Abstract
Clinical and Laboratory Standards Institute interpretive breakpoints for in vitro susceptibility tests that predict mecA-mediated oxacillin resistance in Staphylococcus pseudintermedius isolates from animals have been changed twice in the past decade. Moreover, there are no counterpart recommendations for human isolates of S. pseudintermedius. Individual medical and veterinary laboratories variably use interpretive breakpoints identical to those recommended for use with Staphylococcus aureus or identical to those recommended for use with coagulase-negative staphylococci. The purpose of the current study was to examine correlations between oxacillin disk diffusion, oxacillin gradient diffusion, oxacillin microbroth dilution, and cefoxitin disk diffusion tests used to predict mecA-mediated resistance in S. pseudintermedius and to retrospectively estimate, from disk diffusion zone diameter measurements, the prevalence and rate of increase of oxacillin resistance among canine S. pseudintermedius isolates submitted to a veterinary teaching hospital laboratory. Oxacillin disk diffusion zone diameters of ≤17 mm and oxacillin minimum inhibitory concentrations of ≥0.5 μg/ml were highly correlated with detection of mecA in canine S. pseudintermedius isolates by polymerase chain reaction. MecA-mediated resistance among S. pseudintermedius isolates from dogs increased from less than 5% in 2001 to near 30% in 2007. More than 90% of the methicillin-resistant S. pseudintermedius isolates in 2006 and 2007 were also resistant to representatives of ≥4 additional antimicrobial drug classes. Cefoxitin disk diffusion with the resistance breakpoint set at ≤24 mm significantly underestimated the presence of mecA in S. pseudintermedius.
Introduction
Because of its greater stability, oxacillin has long been used as a surrogate antimicrobial drug for in vitro detection of staphylococcal resistance to the isoxazolyl class of penicillinase-resistant penicillins (typified and first represented by methicillin). Methods and criteria to interpret oxacillin susceptibility tests with Staphylococcus pseudintermedius have varied greatly, both within and between laboratories, making it difficult to recognize broadly emerging patterns of oxacillin resistance in this species. Most veterinary laboratories use disk diffusion or microbroth dilution methods for oxacillin susceptibility testing. Partial-range commercial microbroth dilution susceptibility test panels have not always contained a sufficient range of oxacillin concentrations to detect low levels of oxacillin resistance in S. pseudintermedius according to prevailing breakpoint recommendations. 18 Staphylococcus pseudintermedius is occasionally isolated from humans, and zoonotic transfer from dogs has been implicated. 6,12,22,23 There are currently no recommendations for interpretation of oxacillin susceptibility tests performed on human isolates of coagulase-positive staphylococci other than S. aureus. 8 However, the Clinical and Laboratory Standards Institute (CLSI; formerly National Committee for Clinical Laboratory Standards) recently recommended that oxacillin susceptibility of S. pseudintermedius and other veterinary isolates of coagulase-positive staphylococci be determined using breakpoints equivalent to those recommended for human and veterinary isolates of S. aureus. 7 The cefoxitin disk diffusion screening test also was noted as equal or superior to oxacillin disk diffusion for predicting methicillin (i.e., mecA-mediated) resistance in S. aureus and coagulase-negative Staphylococcus spp. isolated from animals, thus creating uncertainty as to use of cefoxitin disk testing with S. pseudintermedius. The present study examined frequencies of oxacillin resistance in canine isolates of S. pseudintermedius from clinical laboratory submissions from 2001 to 2007 and compared the use of in vitro susceptibility tests as predictors of mecA gene presence in a collection of canine S. pseudintermedius isolates.
Materials and methods
Source of isolates
Staphylococcus pseudintermedius isolates were obtained from clinical samples submitted to the University of Tennessee Clinical Bacteriology Laboratory (UTCVM; Knoxville, TN). An archived collection of >700 strains included selected, predominantly oxacillin-resistant isolates dating back to 1995; all oxacillin-resistant isolates since 2001; and all oxacillin-susceptible isolates from the year 2006. Archived bacteria were stored in cryopreservative at −80°C.
Bacterial identification
Isolates were obtained by routine aerobic bacterial culture as previously described. 13 Suspect colonies on nonselective blood agar plates were usually opaque, off-white, ≥1 mm in diameter, and surrounded by a single or double zone of hemolysis. Presumptive biochemical identification was adapted from standard diagnostic tables using conventional methods. 2,13 Routine tests included tube coagulase; acetoin production; and maltose, lactose, and trehalose fermentation. Occasional isolates were also tested for pyrrolidonyl arylamidase, polymyxin B susceptibility, and with a commercial biochemical test kit. a
Susceptibility testing
Standard reference disk diffusion and microbroth dilution susceptibility tests were performed manually and interpreted visually according to CLSI guidelines. 7 Oxacillin gradient diffusion susceptibility tests were performed according to the manufacturer's specifications. b Antimicrobial-containing disks at CLSI recommended concentrations, 7 , c microbroth dilution panels, d gradient diffusion strips, b cation-adjusted Mueller-Hinton broth, d and agar with e and without c supplemental 2% NaCl were obtained from commercial vendors. Microbroth dilution results were captured for further analytic confirmation and archived with a digital imaging device. f Reference quality control (QC) strains of oxacillin-susceptible S. aureus, oxacillin-resistant (mecA-mediated) S. aureus, and Escherichia coli were obtained from a commercial source. g Susceptibility tests, using QC strains, were performed weekly and with each new lot of media and antimicrobial disks.
