Abstract
Objective:
This study aims to analyze MRSA isolates from pediatric cancer patients, determine the prevalence of PVL genes and assess their clinical implications.
Introduction:
Methicillin-resistant Staphylococcus aureus (MRSA) infections pose significant challenges in pediatric oncology settings. Understanding the prevalence, genotypic characteristics, and antibiotic resistance patterns of MRSA aids in improving patients’ outcomes.
Methods:
A total of 120 S. aureus isolates from patients receiving chemotherapy for treatment of malignant diseases and developing BSIs during febrile episodes were examined. Isolates were identified using Gram staining, biochemical tests, and VITEK 2-Compact 15. Antimicrobial susceptibility was tested using the Kirby-Bauer disk diffusion method. Molecular characterization included PCR assays for 16S rDNA, multiplex PCR for femA, mecA, and PVL genes, and SCCmec type.
Results:
Out of 120 isolates of Staphylococcus aureus, 97 (80%) isolate possessed mecA gene and were identified as MRSA, MRSA bacteremia harboring PVL gene was detected in cancer patients in Egypt at 26.8% of MRSA isolates. The study identified a higher fatality rate in patients aged 6–10 years (26.7%) compared to other age groups (p < 0.044). Deceased patients exhibited higher leukocyte counts and lower platelet counts. Solid tumor patients had significantly higher neutrophil and monocyte counts. Type II of SCCmec correlated with survival and mortality, with the PVL gene being a significant factor. Type III showed a higher prevalence of femA and mecA among survivors, while Type IVb was associated with better outcomes. Antibiotic susceptibility revealed high resistance to cefoxitin, cefepime, and Tazocin, but better sensitivity to ciprofloxacin and gentamicin, particularly in Types IV and V.
Conclusion:
The study highlights the age-related fatality differences and the impact of HCV infection on survival rates. Hematologic parameters and SCCmec types play a crucial role in patient outcomes. The observed antibiotic resistance patterns necessitate the need for targeted therapies based on MRSA SCCmec types.
Background
Staphylococcus aureus (S. aureus) is one of the most common causes of healthcare and community-acquired infections, varying from soft and skin tissue infections (SSTIs) to life-threatening infections such as septicemia, toxic shock, hospital- and community-acquired pneumonia (HAP and CAP). 1 These infections are associated with substantial morbidity and mortality in hospital settings and communities.2,3
Species of staphylococcus that is commonly associated with increasing bacterial resistance to antibiotics is Staphylococcus aureus is currently included in the ESKAPE group (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species).
In the last few decades, methicillin-resistant Staphylococcus aureus (MRSA) has become an increasingly important cause of healthcare-associated infection. Methicillin-resistant S. aureus, widely known as the superbug, was first reported in the early 1960s. Within years, it rapidly spread in the community and health-care settings. At present, MRSA is endemic in health-care settings globally.4–6 Methicillin resistance in staphylococci has emerged with the acquisition of mec genes (mecA, mecC) located on mobile genetic element known as staphylococcal cassette chromosome mec (SCCmec). 7 SCCmec determinants are classified into various types. To date, at least 13 types of SCCmec elements have been recognized and all SCCmec types have individual characteristics. In general, SCCmec type I, II, and III are distributed in the hospital-associated MRSA (HA-MRSA) and type IV and V are present in the community-acquired MRSA (CA-MRSA).8,9 Panton-Valentine Leukocidin (PVL) is a toxin that forms pores in cell membranes and is produced by the lukF-PV and lukS-PV genes. This toxin specifically targets polymorphonuclear leukocytes, monocytes, and macrophages, leading to their lysis. The result is the release of cytotoxic lysosomal granules from these neutrophils, which can induce both apoptosis and tissue necrosis. 10 There is a well-established link between PVL expression and community-acquired methicillin-resistant Staphylococcus aureus (CA-MRSA) infections. Consequently, PVL is considered a key marker for CA-MRSA, influencing its detection, dissemination, and colonization. 11
This study aims to advance the molecular characterization of Staphylococcus aureus isolates by identifying the presence of femA, mecA, and PVL virulence genes. A triplex PCR assay has been developed to distinguish between hospital-acquired MRSA (HA-MRSA) and CA-MRSA strains. Additio-nally, the study will assess the prevalence of these strains, their antimicrobial susceptibility profiles, and the typing of their staphylococcal cassette chromosome mec (SCCmec) types.
