The Clinical and Laboratory Standards Institute published in 2008 new interpretive criteria for the identification of methicillin resistance in staphylococci isolated from animals. The sensitivity of the 2008 interpretive criteria for mecA gene-positive Staphylococcus pseudintermedius, compared with the previous criteria of 2004, was investigated. Thirty clinical isolates of methicillin-resistant S. pseudintermedius from dogs were used. The presence of the mecA gene was determined by polymerase chain reaction. The minimum inhibitory concentration for oxacillin was determined by broth microdilution. The 2008 breakpoint of 4 μg/ml for methicillin resistance resulted in a diagnostic sensitivity of 73.3% (22/30). The 2004 breakpoint guideline of 0.5 μg/ml resulted in a diagnostic sensitivity of 97% (29/30). For oxacillin disk diffusion, the 2008 interpretive criterion of 10 mm for methicillin resistance resulted in a sensitivity of 70% (21/30). If intermediate isolates (11 or 12 mm) were considered resistant, the sensitivity was 93% (28/30). Application of the 2004 interpretive criterion of 17 mm resulted in a diagnostic sensitivity of 100% (30/30). For cefoxitin disk diffusion, the interpretive criterion of 21 mm for methicillin resistance (as used for Staphylococcus aureus) resulted in a diagnostic sensitivity of 6.7% (2/30). The interpretive criterion of 24 mm (as used for coagulase-negative staphylococci) resulted in a diagnostic sensitivity of 43.3% (13/30). With the use of 2008 interpretive criteria, all 3 tests produced what we consider to be an unacceptable level of false negative results. Our findings also suggest that cefoxitin disk diffusion is an inappropriate screening test for methicillin resistance of canine S. pseudintermedius.
Prompted by the release of the 2008 CLSI interpretive criteria, in this study, we investigated whether the new breakpoints would accurately detect oxacillin resistance in mecA-positive S. pseudintermedius isolated from canine patients at The Ohio State University Veterinary Teaching Hospital (Columbus, OH). The goal of this study was to compare the 2008 and 2004 CLSI interpretive criteria for identifying the oxacillin-resistant phenotype of PCR-confirmed, mecA-positive MRSP clinical isolates from dogs.
The specimens included in this study were selected from a pathogen data bank of clinical isolates collected from February 2007 to September 2008 from the clinical microbiology laboratory at the Veterinary Teaching Hospital at The Ohio State University. The bank included all isolates of coagulase-positive staphylococci that were suspected to be methicillin resistant on the basis of resistance or intermediate susceptibility to oxacillin on disk diffusion test and, in some instances, oxacillin salt agar testing. Subsequently, mecA PCR was performed on all banked isolates. Isolates were frozen at −80°C in trypticase soy broth and 20% dimethyl sulfoxide for up to 20 months and were prepared from a single colony.
The criteria used for selection of isolates included that each isolate 1) originated from a unique canine patient, 2) was associated with clinically apparent infection of the skin (wounds, lesions of pyoderma, incisions) or ear (external ear canal, middle ear), 3) was positive for mecA, and 4) was identified as S. pseudintermedius.