Polymerase chain reaction and DNA sequencing
Species identity of S. pseudintermedius was confirmed by DNA sequencing of selected isolates. DNA was extracted from a 1-ml suspension of bacteria (ca. 5 × 10 9 cfu/ml) by cell disruption with glass beads. h Universal eubacterial primers, Broad 1 and Broad 2, 21 were used for polymerase chain reaction (PCR) amplification of the (ca. 1,500 base pair [bp]) portion of the 16S ribosomal RNA (rRNA) gene. The sequence of the entire mecA gene was obtained from selected samples as previously described. 5 The mecA screening was done using primers designed in the current study: GAP-FWD-TCAGAGTTAATGGGACCAAC and GAP-REV-GACTGAACGTCCGATAAAAA. These primers amplified a 562-bp fragment of the mecA gene. Polymerase chain reaction was performed with a commercial Taq polymerase premix i containing reaction buffer and nucleotides. Thermocycler conditions for 16S ribosomal DNA included 1 cycle of denaturation and enzyme activation at 95°C for 1.5 min, 30 cycles of annealing at 50°C for 30 sec, extension at 72°C for 2.5 min, and denaturation for 1 min at 94°C. This was followed by a final cycle of annealing at 50°C for 2 min, extension at 72°C for 5 min, and holding at 4°C. Thermocycler conditions for mecA PCR were 95°C for 90 sec; 55°C for 30 sec, 68°C for 120 sec, 94°C for 30 sec for 35 cycles; then 55°C for 30 sec and 68°C for 5 min. The PCR amplification products were subjected to electrophoresis in 1 % agarose gel containing 0.5 mg/ml ethidium bromide and were visualized by ultraviolet transillumination. The PCR products of expected sizes were treated to destroy single-stranded DNA j and submitted to the University of Tennessee core DNA sequencing facility. The PCR primers were used for direct DNA sequencing of PCR amplification products.
Data analysis
Nucleotide sequence data were analyzed using a commercial software program. k The mecA and 16S rRNA gene sequences of clinical isolates were compared to sequences in the GenBank database using the BLASTn (nucleotide Basic Local Alignment Search Tool; http://www.ncbi.nlm.nih.gov/blast/Blast.cgi) algorithm. 1 Isolates were from multiple body sites, but only 1 isolate of each unique antimicrobial susceptibility profile was included per patient. Statistical analysis was performed using commercially available software. l
Results
All S. pseudintermedius isolates since 2001 (excluding <50 in 2001 and 8 in 2003) were tested for oxacillin susceptibility using the disk diffusion method. Growth inhibition zone diameters were measured and recorded; interpreted results were reported; and selected isolates, predominantly resistant isolates, were archived. In 2006, oxacillin gradient diffusion minimum inhibitory concentrations (MICs) were determined for a subset of 137 isolates (88 resistant, 49 susceptible). The resulting MICs were compared to disk diffusion zone diameters and to mecA gene detection by PCR (Fig. 1). There was a linear correlation (r = 0.68) between disk diffusion zone diameters and gradient diffusion MICs, and both tests showed high categoric agreement with mecA detection. In all isolates interpreted as resistant using a zone diameter breakpoint of ≤17 mm, mecA was found; mecA was also found in all isolates with MICs reported as ≥0.5 μg/ml, and in 5 isolates for which MICs of 0.38 μg/ml were converted to the next highest level for final interpretation, as recommended by the manufacturer. However, mecA was not detected in any isolates that were interpreted as susceptible using gradient diffusion but was present in 2 isolates interpreted as susceptible (breakpoint ≥18 mm) using disk diffusion.
