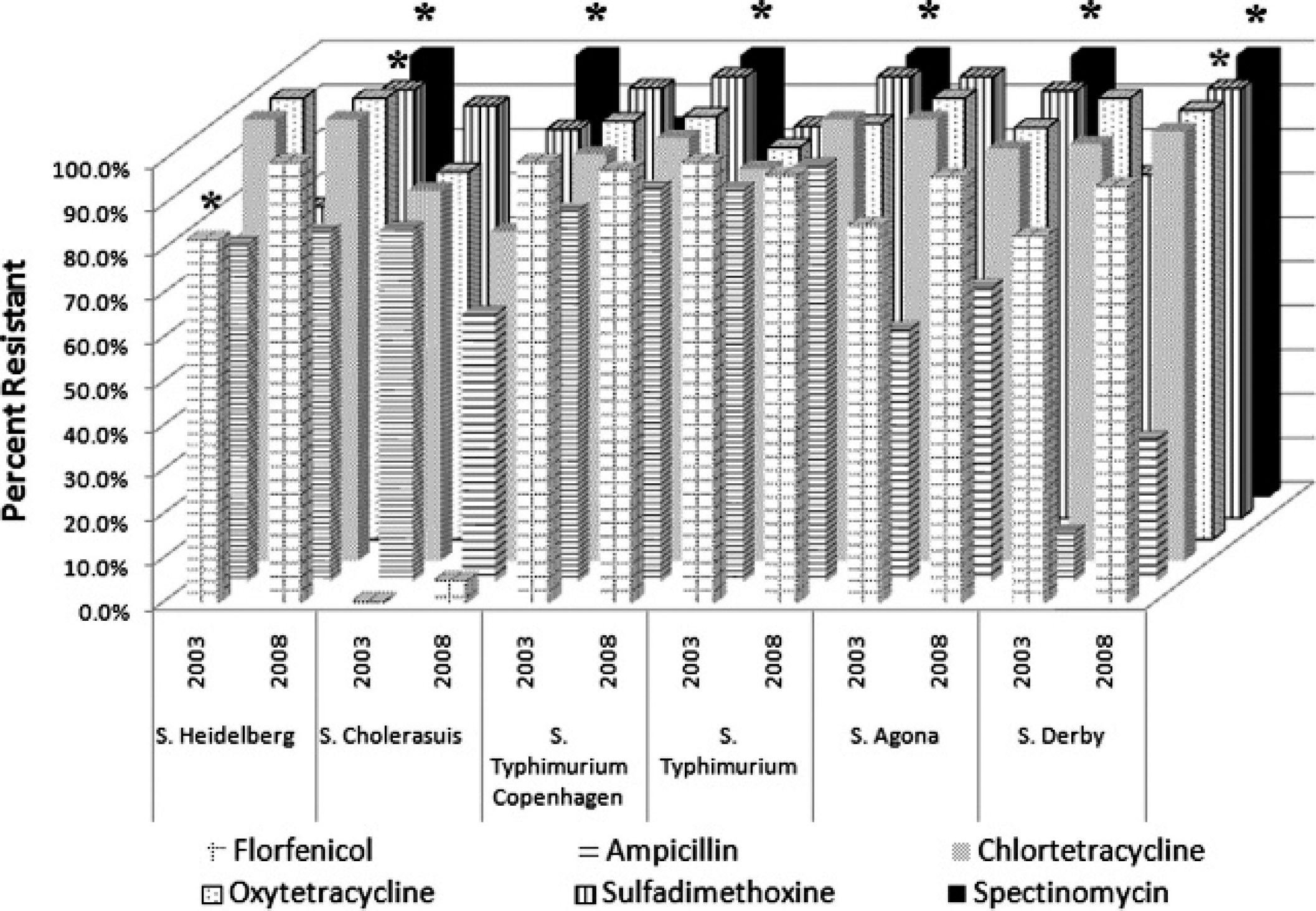
Antimicrobials with high-level resistance patterns to the most frequently isolated Salmonella serotypes recovered from swine in 2003 and 2008. Percent resistant was calculated as the number of isolates classified as “intermediate” or “resistant” divided by the total number of isolates tested and expressed as a percentage. All isolates were resistant to clindamycin, penicillin, tiamulin, and tylosin (data not shown). * Indicates significant difference (P < 0.05) in percent resistant between 2003 and 2008.
