Abstract
Horses are hosts to 2 types of gammaherpesviruses, Equid herpesvirus 2 and 5 (EHV-2 and EHV-5, respectively). Both EHV-2 and EHV-5 are common in horses in Iceland. An Icelandic EHV-5 isolate was recovered by sequential culture in primary fetal horse kidney and rabbit kidney cells. Glycoprotein B, glycoprotein H, and DNA terminase genes of the isolate were fully sequenced, and the DNA polymerase gene was partly sequenced. To date, the glycoprotein B gene of EHV-5 was the only gene that has been reported to be completely sequenced in addition to small parts of the glycoprotein H, DNA polymerase, and DNA terminase genes. The present report, therefore, is a significant addition to previously reported EHV-5 sequences.
Five herpesviruses are known to infect horses; 2 of them, Equid herpesvirus 2 (EHV-2) and Equid herpesvirus 5 (EHV-5), are closely related 1,7,27 (order Herpesvirales, family Herpesviridae, subfamily Gammaherpesvirinae, genus Percavirus). Foals become infected with EHV-2 and EHV-5 at 1–6 months of age, probably via the upper respiratory tract. 3,15,21 After the primary infection, the virus establishes a latent infection, most likely in the case of EHV-2 in B lymphocytes. 10 Equid herpesvirus 2 has been associated with keratoconjunctivitis, respiratory disease, pneumonia, pharyngitis, fever, enlarged lymph nodes, and lack of appetite. 2,6,8,11,18,20,25 Equid herpesvirus 5 was recently associated with multinodular pulmonary fibrosis. 29
Equid herpesvirus 2 has been detected in the majority of horses worldwide, and EHV-5 is also widespread but less common. 5,9,22,24 The 2 viruses have a strong serologic cross reactivity 1,23 and may coexist in the same horse. 7 Less is known about EHV-5 than about EHV-2, and only a few isolates have been described. 4 The 2 viruses have been shown to be common in Iceland. 28
The current report describes the isolation of EHV-5 from peripheral blood mononuclear cells from a 9-year-old mare (isolate BB5). The peripheral blood mononuclear cells from 2 Listeria monocytogenes—infected horses 16 were cocultured with primary fetal equine kidney cells (EqFKC) as previously described. a–c , 28 Two isolates, BB5 and BB11, were shown, with type-specific polymerase chain reaction (PCR), to be mixtures of EHV-2 and EHV-5.
Plaque purification of EHV-5 was attempted in EqFKCs but was not successful until the BB5 and BB11 isolates were alternately passaged in EqFKCs and rabbit kidney cells (RK-13), d which have previously been used for isolation of EHV-5. 4,12 The RK-13 cells were infected with BB5 or BB11 supernatant from EqFKCs and incubated for 8 days. The supernatant from the RK-13 cells was subsequently harvested and used to infect fresh EqFKCs that were incubated until they showed almost 100% cytopathology. Fresh RK-13 cells were then infected as before with this EqFKCs supernatant and incubated for 8 days when the supernatant was harvested and plaque purification carried out in EqFKCs as previously described. 26 In brief, 80–90% confluent EqFKCs were inoculated with the harvested supernatant for 1 hr at 37°C, the supernatant was then removed, and an overlay of 1% agarose in growth medium with 10% fetal bovine serum was added. Plates were incubated in humidified atmosphere of 5% CO2 at 37°C and observed for cytopathic effect. After approximately 7 days, single plaques were harvested.
Five BB11 plaques and 9 BB5 plaques were picked randomly and identified as EHV-2 or EHV-5 by type-specific PCR as previously described, 28 with a slight modification that only the second PCR was carried out at annealing temperatures 55°C and 50°C for EHV-2 and EHV-5, respectively. All of the 9 BB5 plaques had a single strong band of the correct size (76 bp) with the EHV-5 primers (Fig. 1A) but only a weak band (76 bp), and multiple extra bands with the EHV-2 primers (Fig. 1B). In contrast, the 5 BB11 plaques gave a strong band with the EHV-2 primers (Fig. 1D) but no band with the EHV-5 primers (Fig. 1C). Results indicated that the BB5 plaques were EHV-5, but, for confirmation, approximately 600 bp of glycoprotein B (gB) was sequenced for BB5 plaques 1–3 and 9. The sequences e were compared with published EHV-5 and EHV-2 strains (GenBank accession nos. AF050671 and NC_001650, respectively). The 4 BB5 sequences were identical (GenBank accession no. for 1 sequence, GQ325592) and had 99% sequence similarity with EHV-5 and 80% with EHV-2. One of the 4 isolates, BB5-5, and a strain designated #281, 14 were used for further sequencing.