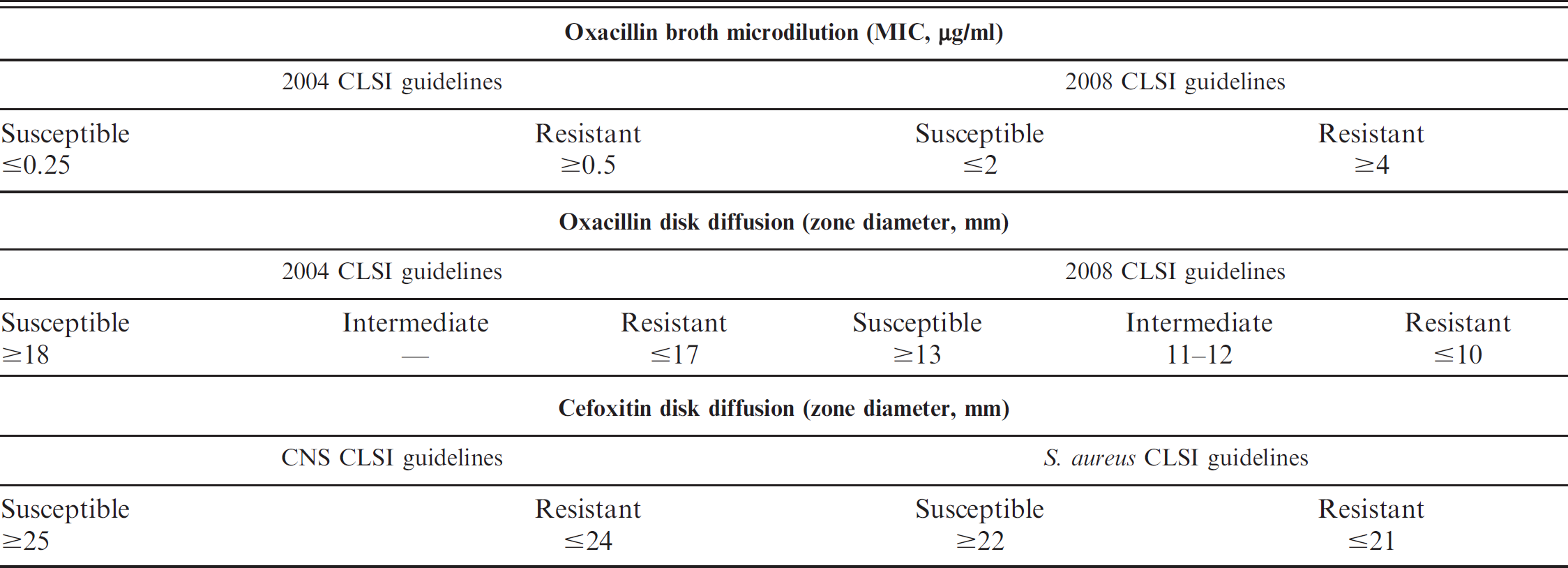
Isolates were evaluated for methicillin resistance using 2008 CLSI S. aureus-based interpretive criteria and 2004 Staphylococcus spp.-based criteria for oxacillin broth microdilution testing and Kirby-Bauer disk diffusion tests. Isolates were also evaluated on the cefoxitin disk diffusion test comparing the interpretive criteria used for S. aureus and coagulase-negative staphylococci (CNS; Table 1).
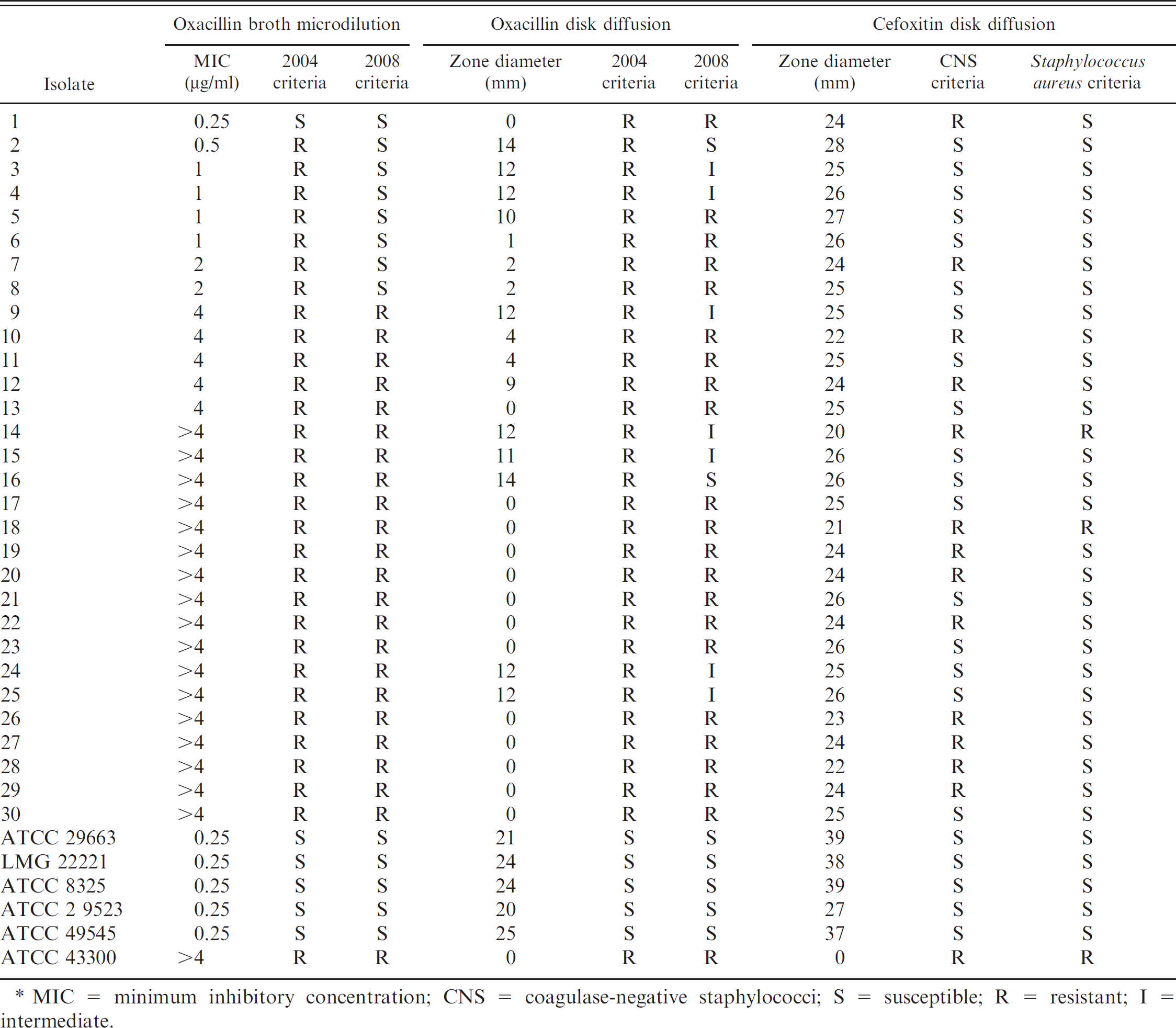
Thirty isolates met all the criteria and thus were included in the study. All isolates were identified as S. pseudintermedius (previously S. intermedius in dogs) and were coagulase positive, VP negative, positive for fermentation of lactose and trehalose, and negative for fermentation of mannitol. The specific results and interpretation of each test for each isolate are presented in Table 2.
The 2008 breakpoint of 4 μg/ml for methicillin resistance resulted in a diagnostic sensitivity of 73.3% (22/30) for mecA-positive S. pseudintermedius. The 2004 breakpoint guideline of 0.5 μg/ml for methicillin resistance resulted in a diagnostic sensitivity of 97% (29/30). The 2008 CLSI interpretive guideline of 10 mm for methicillin resistance resulted in a diagnostic sensitivity of 70% (21/30) for mecA-positive S. pseudintermedius. If intermediate isolates (11- or 12-mm zone of inhibition) are considered resistant (as might occur in clinical interpretations), the diagnostic sensitivity was 93% (28/30).
Application of the 2004 interpretive criterion of 17 mm for methicillin resistance in S. aureus resulted in a diagnostic sensitivity of 100% (30/30). The interpretive criterion of 21 mm for methicillin resistance (as used for S. aureus) resulted in a diagnostic sensitivity of 6.7% (2/30) for mecA-positive S. pseudintermedius. The interpretive criterion of 24 mm for methicillin resistance (as used for coagulase-negative staphylococci) resulted in a diagnostic sensitivity of 43.3% (13/30).
The 2008 interpretive criteria failed to detect methicillin resistance in some clinical mecA-positive isolates of S. pseudintermedius from the skin and ears of dogs. The most sensitive method for phenotypic recognition of methicillin resistance in this limited cohort of mecA-positive S. pseudintermedius isolates was oxacillin Kirby-Bauer disk diffusion under the 2004 interpretive criteria, with 100% sensitivity; the 2008 criteria failed to identify 2 methicillin-resistant isolates. The 2008 interpretive criteria for oxacillin broth microdilution failed to identify methicillin resistance in 26.6% of isolates in this study. The 2004 interpretive criteria were more sensitive, although methicillin resistance was unrecognized in a single isolate (isolate 1).