Methods
Study population and bacterial isolates
One hundred twenty S. aureus isolates from pati-ents admitted to National Cancer Institute (NCI), Cairo University, Egypt a tertiary care hospital with a dedicated oncology setup during the period of 22 March 2023 through Feb 2024 and received in the Microbiology laboratory were included in this study. For each case, clinical and epidemiological information were recorded using a structured proforma from the institute documents including demographics (age from 1 to 18 years) and gender (74 males and 46 females); infection sites; reason for specimen collection, length of hospitalization, and recent hospitalization. The isolates were originally recovered through blood cultures
Bacterial growth and identification of Staphylococcus aureus isolate
Bacterial specimens were collected from blood cultures using BD BACTEC™ 9050 Blood Culture System. All samples were initially cultured on blood agar and MacConkey agar No.3 (Thermo Scientific™ Oxoid™, Thermo Fisher Scientific Inc., USA) plates then, all isolates were cultured on UriSelect™ 4 Medium (Bio-Rad Laboratories, Inc) and confirmed as S. aureus by Gram- staining, catalase test, producing coagulase enzyme (by both slide and tube tests), 12 and growing on mannitol salt agar (MSA; Thermo Scientific™ Oxoid™, Thermo Fisher Scientific Inc., USA). 13 Identifi-cation was carried out morphologically and biochemically by standard laboratory procedures 14 and then confirmed using VITEK 2-Compact 15 (biome Rieux, Inc.) located at the NCI.
Phenotypic antimicrobial susceptibility testing
Susceptibility testing of isolates was carried out utilizing VITEK 2-Compact 15 (biome Rieux, Inc.) dried gram positive MIC/Combo panels for gram-positive organisms. The panel of antibiotics used for gram positive organisms included amikacin, amoxicillin-clavulanate, ampicillin-sulbactam, aztreonam, cefoxitin, cefepime, ceftazidime, ciprofloxacin, gen-tamycin, imipenim, meropenem, piperacillin, piperacillin-tazobactam, clindamycin, erythromycin, gatifloxacin, gentamycin, levofloxacin, linezolid, moxi-floxacin, oxacillin, synercid, tetracycline, and vancomycin. Also, Kirby-Bauer Disk Diffusion method (back-up method) was used for other antibiotic resistance phenotypes in addition to the cefoxitin disk screening test. Results were interpreted according to the Clinical and Laboratory Standards Institute (CLSI) standard M100-S25 guidelines. 15 S. aureus MRSA ATCC 43300, S. aureus ATCC 25923, and Escherichia coli ATCC 35218 were employed as quality control strains in antimicrobial susceptibility determination.
Bacterial DNA extraction from cultured colonies
A sample from cultured colonies for each positive blood culture sample in the present study was subjected to one round version of PCR assay targeting 16S r region of Pan bacterial DNA. Sterile UV-irradiated water was used as amplification target in a fourth reaction as a negative control. Staphylococcus aureus subsp. aureus (ATCC® 43300™) was extracted by the same extraction procedure and used as a positive control in the PCR assay. Staph aureus DNA positive control and negative (water) controls were run in each PCR assay.
Bacterial DNA was extracted using QIAamp® DNA Mini Kit, (QIAGEN GmbH, D-40724, Hilden, Germany) according to the manufacturer’s recommendations, DNA extracts were placed on ice and used immediately for PCR assay or stored at −20°C until PCR analysis. The amount of bacterial DNA was measured by spectrophotometry using a Nano-Drop 2000 spectrophotometer (Thermo Scientific/US, Canada) and DNA template was used in the PCR assays.
The amplified region targeted 16S rDNA gene lying between nucleotide positions 18 and 399 bp of the Pan bacterial genome. 16 Primers were supplied in a lyophilized form. Primers were prepared according to manufacturer’s instructions by adding the required volumes of nuclease-free water (supplied with the master mix) to the vials supplied. After preparation, it was stored at −20°C. The primers for 16S rDNA were prepared as described by Harris and Hartley. 16 Primers were provided by HVD life sciences, Vienna.
Primers 16SFa/16SFb and 16SR amplify a 381 bp hypervariable region at the 5′ end of the bacterial 16S rDNA. Comparison of these primer sequences with bacterial 16S rDNA sequences in the GenBank database shows that they are complementary to a wide range of different bacterial genera and species, as previously described. 16
The amplification was done in a final volume of 25 μl of 5-Prime Master Mix containing 0.2 μM of each primer (16S Fa/Fb, 16SR), using the following thermal cycle program (Perkin-Elmer Cetus). The reactions were performed at the following cycling conditions: 94°C for 3 min, 94°C for 30 s, 63°C for 1 min, and 72°C for 1 min. This was repeated for 40 cycles with final extension was carried out at 72°C for 5 min. The samples were overlaid with mineral oil to prevent evaporation. Then 17 µl of PCR product was subjected to electrophoresis on a 2.5% agarose gel (Sigma) in Tris-Acetate buffer (TAE 1X) pH 8.2, stained with 0.5 μg/ml ethidium bromide and examined under UV transillumination and photographed. Product sizes were estimated by comparison with 50 bp DNA ladder (Amersham, UK). Amplified 16S Pan bacterial amplicon was detected at 381 bp.