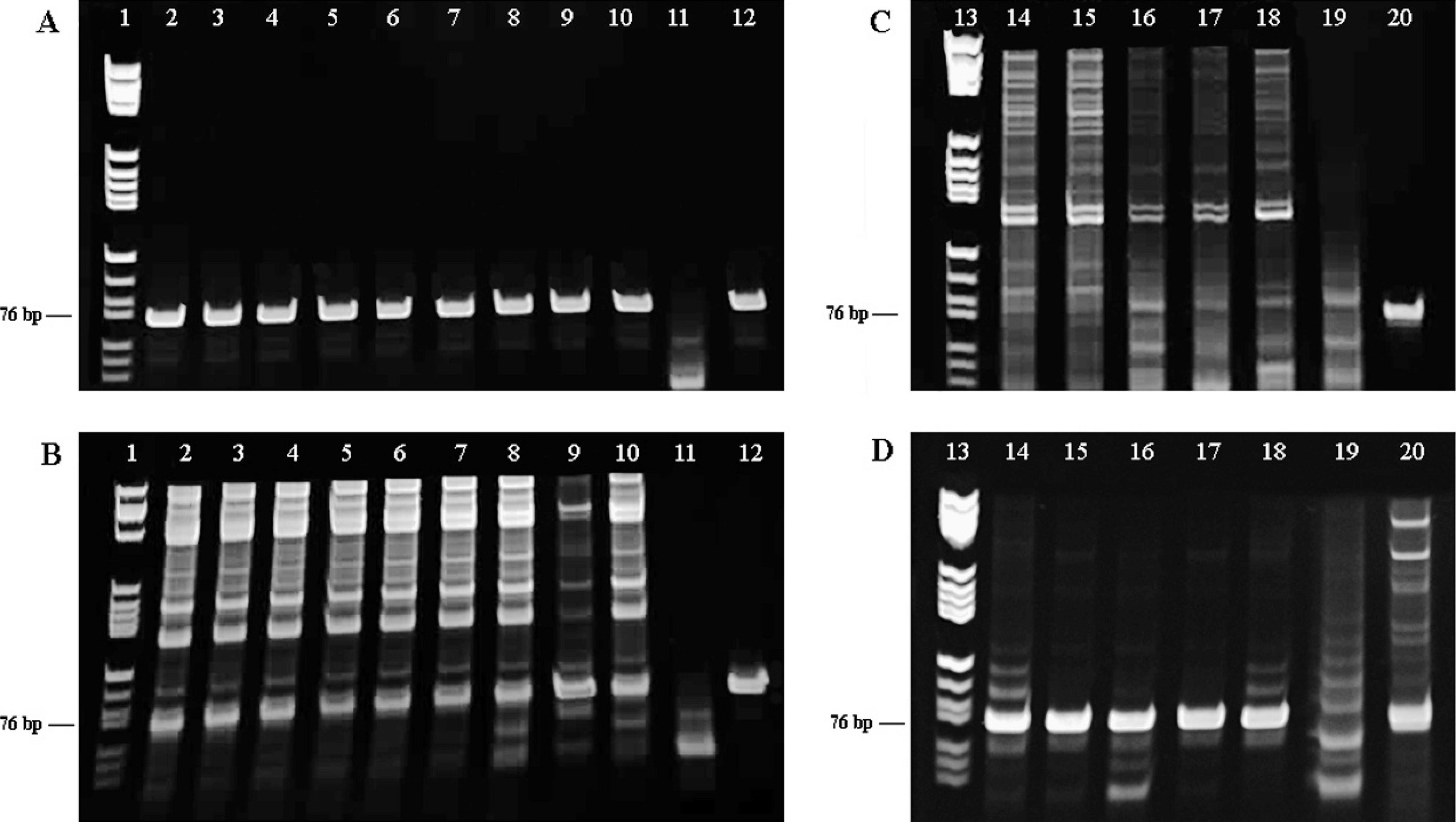
Type-specific polymerase chain reaction for BB11 and BB5 plaques.
Only small fragments of the EHV-5 genome have been published. These are the entire gB 17 and partial sequences for glycoprotein H (gH), 3 DNA polymerase, 13,29 and DNA terminase genes. 19 Primers for the EHV-5 gB were designed from a known EHV-5 sequence (GenBank accession no. AF050671). Amplification for gH, DNA polymerase, and DNA terminase genes was performed with EHV-2 primers, and, if that was successful, then they were used for sequencing. New primers were then designed based on the obtained sequence information. All primers f are listed in Table 1.
