Abstract
Reported cases of herpesvirus-induced disease are uncommon in most species of reptiles, with the majority of reports in chelonians. Two monitor lizards (Varanus spp.) presented for postmortem examination at the Veterinary Medical Diagnostic Laboratory at the University of Missouri. Tan, 1–2-mm foci were grossly visible on the mucosal surface of the intestine and in the liver. Microscopically, there was multifocal necrosis in the lamina propria of the small intestine and in the liver. Many of the degenerate cells contained large, eosinophilic intranuclear inclusions. Enveloped icosahedral virions consistent with herpesvirus were detected by electron microscopy. A 180-bp DNA fragment was amplified by polymerase chain reaction from samples of small intestine and liver using primers that targeted a portion of the herpesvirus DNA polymerase gene. The sequence of the fragment was determined to be most closely related to Varanid herpesvirus 2 (80% nucleotide identity, 82% amino acid identity). Based on histological and molecular findings, a novel pathogenic herpesvirus of lizards in the family Varanidae is proposed.
Two juvenile monitor lizards with a history of acute death were submitted for postmortem examination at the Veterinary Medical Diagnostic Laboratory at the University of Missouri (Columbia, MO). The submitting veterinarian reported that 3 different monitor species (Varanus spp.) were housed on the owner's premises. Four juvenile animals died within 36 hr. Additional clinical and husbandry information was not available. The specimens weighed 191.3 g and 70.3 g (cases 1 and 2, respectively) and were in adequate body condition. The intestinal tract of case 1 contained a moderate amount of yellow–tan viscous material and multiple 2–3-mm, soft, white particles. Moderate amounts of white, thick material were present in the distal colon. The liver was diffusely pale brown with multiple flat, tan pinpoint foci on the capsular surface. In case 2, 1–2-mm, tan foci were disseminated on the mucosal surface of the intestine. The small intestine contained scant viscous white material. The liver was unremarkable.
On histological examination of pooled tissues, multifo-cal, acute coagulative necrosis was observed in the lamina propria of the small intestine (Fig. 1A). The mucosa in these areas was collapsed, with attenuation of villi and loss of tissue architecture. Multiple, nonsegmented, acoelomate trematodes with a thin cuticle and anterior suckers were present in the intestinal lumen. Acute, multifocal, randomly distributed, hepatocellular coagulative necrosis was present. Affected hepatocytes were characterized by hyper-eosinophilic cytoplasm, pyknotic or karyorrhexic nuclei, and a loss of cellular detail (Fig. 1B). Many hepatocytes contained large, nonstaining cytoplasmic vacuoles. Moderate numbers of degenerate cells in the small intestine and liver contained large, pale, eosinophilic intranuclear inclusions, which resulted in peripheralization of the chromatin (Fig. 1C). The lungs, heart, kidney, gonads, stomach, colon, and brain were unremarkable. The histological diagnosis was multifocal necrotizing enteritis and hepatitis with intranuclear viral inclusions. Salmonella enterica serovar Montevideo was isolated from the liver of case 1. Salmonella enterica serovar Albany and S. enterica Montevideo were isolated by bacterial culture from the intestine of case 2.
