Abstract
The objective of the current study was to determine the sensitivity and specificity of real-time polymerase chain reaction (real-time PCR) for feline hemoplasmas when applied to DNA extracted from dried whole-blood smears in comparison to that for DNA extracted from liquid whole blood. Blood samples were collected into ethylenediamine tetra-acetic acid tubes from 305 cats with possible or suspected hemoplasmosis, and dried blood smears from each sample were prepared. DNA was extracted from blood smears and a 160-μl aliquot of each liquid blood sample by using a robotic extractor and was subjected to real-time PCR for feline glyceraldehyde-3-phosphate dehydrogenase (liquid blood), 18S ribosomal RNA (rRNA; dried blood), and “Candidatus Mycoplasma haemominutum”, Mycoplasma haemofelis, and “Candidatus Mycoplasma turicensis” DNA. When using the results for liquid whole blood as the gold standard, the sensitivity of each assay for “Ca. M. haemominutum”, M. haemofelis, and “Ca. M. turicensis” was 49 of 66 (74%), 11 of 13 (85%), and 11 of 20 (55%), respectively. The specificity of each assay was 224 of 234 (96%), 287 of 287 (100%), and 280 of 280 (100%), respectively. When possible, liquid blood samples should be submitted for detection of feline hemoplasmas by using real-time PCR. The improved sensitivity of real-time PCR on blood smears for M. haemofelis compared with that of the other hemoplasma species may reflect the higher organism burdens associated with infection with this species.
Hemotropic mycoplasmas (hemoplasmas) are epierythrocytic mycoplasmas that have never been cultured in vitro. At least 3 species infect cats: Mycoplasma haemofelis and “Candidatus Mycoplasma haemominutum”, as well as a “Candidatus Mycoplasma turicensis”, a new species that was first described in cats in Switzerland and subsequently detected in cats worldwide. 3,7,9,15,16 Mycoplasma haemofelis is capable of causing severe anemia in cats, whereas “Ca. M. haemominutum” has yet to be clearly associated with disease in immunocompetent cats. 2,4,8 Although uncommon, coinfections with multiple hemoplasma species were reported. 4,7,9,14
Cytologic examination of blood smears has low sensitivity and specificity for a diagnosis of hemoplasmosis, and “Ca. M. turicensis” has never been cytologically identified. 9,10,12,14,15 The advent of polymerase chain reaction (PCR) technology has improved the ability to detect hemoplasmas. Recently, quantitative real-time PCR assays were used for specific detection and quantification of “Ca. M. haemominutum”, “Ca. M. turicensis”, and M. haemofelis in whole-blood samples. 1,5,7–9,11,15 The analytical sensitivity of these assays has been as low as between 1 and 10 copies of organism DNA.
Organism loads in pet domestic cats testing positive when using real-time PCR were reported to range from approximately 20 to several million copies per ml of blood, 7,8,14,16 with the vast majority of cats having organism loads exceeding 1,000/ml. Cats infected with “Ca. M. turicensis” appear to have lower median organism loads than cats infected with other hemoplasma species. 14,16 Given that most blood smears are made with at least 10 μl of blood (one drop = 50 μl), it was hypothesized that the volume of blood in a dried blood smear would be sufficient for the detection of hemoplasmas when using real-time PCR. On occasion, blood smears may represent a desirable sample for hemoplasma PCR because of the ease of transport or the lack of availability of larger sample volumes. Often blood smears are stored at room temperature and are available for retrospective analysis when larger blood samples have been discarded. Retrospective analysis may be desirable in cats that have been treated empirically with antimicrobial therapy that would be expected to lead to negative PCR results. The objective of the current study, therefore, was to evaluate the sensitivity of real-time TaqMan PCR assays for “Ca. M. haemominutum”, M. haemofelis, and “Ca. M. turicensis” when used on dried blood smears from cats naturally infected with hemoplasmas compared with larger blood samples.
A total of 305 blood samples that had been submitted to the IDEXX Laboratory (Sacramento, CA) between January 2006 and June 2006 were used in the study. These samples were collected as part of an epidemiological study of cats with possible hemoplasmosis. 9 All blood samples had been anticoagulated with ethylenediamine tetra-acetic acid (EDTA). Two blood smears were made within 6 hr of sample collection and were air-dried. One smear was stained with Wright-Giemsa stain and evaluated cytologically for hemoplasma organisms. Liquid EDTA blood and dried-blood smears were stored at 4°C until processed; all samples were processed within 2 weeks of sample collection. For the dried blood smears, blood was scraped from unstained slides with a disposable cell scraper into 160 μl of phosphate buffered saline solution (PBS). Separate scrapers were used for each slide. DNA was extracted from this solution, along with a 160-μl aliquot of the matched liquid whole-blood sample by using a robotic extractor, a with a final elution volume of 100 μl.