The complete gB, gH, and DNA terminase genes were sequenced in addition to 2,444 bp of the approximately 3,000-bp DNA polymerase gene for the BB5-5 strain. Furthermore, the complete gB, in addition to a large part of gH, DNA polymerase, and DNA terminase were sequenced for the #281 strain (Table 2). This is a significant addition to previously described EHV-5 sequences.
The horse population in Iceland has been isolated for at least 1,000 years. 28 Despite this isolation, only a small variance was observed between BB5-5 and #281 strains. In the gB gene, there was 95% homology, 99% for gH, and 98% for both DNA polymerase and DNA terminase genes, which indicated the stability of these viruses. The authors are currently investigating the genetic diversity of native EHV-2 viruses and comparing them with foreign EHV-2 isolates.
When genomic sequences obtained from the BB5-5 isolate were compared with a known EHV-2 strain (GenBank accession no. NC_001650), there was 72% homology for the gB gene, 69% for gH, 84% for DNA polymerase, and 86% for DNA terminase genes. To the authors' knowledge, this is the first reported isolation of EHV-5 in Europe. Although EHV-5 infection in horses is common, there are few reports of isolation of the virus, and few sequences from the EHV-5 genome have been published. The findings in the present study, therefore, are a substantial addition to the knowledge of the sequence of the EHV-5 genome.
Acknowledgements The authors thank Dr. Marco Franchini for providing the #281 strain.
Nucleotide sequence of primers used in polymerase chain reaction and sequencing.
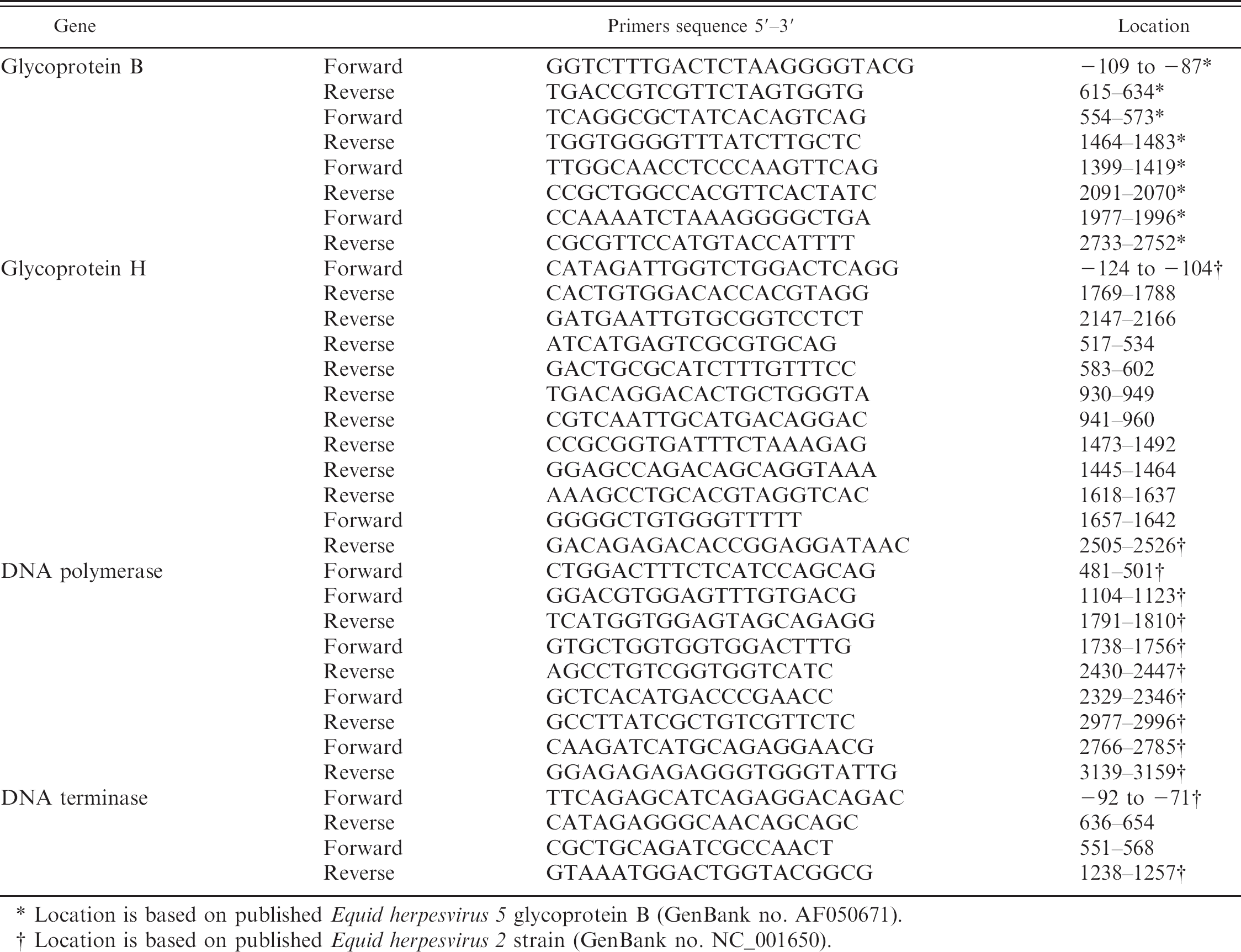
Location is based on published Equid herpesvirus 5 glycoprotein B (GenBank no. AF050671).