Multiplex PCR for detection of femA, mecA, and PVL genes
The Multiplex-PCR method used for the detection of femA gene, mecA gene, and PVL gene17–19 showed an advantage over individual conventional PCR assay as it amplified more than one gene in a single test by using more than one set of primers. Therefore, saving time and avoiding contamination during technical procedure while dealing with individual assay. In Multiplex-PCR method used for the detection of femA gene, mecA gene, and PVL gene. The primers were designed by using the primer sequences from various published studies, that were supplied by (HVD life science, Vienna), (Table 1).
Oligonucleotide sequences for the detection of MRSA genes in multiplex PCR.

Standardization of the Multiplex- PCR conditions was adjusted to ensure that, First the sequences of primers used in the triplex assay was assessed using ncbi blast to ensure specificity of each primer and oligoanalyzer 3.1 software (IDTDNA, 2015) to identify proper annealing temperatures and if there were any secondary structures for the selected primers may affect performance of the assay. Second the concentrations of the forward and reverse primers of each gene (PVL—0.1 μM each, mecA—0.2 μM each, and femA—0.2 μM each) were optimized for the multiplex PCR and Third the annealing temperature for the triplex PCR was standardized by a gradient technology. The PCR was carried out in a 25 μl reaction mix which contained 200 μM of dNTPs, 1X PCR buffer, and 0.5U Taq DNA polymerase.
The amplification was done by using a Biometra Gradient (USA) under the following cycling conditions (one cycle of initial denaturation at 94°C for 4 min, 25 cycles of denaturation at 94°C for 30 s, annealing for 30 s at 54°C, and extension at 72°C for 30 s, followed by a final extension at 72°C for 5 min. The Triplex PCR was found to produce excellent results with 1.5 mM MgCl2, 0.5U Taq DNA polymerase, 200 μM dNTPs, 10 ng of DNA, and a 55°C annealing temperature. Each PCR run included positive controls S. aureus ATCC 49775 was used as a positive control for the femA and mecA genes and regarding the PVL gene positive control, the samples are screened for clear positive PVL sample, so it was used as positive control in each run for the PVL gene. As a negative control, the reaction mixture was tested with sterile water. Each gene run in separate tube under same PCR conditions then positive samples were subjected to run in same tube with same conditions.
The amplified products were separated on a 1.5% low melting agarose gel (Sigma). They were electrophoresed (0.5X TAE buffer at 150 V and 90 mA for 30 min), stained with 0.5% ethidium bromide, visualized and recorded by using gel documentation system (BIO-RAD). Fifty bp or one hundred bp ladders (Applied Biosystem-USA) were run as molecular markers. Each PCR run included positive controls S. aureus ATCC 49775 was used as a positive control for the fem A and mec A genes and regarding the PVL gene control, The PVL-positive methicillin susceptible S. aureus (MSSA) strains ATCC49775 and 2002-66, isolated from furuncle (kindly provided by O. Yamasaki), were used as a PVL positive control strain. HA-MRSA strain ATCC700699 (Mu50), whose genome sequence has been determined, was used as a PVL-negative control strain. Laboratory standard strains of MSSA, ATCC29213, and 209P were also used.
Typing of staphylococcal cassette chromosome mec (SCCmec), for all CA- and HA-MRSA isolates
The type of SCCmec mobile genetic resistance element is determined by the combination of the type of ccr gene complex that are represented by the type of the ccr and the class of the mec gene complex. 20 Detection, molecular characterization and associated subtyping of SCCmec types for all MRSA isolates were determined by the method of multiplex PCR (M-PCR) developed by Zhang et al. 20 The amplified product sizes were estimated by comparison with AccuRuler 100 bp Plus DNA RTU Ladder cat.02002-500, size 50 mg/500 ml (Maestrogen, USA) using 2% ethidium bromide-stained agarose gel.
Screening for viral hepatitis biomarkers
All cases were subjected to screening for hepatitis C virus (HCV) and hepatitis B virus (HBV) biomarkers. The screening was conducted using Elecsys HBsAg II (ref. 08814856190) for the detection of HBV surface antigen and Elecsys Anti-HCV II (ref. 08836981190) for HCV antibodies, both supplied by Roche Diagnostics GmbH, Sandhofer Mannheim. The tests were performed on the Cobas® e411 analyzer, provided by Roche Diagnostics, Rotkreuz, Switzerland.