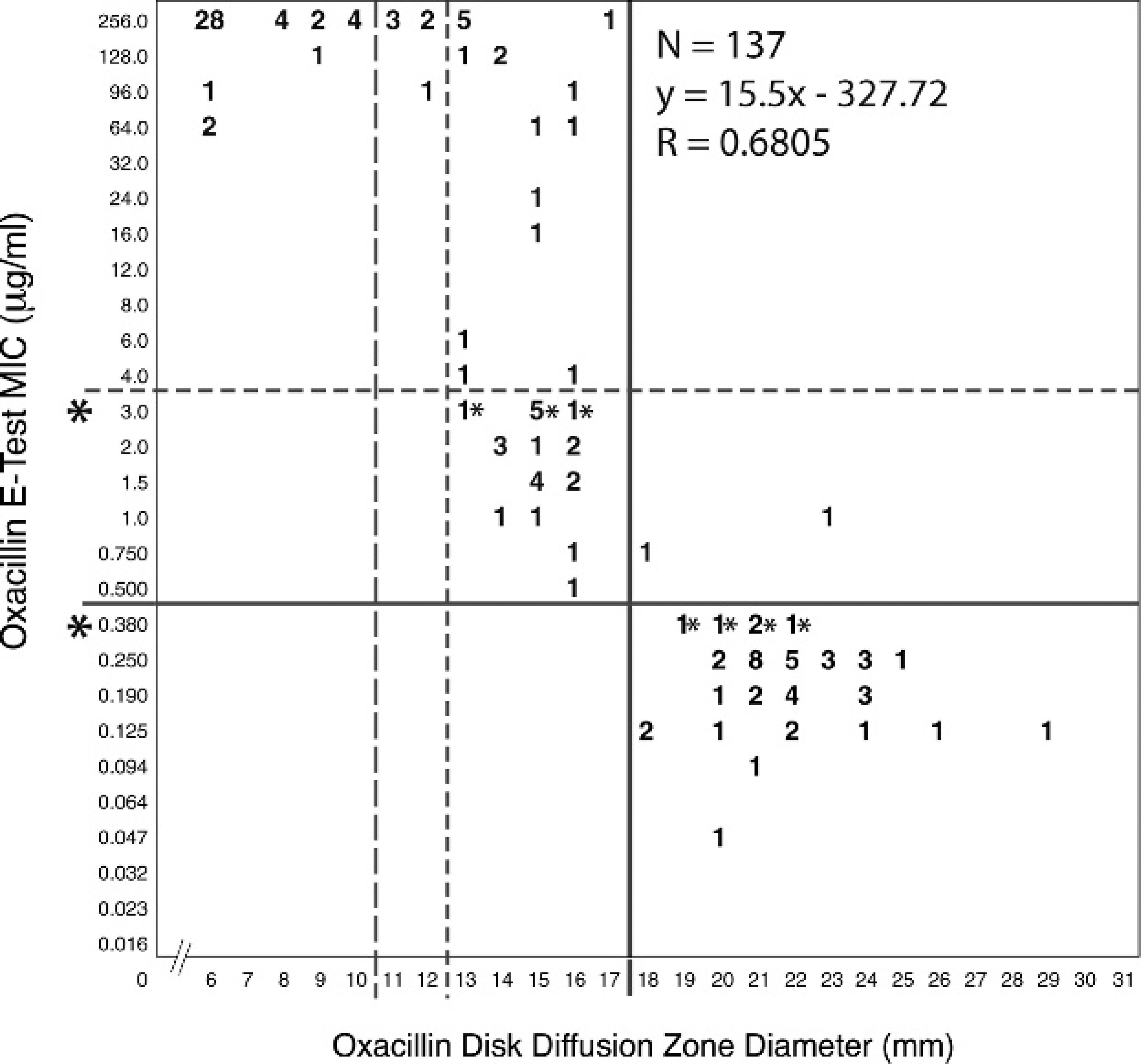
Correlation between gradient diffusion minimum inhibitory concentration (MIC) and disk diffusion zone diameters for oxacillin. Solid lines = former (2004–2008) Clinical and Laboratory Standards Institute (CLSI) breakpoints; broken lines = current (2008) CLSI breakpoints. *Gradient diffusion MICs that lie between values found in the standard 2-fold dilution series are rounded to the next highest value for interpretation.
Oxacillin disk diffusion susceptibility records from 2001 to 2007 were examined using ≤17 mm as the interpretive breakpoint (Fig. 2). The frequencies of oxacillin resistance among canine S. pseudintermedius isolates increased from <5% in 2001 to near 30% in 2008.
Cefoxitin disk diffusion zone diameters were compared with those of oxacillin for 292 canine S. pseudintermedius isolates obtained in 2006 (Fig. 3). Cefoxitin disk diffusion test sensitivity and specificity for predicting oxacillin susceptibility in this data set, with breakpoints at ≤24 mm for resistant and ≥ 25 mm for susceptible, were 100% and 54%, respectively. The positive predictive value was 84%, and the negative predictive value was 100%.