Location is based on published Equid herpesvirus 2 strain (GenBank no. NC_001650).
Sequenced genes of Equid herpesvirus 5 isolates.
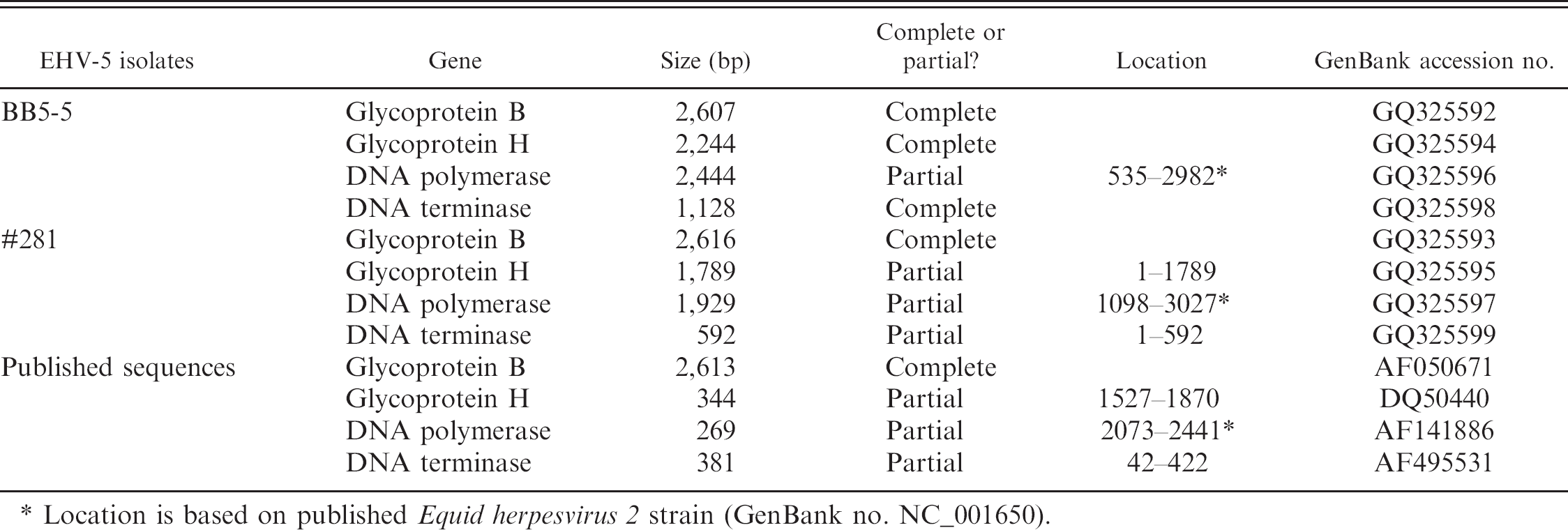
Location is based on published Equid herpesvirus 2 strain (GenBank no. NC_001650).
Footnotes
a.
Culture flasks, Nunc A/S, Roskilde, Denmark.
b.
DMEM, Gibco, Invitrogen, Paisley, Renfrew, UK.
c.
FCS, Gibco, Invitrogen, Paisley Renfrew, UK.
d.
RK-13, American Type Culture Collection, Manassas, VA.
e.
ABI PRISM® 310 Genetic Analyzer, Applied Biosystems, Foster City, CA.
f.
Primers, TAG Copenhagen, Copenhagen, Denmark.