Statistical analysis
The statistical analysis in this study included several methods to assess differences and correlations, with careful consideration of error rates and assumptions at each stage to ensure robustness of results. Descriptive statistics were first computed, summarizing the data using mean ± standard error (SE) for continuous variables. For comparing differences between groups in quantitative data, Analysis of Variance (ANOVA) was employed to test for overall group effects. Following the ANOVA, post-hoc pairwise comparisons were conducted using the Least Significant Difference (LSD) test, which controls for Type I error when making multiple comparisons. The p-value threshold for statistical significance was set at 0.05.
For non-parametric data, Chi-square tests and cross-tabulations were used to assess the relationships between variables. The Chi-square test was performed to evaluate whether observed frequencies differed significantly from expected frequencies under the null hypothesis. The p-value threshold for significance in Chi-square tests was also set at 0.05. To examine the strength and direction of linear relationships between continuous variables, Pearson’s correlation coefficient (r) was used. The correlation coefficient was interpreted with the guideline that values of r > 0.7 indicate strong correlations, values between 0.3 and 0.7 indicate moderate correlations, and values <0.3 indicate weak correlations. All statistical analyses were performed using SPSS for Windows, version 16. A significance level of 0.05 was maintained throughout all tests, and error rates were controlled through the application of appropriate correction methods to minimize both Type I and Type II errors.
Results
Demographic data, fatality, and stage of treatment of cancer patients
The study included 120 cancer patients: 72 with leukemia, 18 with lymphoma, and 30 with solid tumors. The analysis indicates that the fatality rate is significantly higher in the 6–10 years age group at 26.7%, compared to 15.1% in the 1–5 years age group and 17.6% in the 11–18 years age group (p < 0.044). Regarding tumor type, a significant gender disparity is evident: males have a higher proportion of leukemia cases (68.6%) compared to females (48.0%), with this difference being statistically significant (p < 0.005). Conversely, among the 120 patients, 14 showed evidence of HCV infection, which is associated with a significantly lower fatality rate of 7.1% compared to 19.8% in those without HCV (p < 0.005; Table 2).
Demographic data fatality and stage of treatment of cancer patients in the current study.

Chi-Square 0.044, Pearson correlation (p < 0.14, R = 0.224).
Chi-Square 0.005 for the type of tumor.
Effect of fatality, cancer type, and stage of chemotherapy on the hematologic parameters
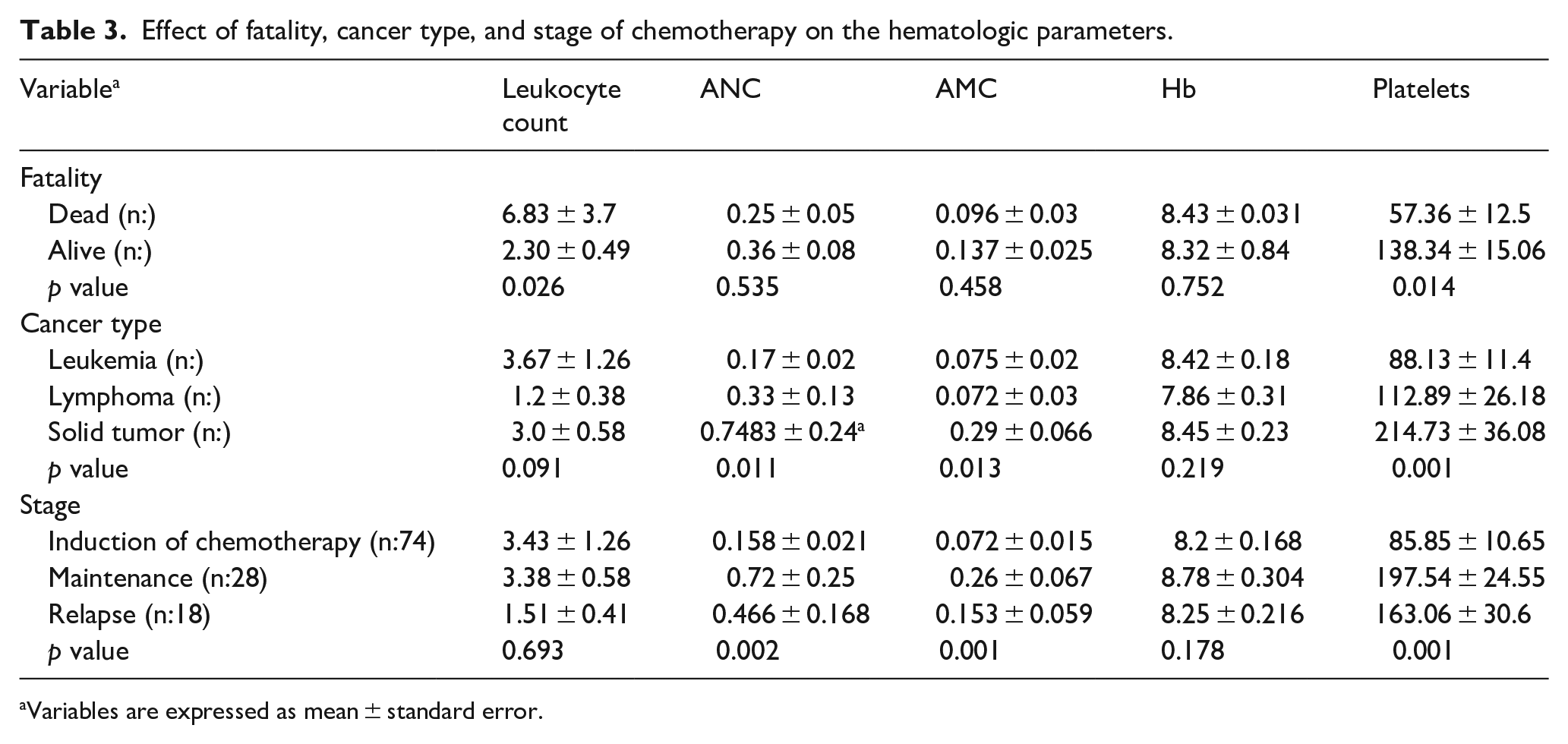