Real-time PCR assays for detection of the 16S ribosomal RNA (rRNA) gene of “Ca. M. haemominutum”, 8 M. haemofelis, 8 and “Ca. M. turicensis” 15 were performed as described. Real-time PCR assays for feline glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and eukaryotic 18S rRNA were used as controls in liquid whole-blood samples and dried-blood samples, respectively, to indicate the successful purification of DNA and the absence of PCR inhibitors, as described. 8 , b Amplification was performed with the default cycling parameters of the manufacturer's real-time PCR assay platform b and consisted of 2 min at 50°C, 10 min at 95°C, and 40 cycles of 15 sec at 95°C and 60 sec at 60°C. Data were collected and processed with sequence detection system software. b Positive controls for each species and negative extraction controls (160 μl PBS) were included with each PCR run. Cycle threshold values of < 40 were considered positive.
To compare the limit of organism detection from liquid whole blood to that from dried-blood smears, dried-blood smears were also made from 10-fold serial dilutions of whole blood derived from 3 blood samples. One sample had tested positive when using real-time PCR for “Ca. M. haemominutum” alone, one had tested positive for M. haemofelis alone, and the remaining had tested positive for “Ca. M. turicensis” alone. The results of real-time PCR when applied to serial dilutions of these liquid whole-blood samples were compared with the results of real-time PCR when applied to the dried-blood smears.
The D'Agostino and Pearson normality test was used to confirm normality among data from clinical cases. For the samples that tested positive on liquid whole blood, the unpaired t-test was used to compare cycle threshold (Ct) values for each pathogen for liquid whole-blood samples between samples that tested positive on dried whole-blood smears and samples that tested negative on dried whole-blood smears. Chi-square analysis was used to compare the assay sensitivities for each species. All analyses were performed with standard statistical software c ; P values of < 0.05 were considered significant.
Five of the 305 dried whole-blood samples analyzed tested negative for 18S DNA and were negative for all 3 hemoplasma species. These were excluded from the study because of the possibility that inhibitors present in these samples prevented amplification of hemoplasma DNA. Nevertheless, the PCR assays for feline GAPDH in the corresponding liquid whole-blood samples from the 5 cats were positive.
Results of the real-time PCR assay for “Ca. M. haemominutum” DNA on the liquid whole-blood samples were positive for 66 of the 300 cats (22%). On the dried whole-blood samples, results of the real-time PCR assay for “Ca. M. haemominutum” DNA were positive for 49 of these 66 cats, and for 10 of the 234 negative cats, which yielded a sensitivity of 74% and a specificity of 96% when compared with real-time PCR on liquid whole blood. The Ct values for the liquid whole-blood samples were lower in the samples that also tested positive when using dried whole blood than the Ct values for the liquid whole-blood samples that tested negative when using dried whole blood (Fig. 1A). The Ct values in the samples that tested positive on dried whole blood but not on liquid whole blood ranged from 29.0 to 39.9 (median: 39.1).