Nested polymerase chain reaction (PCR) and electron microscopy were used to evaluate the tissues for viral pathogens. A 180-bp DNA fragment of the herpesvirus DNA polymerase gene, which is highly conserved among many herpesviruses, was amplified from fresh samples of small intestine and liver and sequenced using consensus primers described previously. 8 Briefly, primary PCR mixtures contained 2 upstream primers (DFA: 59-GAYT-TYGCNAGYYTNTAYCC-39 and ILK: 59-TCCTGGA-CAAGCAGCARNYSGCNMTNAA-39) and 1 downstream primer (KG1: 59-GTCTTGCTCACCAGNTC-NACNCCYTT-39). Nested PCR assays were performed on 2 μl of the primary PCR mixture in a 20-μl volume with upstream primer TGV (59-TGTAACTCGGTGTAYG-GNTTYACNGGNGT-39) and downstream primer IYG (59-CACAGAGTCCGTRTCNCCRTADAT-39). Secondary PCR products were analyzed on a 2.0% agarose gel containing 0.5 mg of ethidium bromide per milliliter. Postamplification purification of the DNA was performed, a and the amplicons were sequenced b at the University of Missouri DNA Core Facility (Columbia, MO). The amplicon sequence subsequently was analyzed using commercial software, c and the consensus sequence was used for subsequent manipulations. A search of GenBank revealed that the sequence was most closely related to Varanid herpesvirus 2 (VHV-2; unpublished data, Japan; GenBank accession nos. BAF03633 and AB189433), with 80% nucleotide identity (data not shown) and 82% amino acid identity. Multiple chelonian viruses (Olive Ridley turtle herpesvirus, GenBank accession no. AAC26684; Florida green turtle herpesvirus, GenBank accession no. AAC26682), Varanid herpesvirus 1 (VHV-1; GenBank accession no. AAS17072), Lacerta viridis herpesvirus 1 (Gen Bank accession no. ACD64983), and Gerrhosaurid herpesviruses 1, 2, and 3 (GenBank accession nos. AAL13288, AAL13287, and AAL13289, respectively) demonstrated less than 57% amino acid identity (Fig. 2 and Table 1). These reptilian herpesviruses are classified in the subfamily Alphaherpesvirinae. A dendrogram was prepared using PHYLIP (Phylogeny Inference Package, version 3.5c) 2 to represent the interrelatedness of select currently characterized reptilian alphaherpesviruses (Fig. 3).
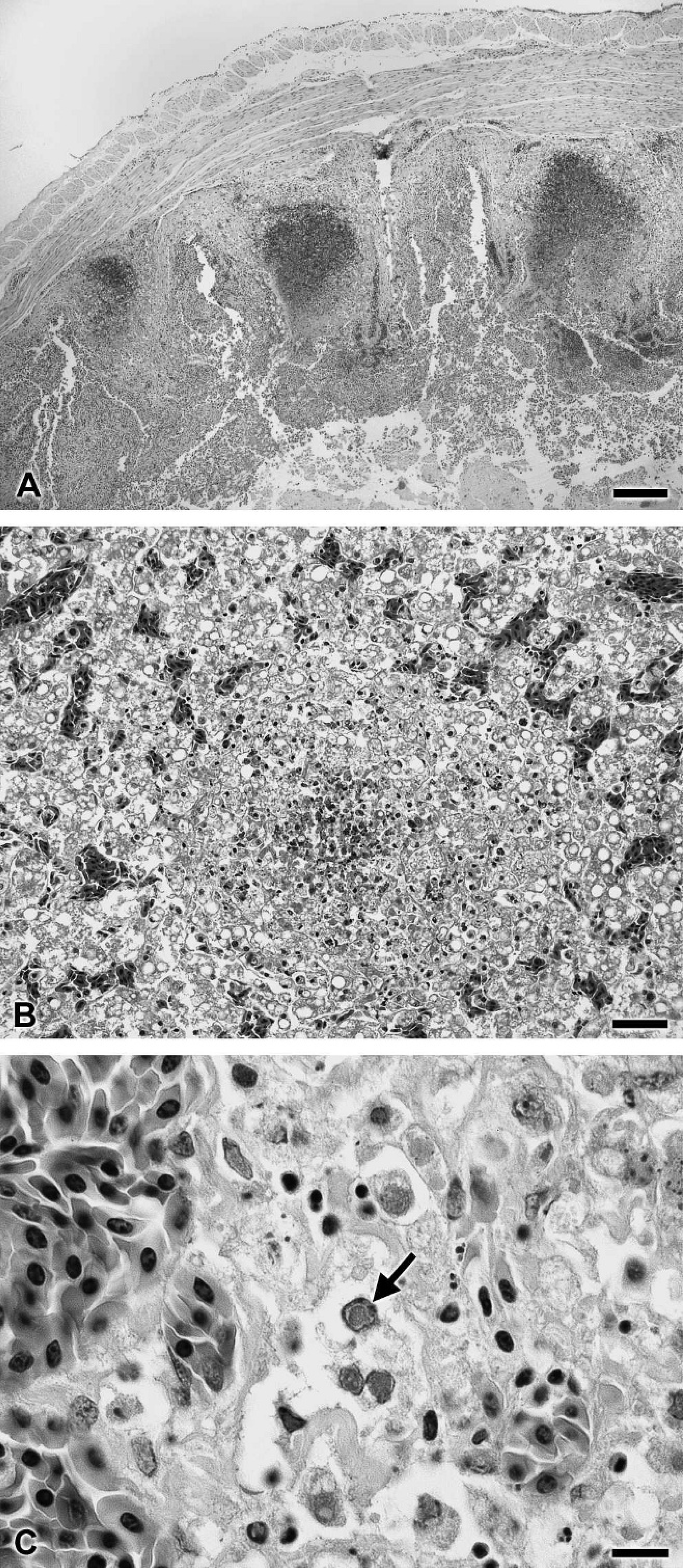
Multifocal necrosis in the small intestine (