The average leukocyte count was significantly higher in deceased patients (6.83 ± 3.7) compared to those who survived (2.30 ± 0.49), with a p-value of 0.026, indicating statistical significance. Conversely, platelet counts were notably lower in deceased patients (57.36 ± 12.5) compared to survivors (138.34 ± 15.06), with a p-value of 0.014 (Table 3).
Effect of fatality, cancer type, and stage of chemotherapy on the hematologic parameters.
aVariables are expressed as mean ± standard error.
The analysis revealed differences in hematologic parameters across different cancer types. Solid tumor patients exhibited significantly higher absolute neutrophil counts (ANC), and absolute monocyte counts (AMC) compared to those with leukemia (p < 0.001 for ANC and AMC) and lymphoma (p < 0.048 for ANC). Platelet counts were also significantly higher in patients with solid tumors (214.73 ± 36.08) compared to those with leukemia (88.13 ± 11.4) and lymphoma (112.89 ± 26.18), with p-values of <0.001 and <0.01, respectively (Table 3).
The stage of chemotherapy had a significant impact on ANC, AMC, and platelet counts. Patients undergoing maintenance chemotherapy had significantly higher ANC, AMC, and platelet counts compared to those in induction chemotherapy (p < 0.001 for all). However, total leukocyte counts did not differ significantly between the chemotherapy stages (p = 0.693), indicating that while ANC, AMC, and platelets vary significantly with the type of chemotherapy, total leukocyte counts remain stable (Table 3).
Distribution of SCCmec types, femA, mecA, and PVL presence in MRSA isolates by survival status of infected cancer patients
Table 4 provides a detailed view of SCCmec types among cancer patients with Staphylococcus aureus infections, highlighting the association between these types and patient outcomes (alive vs dead) as well as the presence of specific types: femA, mecA, and PVL (Figure 1). All patients were tested positive for Staphylococcus aureus using panbacterial 16S rDNA and the femA gene. Among these patients, 97 had MRSA and 23 had Methicillin-Sensitive Staphylococcus aureus (MSSA). Of these, 22 patients died following a confirmed Staphylococcus aureus infection: 21 cases due to MRSA and 1 case due to MSSA (Table 4).
Distribution of SCC mec Types, femA, mecA, and PVL in Staphylococcus aureus isolates by survival status of cancer patients.

SCCmec types refer to the staphylococcal cassette chromosome mec types. femA, mecA, and PVL are genes associated with Staphylococcus aureus. “No” and “Yes” columns indicate the presence or absence of the respective genes. Total counts for each category are provided. The values represent the number of isolates with each gene combination within the SCCmec genotype.

Gel electrophoresis of MRSA using triplex PCR assay.
By examining the distribution of these types, we can gain insights into their potential impact on patient survival. The data reveals distinct patterns in patient outcomes across different SCCmec types. For type I, the distribution of deaths and survivals does not show significant variation based on the presence of femA or mecA.