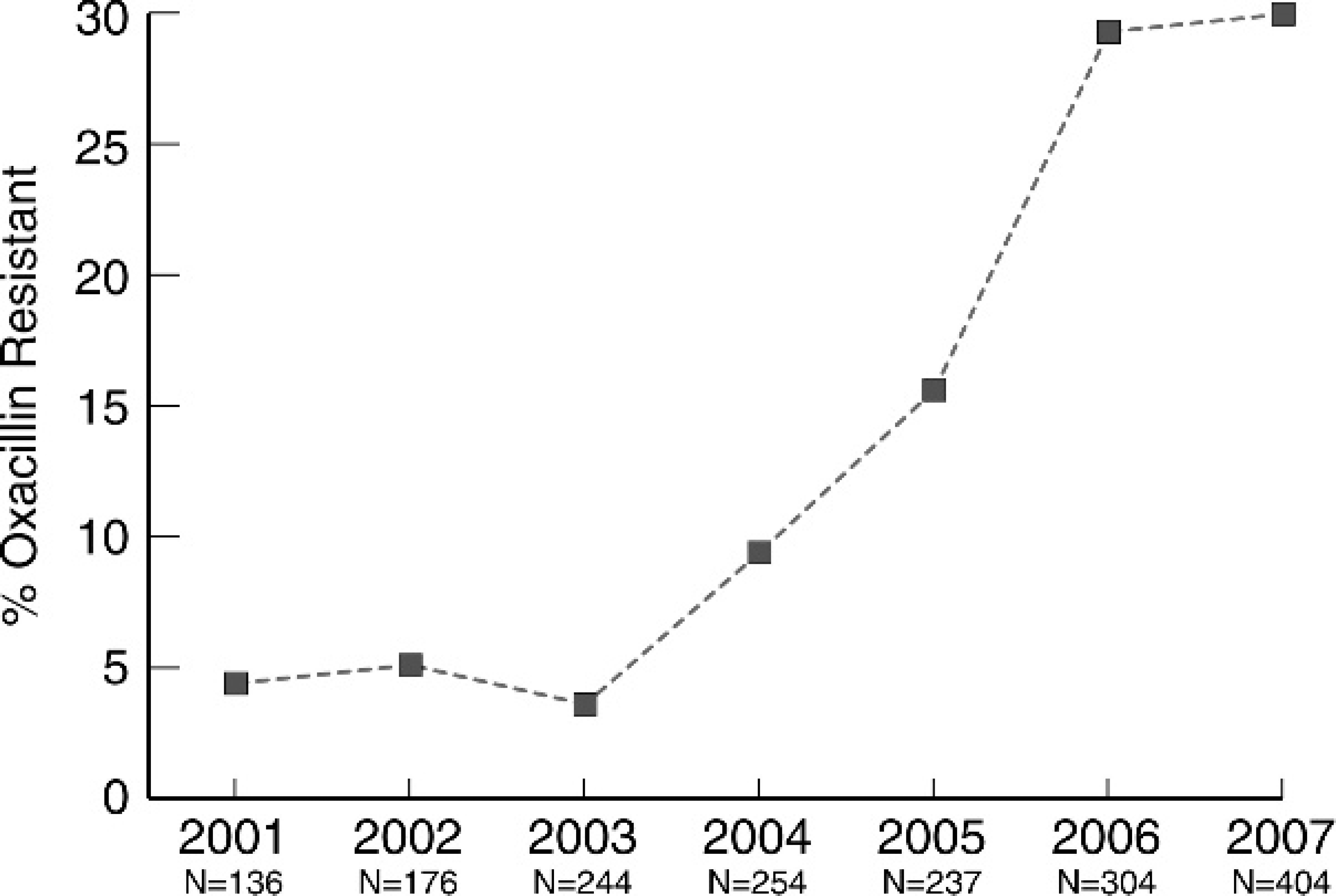
Frequencies of oxacillin resistance among Staphylococcus pseudintermedius isolates from dogs from 2001 to 2007. Disk diffusion method; resistant: ≤17 mm, susceptible: ≥18 mm.