Amino acid alignment for select herpesviruses of reptiles. VHV' = new Varanid herpesvirus isolate; VHV1, VHV2 = Varanid herpesviruses 1 and 2, respectively; LHV = Lacerta viridis herpesvirus 1; RHV = Olive Ridley turtle herpesvirus; GHV1, GHV2, GHV3 = Gerrhosaurid herpesviruses 1, 2, and 3, respectively; FHV = Florida green turtle herpesvirus.
Electron microscopy was performed on homogenized, fresh tissues, diluted with 2% phosphotungstic acid. A drop of the solution was absorbed onto a carbon-coated, 400-mesh copper grid and viewed by transmission electron microscopy. Enveloped, 100-nm, icosahedral viral particles consistent with herpesvirus were detected (Fig. 4).
Herpesviruses are enveloped, linear, double-stranded DNA viruses with a 100- to 110-nm icosahedral capsid composed of 162 capsomeres. 1 The host range includes a wide variety of mammals, fish, amphibians, and reptiles. Although numerous herpesviruses have been identified in chelonians, there are few reports in other reptiles. The cases in lizards are limited to green iguanas (Iguana iguana), green lizards (Lacerta viridis), 6 2 red-headed agama lizards (Agama agama), 9 3 plated lizards (Gerrhosaurus major, Gerrhosaurus nigrolineatus), 11 and 3 green tree monitor lizards (Varanus prasinus). 10 In a group of laboratory-reared green lizards, a herpes-like virus was isolated from papilloma-like growths on the dorsum. 6 The 2 red-headed agama lizards were acquired by the Detroit Zoological Park from an animal dealer; necrotic foci were identified in the lung, liver, and spleen with associated intranuclear eosinophilic inclusion bodies. Viral particles consistent with herpesvirus were identified by electron microscopy. 9 The plated lizards were part of a private collection in Minnesota, in which 3 lizards developed stomatitis. Three different herpesviruses, Gerrhosaurid herpesviruses 1, 2, and 3, were identified by genetic sequencing of the DNA polymerase. 11 In that study, Gerrhosaurid herpesviruses 1 and 2 were most closely related to Gallid herpesvirus 2, and Gerrhosaurid herpesvirus 3 was most closely related to Bovine herpesvirus 2. All 3 viruses were classified as alphaherpesviruses. Three green tree monitor lizards from 2 separate collections at the Toronto Zoo and in Florida were examined for proliferative stomatitis (1 case) and squamous cell carcinoma (2 cases). 10 Nearly identical alphaherpesviruses were detected in each of the 3 animals by genetic sequencing of the DNA polymerase and by DNA in situ hybridization. The virus isolated from that study was designated Varanid herpesvirus 1.