Specifically, all patients with type I either lack or have these factors equally distributed between alive and dead groups. type II stands out with a clear correlation to survival outcomes; all patients with this type are either alive or dead, and the presence of PVL is consistently observed in both outcomes. In contrast, type III, which includes 11 deaths and 31 survivors, shows that the presence of femA and mecA is more prevalent among survivors. Additionally, the presence of PVL is associated with both survival and mortality, implying that it is a significant factor in patient outcomes across different survival statuses. Types IV (a, b, c, and d) include very few cases, and none of the patients with types IVa, c, or d are dead. Specifically, type IVb shows that all patients are alive, hinting at a potentially favorable prognosis.
MRSA bacteremia harboring the PVL gene was detected in 26 out of the total 97 (26.8%) MRSA isolates from cancer patients in Egypt (Table 4). Regarding the duration of the episodes, it was significantly longer in PVL-positive MRSA compared to PVL-negative strains (data not shown).
The analysis of the results revealed some interesting associations. femA did not show significant differences between alive and dead patients.
In contrast, mecA was positively correlated with survival, as indicated by a Spearman correlation coefficient of 0.244 (p < 0.015). PVL demonstrated a robust association with patient outcomes. The results from the Chi-square test (p < 0.001) and Spearman correlation (r = 0.54 for dead and r = 0.781 for alive) showed PVL is a significant gene in predicting survival.
CA-MRSA strains were associated exclusively with SCCmec types IVa, IVb, IVc, IVd, and V and characterized by the presence of femA, mecA, and PVL genes. Conversely, HA-MRSA strains were linked to SCCmec types I, II, and III and notable for the absence of the PVL gene. However, a high proportion of HA-MRSA strains exhibit elevated rates of femA and mecA. Specifically, in type I, 5 out of 8 strains (62.5%) were positive for mecA.
For type II, 35 out of 44 strains (79.5%) possed the mecA gene, and in type III, 11 out of 42 strains (73.8%) were mecA-positive. Regarding the femA gene, a substantial proportion of HA-MRSA strains were positive: 7 out of 8 strains (87.5%) in type I, 43 out of 44 strains (97.7%) in type II, and 39 out of 42 strains (92.9%) in type III. Overall, 89 out of 94 HA-MRSA strains (94.7%) carried the femA gene (Table 5).
Distribution of SCCmec types, femA, mecA, and PVL presence in Staphylococcus aureus isolates classified by healthcare-associated (MRS-HA) and community-associated (MRS-CA) infections.

SCCmec types refer to the staphylococcal cassette chromosome mec types. femA, mecA, and PVL are genes associated with Staphylococcus aureus. “No” and “Yes” columns indicate the presence or absence of the respective genes. Total counts for each category are provided. The values represent the number of isolates with each gene combination within the SCCmec genotype.
Antibiotic sensitivity patterns of MRSA Staphylococcus aureus classified based on the SCCmec type
The table illustrates the sensitivity patterns of MRSA Staphylococcus aureus strains with different SCCmec types to various antibiotics. Cefoxitin and cefepime showed low sensitivity across all Types, with only 16.7% of strains sensitive to each, reflecting high resistance rates of up to 100% in some Types and cumulative resistance rates of 83.3% and 82.5%, respectively. Ciprofloxacin, however, demonstrated higher sensitivity with a cumulative rate of 60.8%, particularly in Types IV and V, where sensitivity reaches up to 100%.
Gentamicin sensitivity is moderate, with 38.3% of strains being sensitive and variability observed across Types. Tazocin shows the lowest sensitivity, with only 25.8% of strains responsive and a notable intermediate susceptibility of 9.2%, as indicated by a p-value of <0.001. Clindamycin exhibited moderate sensitivity at 60%, with Types I and II showing the highest sensitivity (Table 6).
Antibiotic sensitivity patterns of MRSA Staphylococcus aureus classified based on the SCCmec genotype.

Tazocin p < 0.001.
Discussion
This study highlights the significant prevalence of methicillin-resistant Staphylococcus aureus (MRSA) among cancer patients, reflecting an ongoing challenge in managing infections in this high-risk group. The observed high fatality rate, especially in the 6–10 years’ age group, highlights the severe impact of MRSA infections in children undergoing cancer treatment. This aligns with previous studies indicating that pediatric patients with malignancies are particularly susceptible to severe infections due to their compromised immune systems and the intensive treatments they undergo.21,22 Effective infection control measures and tailored antimicrobial therapies are crucial in mitigating these risks.