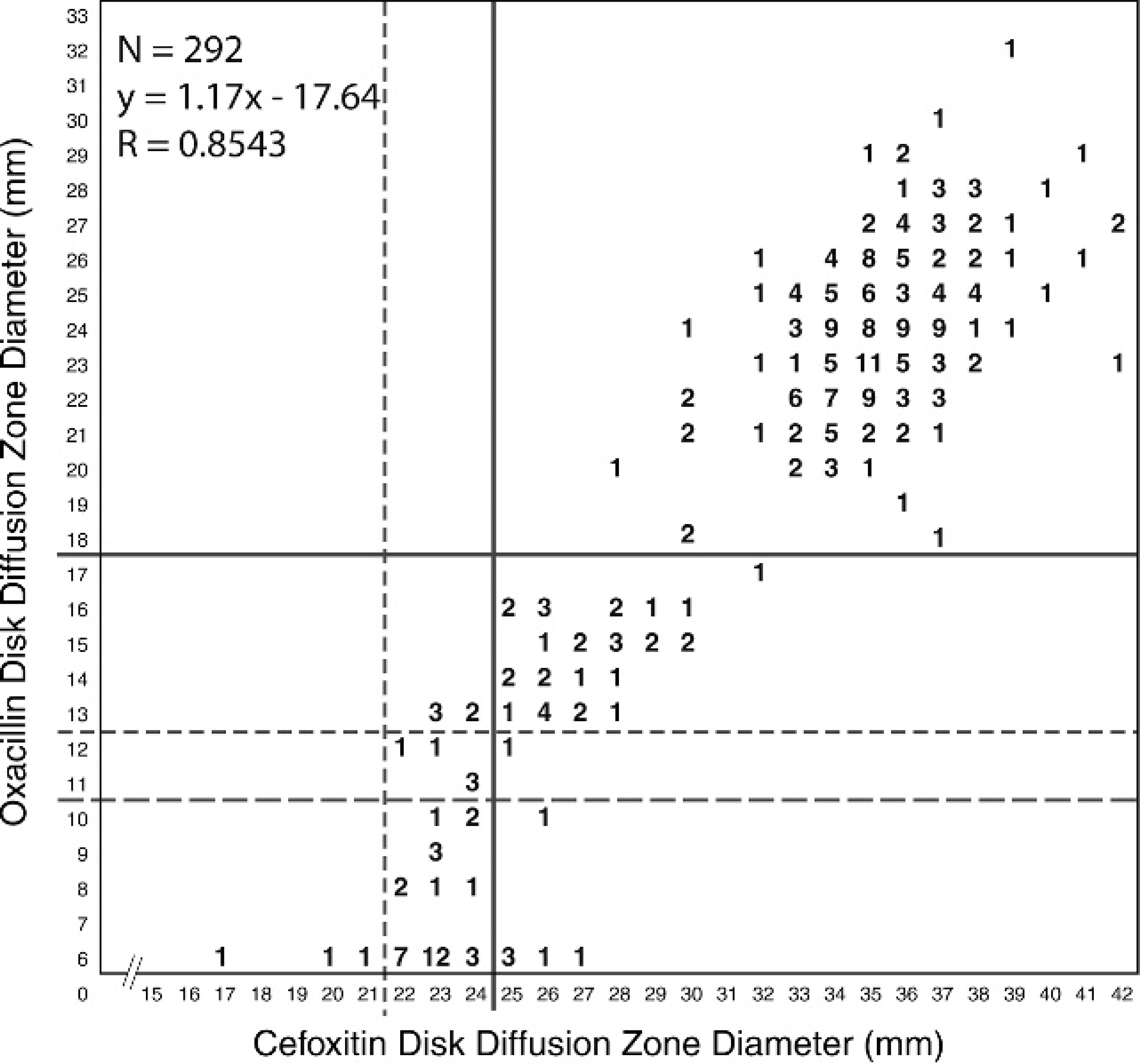
Correlation between oxacillin disk diffusion zone diameters and cefoxitin disk diffusion zone diameters. For oxacillin: solid lines = former (2004–2008) Clinical and Laboratory Standards Institute (CLSI) breakpoints; broken lines = current (2008) CLSI breakpoints. For cefoxitin: solid lines = CLSI breakpoints used for coagulase-negative staphylococci; broken lines = CLSI breakpoints used for Staphylococcus aureus.
The mecA PCR was performed on all S. pseudintermedius isolates from dogs during 2006 and on all resistant isolates through the first quarter of 2008. All available data relating mecA detection by PCR to zone diameters around disks for oxacillin and cefoxitin were analyzed by the error-rate bounding method 16 usually used to maximize intermethod agreement for MIC breakpoints established by CLSI. 7 Such calculations are intended to minimize false-susceptible (very major) errors for the disk diffusion to ≤1.5% and intermethod minor and total errors to ≥10.0% (Table 1). 16 Using current (2008) CLSI disk diffusion breakpoints, the rates of very major and total errors for oxacillin were 14.9% and 20.8%, respectively. Use of CLSI breakpoints for oxacillin disk diffusion that were in effect from 2004 to 2008 resulted in 0.9% very major errors and 1.5% total errors. A group of 116 archived isolates, with previously determined oxacillin disk diffusion zone diameters of 11–17 mm, was selected for further comparisons among tests to predict mecA-mediated resistance. Oxacillin disk diffusion and gradient diffusion test results showed high (>93%) categoric agreement with those of both the reference microbroth dilution and the “gold standard” mecA PCR, when the MIC resistant breakpoint was ≥0.5 μg/ml and the disk diffusion resistant breakpoint was ≤17 mm (Table 2). Categoric agreement between cefoxitin disk diffusion test results and mecA detection in these isolates was 34% when the resistant breakpoint was ≤24 mm. Greater than 90% of the oxacillin-resistant S. pseudintermedius isolates from dogs were also resistant to representative drugs from 4 non–beta-lactam drug classes (Table 3).