The significance of the Salmonella spp. and the trematodes isolated from the monitors in the current report is uncertain. No associated lesions were detected by microscopic examination of fixed tissues. Salmonella spp. are frequently isolated from reptiles, including lizards. 4,5,7 The infection, however, is not typically associated with disease. 3 Similarly, many helminth parasites, including trematodes, have been detected in lizards, but the clinical significance of the infection is not well understood. 3
. Amino acid identity comparison of DNA polymerase fragments from select reptilian herpesviruses. *
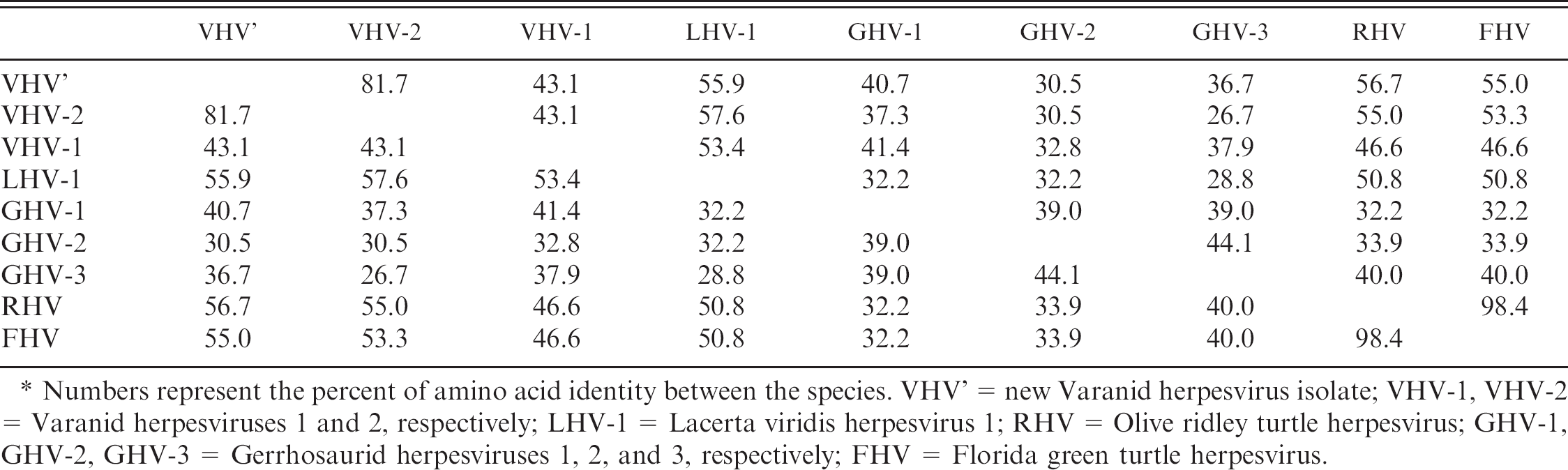
Numbers represent the percent of amino acid identity between the species. VHV' = new Varanid herpesvirus isolate; VHV-1, VHV-2 = Varanid herpesviruses 1 and 2, respectively; LHV-1 = Lacerta viridis herpesvirus 1; RHV = Olive ridley turtle herpesvirus; GHV-1, GHV-2, GHV-3 = Gerrhosaurid herpesviruses 1, 2, and 3, respectively; FHV = Florida green turtle herpesvirus.
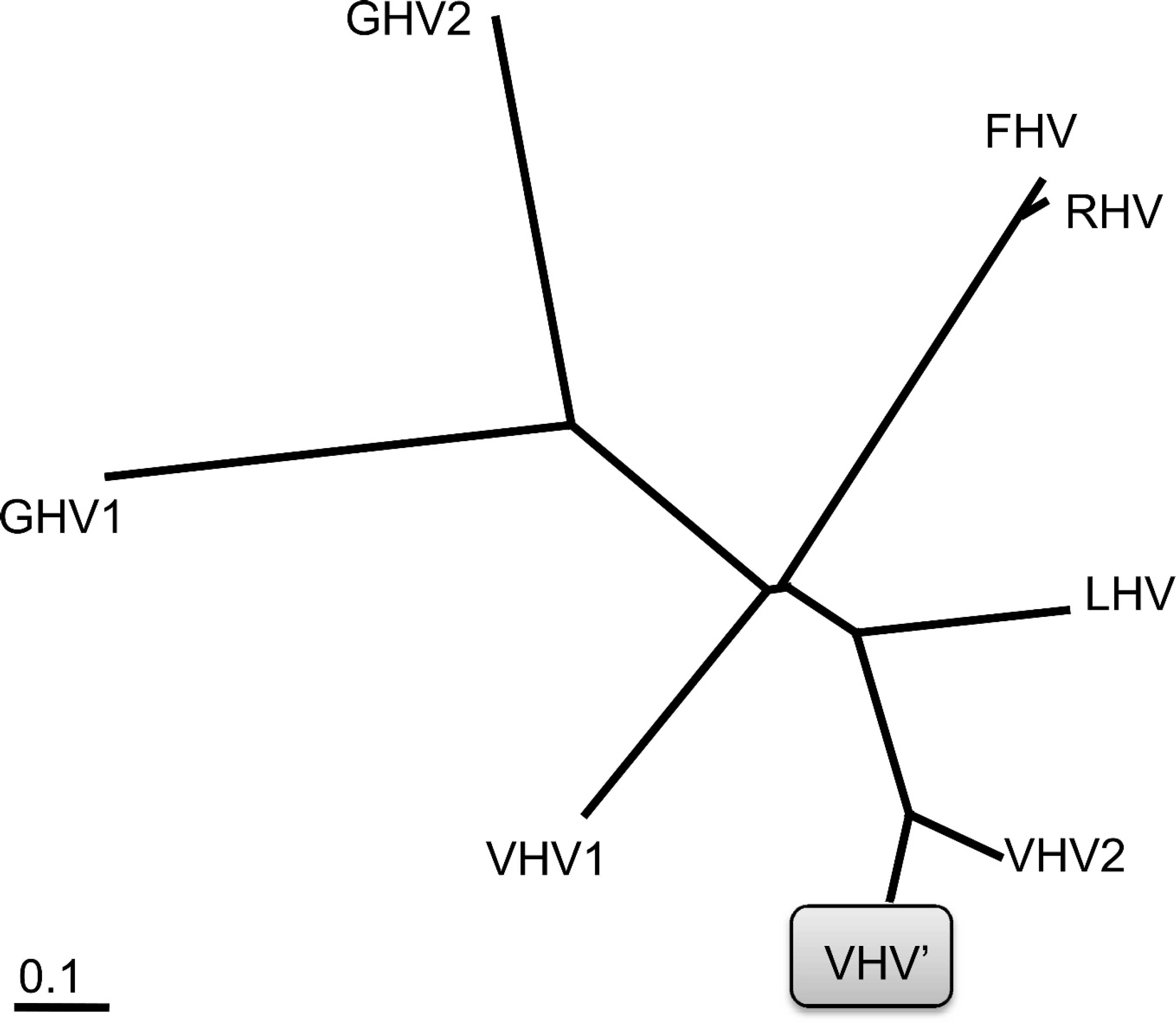
Dendrogram demonstrating relatedness of select reptilian herpesviruses based on GenBank amino acid sequences. VHV' = new Varanid herpesvirus isolate; VHV1, VHV2 = Varanid herpesviruses 1 and 2, respectively; LHV = Lacerta viridis herpesvirus 1; RHV = Olive Ridley turtle herpesvirus; GHV1, GHV2 = Gerrhosaurid herpesviruses 1 and 2, respectively; FHV = Florida green turtle herpesvirus.
The present report describes the partial sequence from a third herpesvirus in a monitor lizard. Based on gross, histological, and molecular diagnostics, the virus is distinct from other reported lizard herpesviruses, including those isolated from other Varanid spp. A novel herpesvirus, Varanid herpesvirus 3, is therefore proposed. The viral sequence is genetically most closely related to VHV-2, which was reported in Japan, but is quite distinct from VHV-1, reported in the United States. To the authors' knowledge, further details regarding VHV-2 have not been published. The virus in the current report was associated with hepatitis and enteritis, whereas VHV-1 was associated with stomatitis and oral squamous cell carcinoma. The origin of infection and evolutionary relationship of reptilian herpesviruses are poorly understood, and further investigation is required to determine the consequence of herpesviral infection in lizards.
Acknowledgements. The authors are grateful to Sunny Younger for the electron microscopy and imaging support and to Howard Wilson for his assistance with preparation of the figures and tables.