The gender disparity observed in leukemia cases, with a higher prevalence among males, corroborates findings from other studies which have noted sex-based differences in cancer incidence and outcomes. 23 The association between HCV infection and a lower fatality rate is intriguing and warrants further investigation. Some studies suggest that HCV infection might influence immune responses or interact with cancer treatments in ways that affect overall patient outcomes.24–27 This association could reflect complex interactions between viral infections and cancer treatments, affecting disease severity and patient survival.
The significant differences in hematologic parameters between deceased and surviving patients highlight the importance of monitoring these metrics as part of comprehensive patient management. Ele-vated leukocyte counts and decreased platelet counts in deceased patients suggest severe systemic involvement or complications related to MRSA infection, consistent with the literature linking high leukocyte counts and low platelet counts to poor outcomes in severe infections.28,29 The observed variations in hematologic parameters based on cancer type and chemotherapy stage further emphasize the need for individualized patient care and monitoring strategies.30,31
The data illustrate that SCCmec type I dose not exhibit a significant correlation between patient survival and the presence of femA or mecA. This finding suggests that in type I, these genes do not influence survival outcomes, as patients with this type display a balanced distribution of these genes across both alive and dead groups. This is consistent with earlier research indicating that the impact of specific genetic factors on MRSA virulence may vary depending on the SCCmec type. 32
Type II presents a different scenario, with a clear association between type and survival outcomes. All patients with type II are either alive or dead, and the consistent presence of PVL in both outcome groups suggests that PVL is a significant gene in determining survival. It was significantly longer in PVL-positive MRSA compared to PVL-negative strains. The durations of the episodes were longer with PVL-positive strains, which may indicate greater virulence compared to PVL-negative strains. Further studies are needed to confirm this hypothesis. This is supported by studies showing that PVL is linked to more severe disease manifestations, with more invasive disease with poor prognosis and could be a predictor of poor outcomes in MRSA infections.33,34
In type III, the distribution of femA and mecA is notably higher among survivors compared to those who did not survive. The presence of PVL, observed in both survival and mortality outcomes, underscores its role as a crucial factor influencing patient outcomes. This aligns with findings that PVL enhances MRSA virulence, which can complicate infection outcomes. 35
The minimal cases of Types IV (a, b, c, and d) and the absence of deaths among patients with Types IV a, b, and d suggest a potentially favorable prognosis for these genotypes. Specifically, Type IVb, with all patients alive, indicates a possible protective effect associated with this type. This observation may be reflective of the lower virulence associated with certain SCCmec types. 36
PVL, demonstrates a strong association with patient outcomes. indicate that PVL is a significant factor in predicting patient survival. This supports previous research highlighting PVL as a major virulence factor contributing to the severity of MRSA infections and its impact on patient outcomes.32,33
The genetic characteristics of healthcare-associated MRSA (HA-MRSA) and community-associated MRSA (CA-MRSA) reveal significant differences in their epidemiological profiles and associated virulence factors. In our study, CA-MRSA strains are exclusively associated with SCCmec types IVa, IVb, IVc, IVd, and V. These strains are notably characterized by the presence of the femA, mecA, and Panton-Valentine leukocidin (PVL) genes.
The presence of the PVL gene is particularly significant, as it contributes to the enhanced virulence of CA-MRSA, leading to severe skin and soft tissue infections as well as more invasive diseases such as respiratory tract infections, with or without bacteremia, and septic shock.33,35 Although most CA-MRSA strains are generally sensitive to non-beta-lactam antibiotics, there are multidrug-resistant variants, reflecting the adaptability and potential severity of these infections. 33
In contrast, we found that HA-MRSA strains are linked to SCCmec types I, II, and III. These strains are predominantly multidrug-resistant, with SCCmec type II being particularly common among HA-MRSA strains. 32 Notably, HA-MRSA strains lack the PVL gene, differentiating them from CA-MRSA strains. Despite the absence of PVL, HA-MRSA strains exhibit high rates of the femA and mecA genes, which are essential for their antibiotic resistance and survival. Specifically, a significant proportion of HA-MRSA strains are positive for the mecA gene: 62.5% in Type I, 79.5% in Type II, and 73.8% in Type III. Similarly, the femA gene is found in most HA-MRSA strains, with 87.5% of Type I, 97.7% of Type II, and 92.9% of Type III strains being positive for femA.