. Disk diffusion breakpoints (current and former) for oxacillin and cefoxitin with summary of intermethod error rates.*
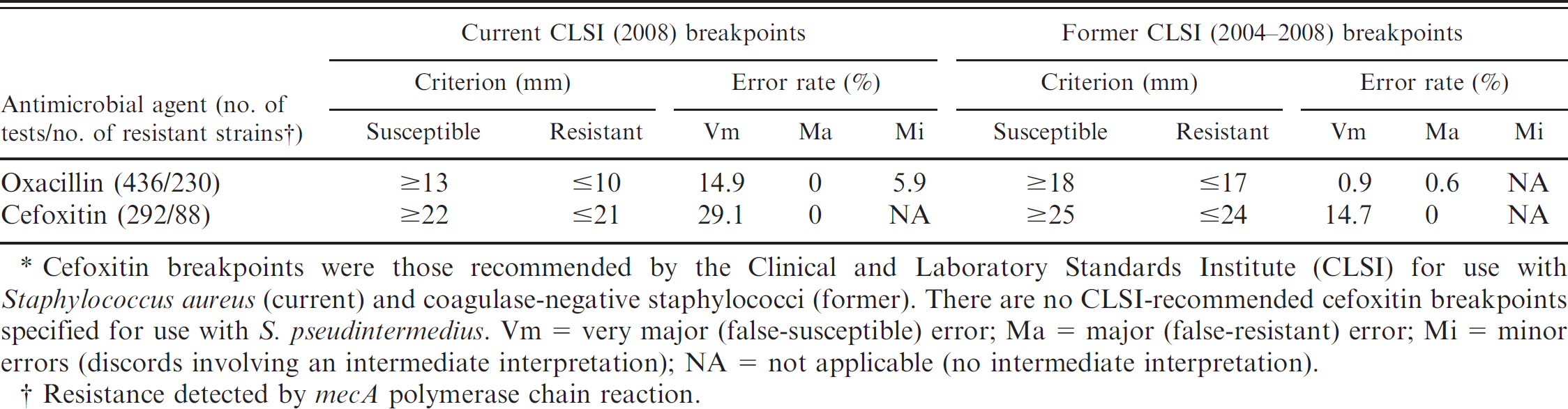
Cefoxitin breakpoints were those recommended by the Clinical and Laboratory Standards Institute (CLSI) for use with Staphylococcus aureus (current) and coagulase-negative staphylococci (former). There are no CLSI-recommended cefoxitin breakpoints specified for use with S. pseudintermedius. Vm = very major (false-susceptible) error; Ma = major (false-resistant) error; Mi = minor errors (discords involving an intermediate interpretation); NA = not applicable (no intermediate interpretation).
Resistance detected by mecA polymerase chain reaction.
The identities of more than 50 representative isolates were confirmed as S. pseudintermedius throughout the current study by partial 16S rRNA gene sequencing. Complete 16S rRNA and mecA gene sequences were determined for 4 isolates (2 from 2005 and 2 from 2006). The 16S rRNA gene sequences were identical in each of the 4 strains and were identical to those reported for S. pseudintermedius (GenBank accession nos. AJ780976, AJ780977, AJ780978, AJ780979). The mecA gene sequences were also identical in each strain tested and identical to those reported for S. haemolyticus (GenBank accession no. AP006716); they differed from those reported for S. pseudintermedius (GenBank accession nos. AM904732, AM904731) by 1 and 2 nucleotide bases, respectively.