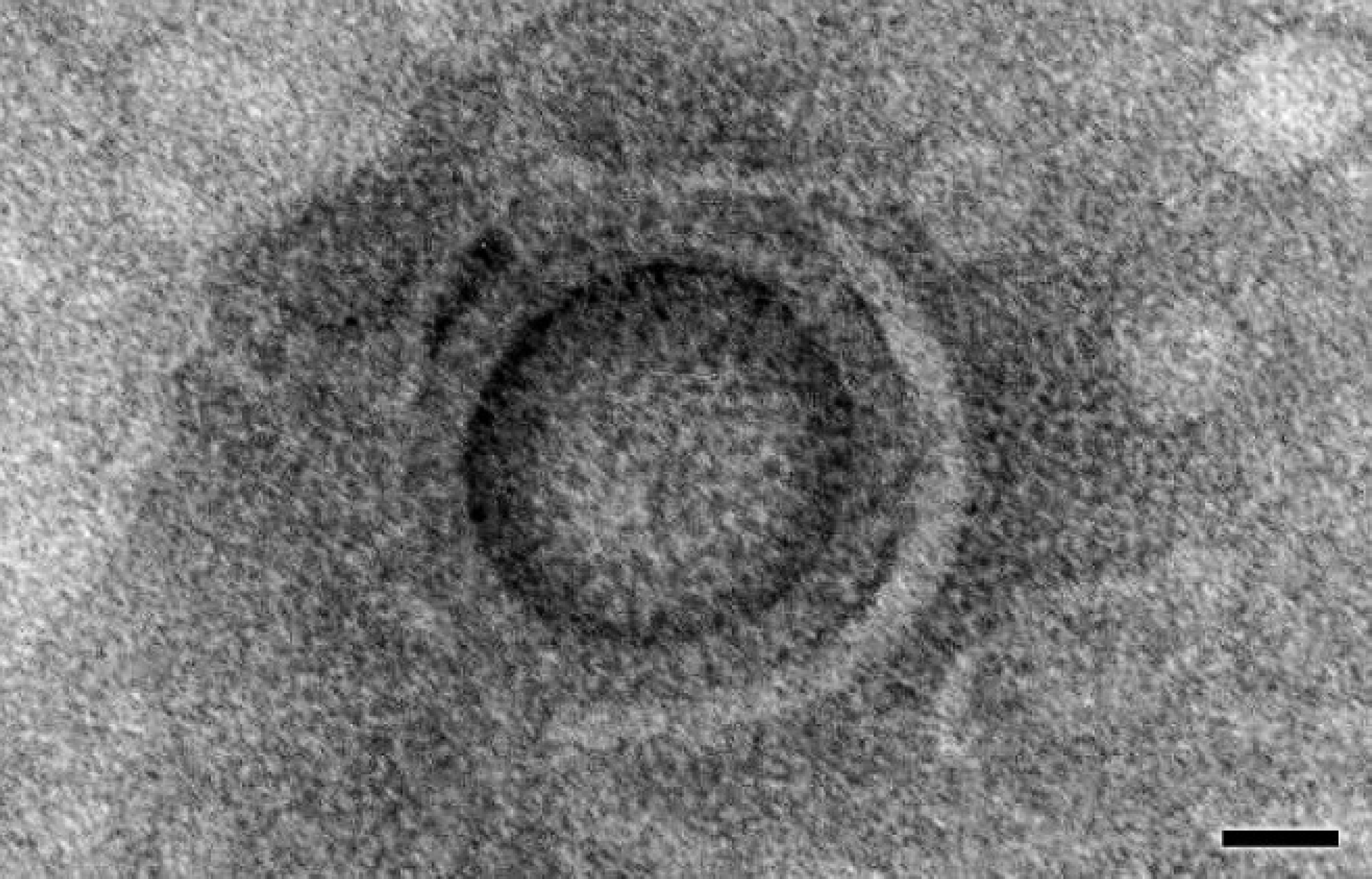
Small intestine. Enveloped, 100-nm, icosahedral viral particle consistent with herpesvirus. Electron micrograph. Bar = 25 nm.
Footnotes
a.
Wizard® SV Gel and PCR Clean-up System, Promega Corp., Madison, WI.
b.
3730 DNA Analyzer, Applied Biosystems, Foster City, CA.
c.
DNASTAR Inc., Madison, WI.