Isolate 1, though mecA-positive, was not identified as methicillin resistant with oxacillin broth microdilution 2004 nor 2008 interpretive criteria. Discordance between the presence of mecA and the absence of a corresponding oxacillin-resistant phenotype has been recognized previously in veterinary isolates of S. aureus, S. intermedius, and S. schleiferi,
3
as well as human isolates of S. aureus.
11
However, in isolate 1, methicillin resistance was detected via oxacillin disk diffusion (2004 and 2008 interpretive criteria) and via cefoxitin disk diffusion (2004 CNS interpretive criteria), which is consistent with a PBP2a-positive phenotype.
mecA-negative clinical isolates were not evaluated in the current study. Therefore specificity and expectation for false positive results for these methods was not determined. A cohort of 44 mecA-negative canine S. pseudintermedius isolates was recently evaluated.
2
The specificity of oxacillin broth microdilution was 99.4%, and 100% with 2004 and 2008 criteria, respectively.
2
The superior sensitivity of the 2004 oxacillin Kirby-Bauer disk diffusion criteria was similarly demonstrated in 230 mecA-positive canine S. pseudintermedius isolates in a recent study.
2
Oxacillin Kirby-Bauer disk diffusion resulted in a sensitivity of 99.1% and 85.1% with 2004 and 2008 criteria, respectively.
2
Bemis et al.
2
evaluated cefoxitin disk diffusion in 88 mecA-positive and 204 mecA-negative canine S. pseudintermedius isolates. Although highly specific (100%), the cefoxitin Kirby-Bauer disk diffusion method was also found to be the least sensitive method; sensitivities were 85.3% and 70.9% under 2004 and 2008 criteria, respectively.
2
Of note, the diagnostic sensitivities for cefoxitin disk testing reported previously
2
(85.3% and 70.9%) are higher than that of the work presented here (6.7% and 43.3%). The reason for this difference is speculative and might reflect increased accuracy via larger cohort size of that study or regional strain differences. The study also did not restrict number of isolates included per patient; however, the frequency of such inclusions was not reported. Inclusion of concurrent or subsequent isolates from the same individual might have inadvertently selected for clonal isolates that were cefoxitin-sensitive. However, clonality is not restricted to the individual, and evaluation of clonality was not undertaken in this study, nor in a previous study.
2
Despite this disagreement in sensitivity, the work presented in the current study confirms the superior sensitivity of oxacillin Kirby-Bauer disk diffusion interpreted with 2004 CLSI criteria and the inferior sensitivity of cefoxitin disk diffusion in the evaluation of methicillin resistance in canine-origin S. pseudintermedius isolates in a geographically separate population.
With the use of 2008 interpretive criteria, all 3 test methods evaluated would have given what we consider to be an unacceptable level of false-negative results, which could have led to inappropriate treatment if susceptibility to other β-lactam antibiotics was identified and reported in isolates falsely determined to be methicillin susceptible. Our study suggests that application of the 2004 CLSI criteria for oxacillin disk diffusion and oxacillin broth microdilution test produces superior sensitivity for the detection of methicillin-resistant S. pseudintermedius compared with the 2008 criteria. Furthermore, the findings presented in the current study demonstrate that an unacceptably higher rate of false-negative results occur with application of the 2008 CLSI criteria, consistent with recent work.
2
Therefore, it is advised that veterinary diagnostic laboratories abstain from using the 2008 CLSI criteria when evaluating methicillin resistance in canine S. pseudintermedius. The 2004 CLSI criteria for oxacillin Kirby-Bauer disk diffusion and oxacillin broth microdilution should be used to evaluate methicillin resistance in canine S. pseudintermedius isolates. In addition, findings in the present study confirm that cefoxitin disk diffusion, regardless of CNS or S. aureus CLSI interpretive guidelines, is an inappropriate screening test for methicillin resistance in S. pseudintermedius isolated from dogs.
Acknowledgements. The authors thank Narry Tiao and Jennifer Mathews for performing mecA PCR and Nancy Martin for assistance with antimicrobial susceptibility.