. Percent categoric agreement among tests to predict mecA-mediated resistance in Staphylococcus pseudintermedius isolates with oxacillin disk diffusion zone diameters from 11 mm to 17 mm (N = 116).*
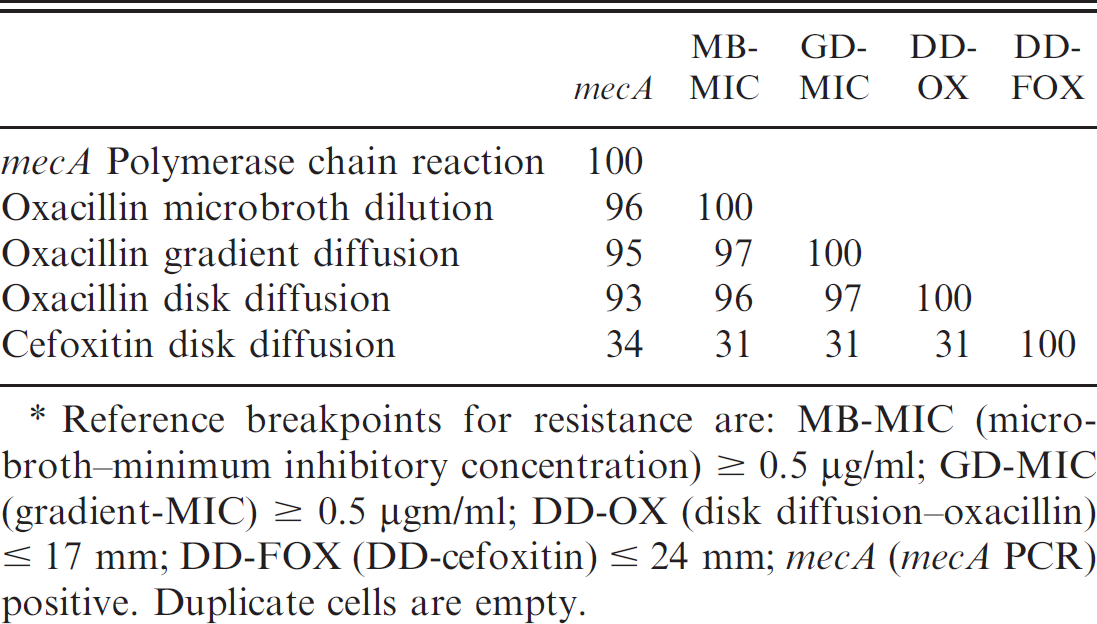
Reference breakpoints for resistance are: MB-MIC (microbroth-minimum inhibitory concentration) ≥ 0.5 μg/ml; GD-MIC (gradient-MIC) ≥ 0.5 μgm/ml; DD-OX (disk diffusion–oxacillin) ≤ 17 mm; DD-FOX (DD-cefoxitin) ≤ 24 mm; mecA (mecA PCR) positive. Duplicate cells are empty.
Discussion
The present study demonstrated that currently recommended CLSI interpretive criteria for oxacillin disk diffusion tests are not sufficient to detect all mecA-associated resistance in S. pseudintermedius. Detection of the mecA gene by PCR or gene probe or detection of its gene product by immunologic methods are considered to be the gold standards for confirmation of mecA-mediated resistance in staphylococci. 7 Disk diffusion tests provide simple and cost-effective means for laboratories to screen for oxacillin-resistant staphylococci. Discrepancy rate calculations used in the current study demonstrated that the use of current (2008) CLSI breakpoints resulted in unacceptable levels of very major and total errors in oxacillin disk diffusion tests. The present study also demonstrated that the results of oxacillin disk diffusion tests using former (2004–2008) CLSI breakpoints are in close agreement with those of gradient diffusion, microbroth dilution, and mecA gene detection by PCR. Oxacillin disk and MIC breakpoints of ≤17 mm and ≥0.5 μg/ml, respectively, were accurate indicators of mecA-mediated resistance in S. pseudintermedius. These former (2004) veterinary CLSI breakpoints 18 are identical to those currently recommended for human isolates of coagulasenegative staphylococci (CNS). 8 It is recommended that veterinary diagnostic laboratories use these former oxacillin interpretive criteria to improve detection of mecA-mediated resistance in clinical isolates of S. pseudintermedius.
. Multidrug resistance in Staphylococcus pseudintermedius isolates from dogs (2006).
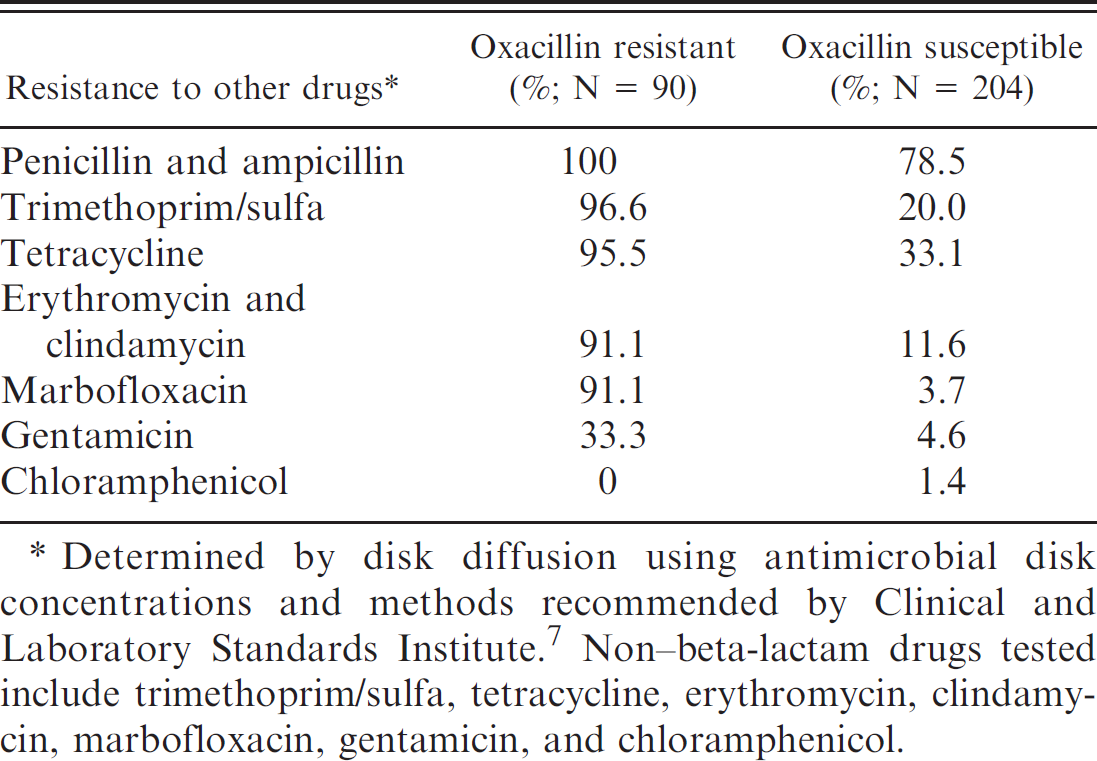
Determined by disk diffusion using antimicrobial disk concentrations and methods recommended by Clinical and Laboratory Standards Institute. 7 Non-beta-lactam drugs tested include trimethoprim/sulfa, tetracycline, erythromycin, clindamycin, marbofloxacin, gentamicin, and chloramphenicol.
As noted in the current and previous studies. 4,15 cefoxitin disk testing using current established CLSI criteria for both S. aureus and CNS underestimate mecA-mediated resistance in S. pseudintermedius. Currently, there are no recommendations for use of cefoxitin disk testing to predict mecA-mediated resistance in S. pseudintermedius. However, the current CLSI document applicable for veterinary isolates 7 has adopted recommendations from the counterpart document for human isolates 17 for cefoxitin disk testing with S. aureus and CNS. Veterinary diagnosticians should be cautioned that extrapolated use of cefoxitin disk testing with S. pseudintermedius may result in very major susceptibility testing errors.