Additionally, the observation that Salmonella Typhimurium var. 5— appears to be more frequently associated with clinical disease in swine has been reported by other diagnostic laboratories. 9
Although the NAHMS survey focused on resistance patterns against human antimicrobial drugs and not those labeled for use in food animals, comparisons based on antimicrobial class demonstrate similar results to this study. The NAHMS study found that tetracycline (78.6%), sulfa drugs (67.3%), and spectinomycin (53.3%) demonstrated the highest levels of resistance across all Salmonella serovars; that resistance to ceftiofur (14.6%), potentiated sulfonamides (7.5%), and gentamicin (2.1%) remains relatively low; and that resistance to fluoroquinolones (nalidixic acid [0%] and ciprofloxacin [0%]) was not identified. Veterinary antimicrobials evaluated in the present study showed similar resistance patterns in clinical isolates. Resistance to tetracyclines (chlortetracycline, oxytetracycline) ranged from 74.6% to 100%, while resistance to fluoroquinolones remained very low (0–8.8%).
Assessments using livestock-approved antimicrobials provide the ability to detect emerging patterns of resistance development due to use of these antibiotics over time. In the current study, increased resistance to spectinomycin was the most widespread resistance pattern detected; however, the interpretive criteria used to define resistance to spectinomycin in 2003 were different from those used in 2008. In 2003, MIC values of <64 were considered “susceptible”; 2008 criteria identified MIC values of 16 and 32 as “intermediate” and values of <16 as “susceptible.” For the purposes of this evaluation, MIC interpretations of “intermediate” were counted as resistant in order to apply the most stringent criteria to the assessment of resistance development. A review of MIC values from 2003 indicated that approximately 67% of isolates defined as “susceptible” using those criteria would be considered “resistant” by 2008 standards.
While most Salmonella serovars are resistant to many of the antimicrobials available for veterinary treatment, resistance levels have not changed appreciably over the 5-year period from 2003 to 2008. Of primary concern is the increased resistance to sulfadimethoxine and spectinomycin demonstrated by Salmonella Derby, since this serovar is the most common one isolated from healthy swine and pork products and may represent an emerging risk to human food safety. Additionally, Salmonella Heidelberg showed the highest levels of resistance to all veterinary antimicrobials evaluated in this study.
Very few isolates of any of the serovars demonstrated any resistance to enrofloxacin, the only labeled fluoroquinolone in use during both time periods. Mutations in the bacterial gyrA gene that encodes for DNA gyrase represent the most commonly identified mechanism of resistance development to fluoroquinolones in Campylobacter 17 and had been demonstrated in Salmonella. 8 Escalating fluoroquinolone resistance reported in human Salmonella cases 13 was not detected against enrofloxacin in these and other swine-recovered isolates. 9 In addition, resistance to the newer fluoroquinolones, such as danofloxacin, evaluated against the 2008 ISU VDL isolates and by a previous study in Indiana, 9 and ciprofloxacin, evaluated in the 2006 NAHMS study, has not been demonstrated.
The present study identified the changing patterns in antimicrobial resistance and in clinical swine Salmonella serovars identified at the ISU VDL. While these isolates do not represent a random sample of the livestock population and are biased toward animals that had received prior antimicrobial therapy, this review does provide information on the bacteria most likely to be placed under antibiotic selection pressure. Critical monitoring of these patterns is necessary to elucidate changing dynamics of Salmonella pathogens in swine and to protect the human food supply.
Footnotes
a.
Sensititre, Trek Diagnostic Systems, Cleveland, OH.
b.
National Committee for Clinical Laboratory Standards (NCCLS), Wayne, PA.
c.
Clinical Laboratory Standards Institute (CLSI; formerly NCCLS), Wayne, PA.
d.
Version 9.1, SAS Institute, Inc., Cary, NC.