Overall, 94.7% of HA-MRSA strains carry the femA gene. These findings highlight the distinctive genetic and clinical profiles of HA-MRSA and CA-MRSA. The prevalence of SCCmec types IV and V in CA-MRSA strains, coupled with the presence of the PVL gene, highlights the specific virulence factors contributing to their pathogenicity. On the other hand, the association of SCCmec types I through III with multidrug resistance in HA-MRSA strains, along with the absence of the PVL gene, reflects the adapted nature of these strains to healthcare environments and their role in severe hospital-acquired infections.34,36
Typing of SCCmec elements provides valuable insights into the diversity and resistance profiles of MRSA strains. Our findings that Type II and III correlate with improved survival outcomes suggest these genotypes may have distinct pathogenic characteristics or be associated with less aggressive disease courses compared to other genotypes. This is consistent with previous research indicating that different SCCmec types can influence the virulence and clinical impact of MRSA strains.37,38
The presence of the PVL gene, associated with both survival and mortality, highlights its role in influencing the severity of MRSA infections. PVL-positive MRSA strains are often linked to more severe skin and soft tissue infections, and their impact on systemic infections can vary depending on the host’s immune status and underlying conditions. 39
Different MRSA Types exhibit varying levels of virulence and pathogenicity, which can significantly impact clinical outcomes. Type I, typically associated with community-acquired MRSA, has been linked to less severe infections compared to other types, but its role in virulence remains debated. 40 In contrast, Type II, often found in hospital-acquired MRSA, is associated with more severe infections and higher resistance rates, reflecting its adaptation to hospital environments and prolonged antibiotic exposure. 41 Type III, which includes both community and hospital-acquired strains, demonstrates a complex pattern of resistance and virulence, making it a challenging Type to manage. 42
The lack of fatalities among Type IV strains in our study may indicate that these Types are less virulent or better adapted to certain host conditions. However, this needs further validation, as the low prevalence of these Types in our cohort may limit the ability to draw definitive conclusions. 41
Antibiotic resistance patterns associated with different Types further highlight the need for targeted treatment strategies. The high resistance rates to cefoxitin and cefepime observed across all Types reflect the broader trend of MRSA strains exhibiting multi-drug resistance, making effective treatment more challenging. 43 Conversely, the higher sensitivity of certain Types to ciprofloxacin and gentamicin suggests potential alternatives for treatment but also underscores the need for ongoing susceptibility testing.
Limitation
A key limitation of the study was the small sample size (120 patients) and being a single-center design without involving patients from centers other than NCI, restricts generalization of the findings and include heterogeneous cancer types among participants with different stages and chemotherapeutic protocols. Despite this limitation, our findings provide valuable insights that should be followed by Multi-center study including large number of patients to confirm and approve our results.
Conclusion
In this study, we observed that the highest fatality rate occurred in the 6–10 years age group, with males being more severely affected by leukemia. HCV-infected patients, on the other hand, demonstrated lower fatality rates. Among deceased patients, elevated leukocyte counts, and decreased platelet counts were noted. In contrast, patients with solid tumors had higher neutrophil, monocyte, and platelet levels. Furthermore, maintenance chemotherapy was associated with improved hematological profiles. Regarding bacterial factors, we found that SCCmec Type II was significantly correlated with both survival and mortality, with Panton-Valentine leukocidin (PVL) being a notable factor influencing these outcomes.
Among SCCmec types, Type III showed a higher prevalence of femA and mecA in survivors, whereas Type IV, especially subtype IVb, was linked to better patient outcomes. In terms of antibiotic resistance, MRSA strains exhibited high resistance to cefoxitin, cefepime, and Tazocin, but showed greater sensitivity to ciprofloxacin and gentamicin, particularly in Types IV and V. These findings highlight the complex interplay between age, gender, underlying conditions, and bacterial factors in influencing patient outcomes, emphasizing the importance of tailored treatment strategies to improve survival rates and manage antibiotic resistance effectively.
Footnotes
Acknowledgements
The authors would like to acknowledge Deanship of Graduate Studies and Scientific Research, Taif University, Taif, Saudi Arabia for funding this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by Taif University, Saudi Arabia, Project Number (TU-DSPP-2024-125).
Ethical approval
Approval for the study was granted by the Ethics Committee Approval (CP2302-503-44) from review board (IRB) of NCI, Cairo University. All the measures were performed, and it adhered to the ethical principles outlined in the Declaration of Helsinki. Prior to the commencement of the study, all participants willingly provided written informed consent.
Informed consent
Informed written consent was obtained from all participants after the study objectives were explained and before blood sampling. Confidentiality of patient data was guaranteed.
Consent for publication
The authors declare no conflict of interest.
Trial registration
Not applicable.
Availability of data and materials
All data and materials are available and can be submitted when needed, Corresponding Author is responsible person who should be contacted if someone wants to request the data from this study.