The mecA gene sequences found in S. pseudintermedius are essentially identical to those found in methicillin-resistant S. aureus, 14 and penicillin binding protein 2a has been demonstrated by immunologic methods in methicillin-resistant S. pseudintermedius (MRSP). 4,11 It is presumed that expression of mecA in S. pseudintermedius has the same potential to confer broad clinical resistance to all beta-lactam drugs (penicillins, cephalosporins, carbapenems) as it does in S. aureus and other Staphylococcus species; however, this requires further study. The extent and level of resistance now being observed in oxacillin-resistant S. pseudintermedius isolates against representatives of other drug classes is of concern. What, if any, role the genomic island that contains mecA (staphylococcal cassette chromosome mec ) plays in multiple resistance of S. pseudintermedius is unknown. Investigations of resistance mechanisms and their distribution among staphylococcal populations have been conducted, but further work is needed to fully evaluate their clinical relevance. 3,9 The current study and others also confirm that relative resistance of S. pseudintermedius to oxacillin has increased in recent years. 10,13,15,19,20 Efforts to standardize detection methods, establish shared databases, promote collaborative research, and create venues for sharing knowledge of MRSP will help improve surveillance and reduce the spread of MRSP.
Acknowledgements
The authors thank Mary Jean Bryant, Brian Johnson, Rupal Brahmbhatt, Laura Eberline, Chad Black, Michael Ofori, and Christine Campion for technical assistance; Barry Gaddes for equipment and technical support; and Deborah Haines for graphic illustrations. Supported in part by UTCVM Center of Excellence grant and summer UTCVM student research fellowship program.
Footnotes
a.
BBL™ Crystal™ 2 Gram-Positive ID Kit, BD, Sparks, MD.
b.
Etest®, AB Biodisk, Solna, Sweden.
c.
BBL™, BD, Sparks, MD.
d.
Sensititre®, Companion Animal Panel COMEQ3F; TREK Diagnostic Systems Inc., Cleveland, OH.
e.
Remel, Lenexa, KS.
f.
Sensititer Vizion System, TREK Diagnostic Systems Inc., Cleveland, OH
g.
ATCC strains 25923, 43300, and 25922; American Type Culture Collection, Rockville, MD.
h.
BioSpec Products Inc., Bartlesville, OK.
i.
Premix Taq™, Takara Bio Inc., Otsu, Shiga, Japan.
j.
ExoSAP-IT®, USB Corp., Cleveland, OH.
k.
Lasergene® version 7.2, DNASTAR Inc., Madison, WI.
l.
Microsoft® Excel® 2007, Microsoft Corp., Redmond, WA.