Results of the real-time PCR assay for M. haemofelis DNA on the liquid whole-blood samples were positive for 13 of the 300 cats (4.3%). On the dried whole-blood samples, results of the real-time PCR assay for M. haemofelis DNA were positive for 11 of these 13 cats and for none of the 287 negative cats, that yielded a sensitivity of 85% and a specificity of 100% when compared with real-time PCR on liquid whole blood. The Ct values on liquid whole blood for the cats testing positive by using both sample types were 27.2 ± 3.4 mean (SD). The 2 samples that were negative on dried-blood smears had Ct values of 30.2 and 36.6 (Fig. 1
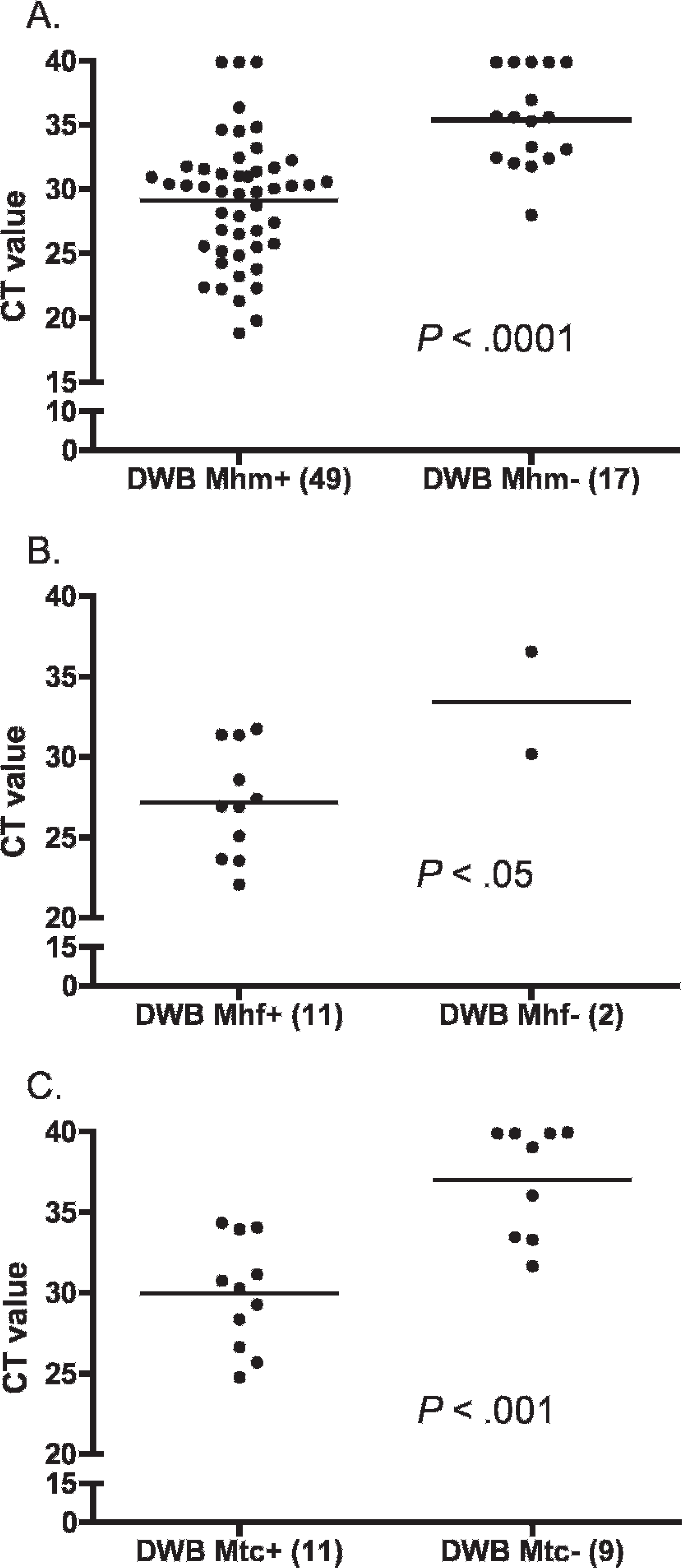
Comparison of real-time polymerase chain reaction (real-time PCR) cycle threshold values on liquid whole blood for cats that tested negative for hemoplasmas when using real-time PCR on dried whole-blood smears (right) and cats that tested positive when using real-time PCR on dried whole-blood smears (left).
Results of the real-time PCR assay for “Ca. M. turicensis” DNA on the liquid whole-blood samples were positive for 20 cats (6.6%). On the dried whole-blood samples, results of the real-time PCR assay for “Ca. M. turicensis” DNA were positive for 11 of these 20 cats and for none of the 280 negative cats, which yielded a sensitivity of 55% and a specificity of 100% when compared with real-time PCR on liquid whole blood. The Ct values for the liquid whole-blood samples were lower in the samples that also tested positive when using dried whole blood than the Ct values for the liquid whole-blood samples that tested negative when using dried whole blood (Fig. 1C).
Hemoplasmas were identified during cytologic examination of blood smears from 8 cats. Based on the results of conventional PCR and sequencing, 2 of these cats were infected with a variant strain of “Ca. M. haemominutum” not expected to be detected when using the real-time PCR assay for “Ca. M. haemominutum”. 9 Real-time PCR on DNA extracted from the dried blood smears was positive for M. haemofelis in 5 samples, including 1 sample that was also positive for “Ca. M. turicensis”. For the remaining cat, the result was negative.
When using real-time PCR on the liquid whole-blood samples, mixed infections were detected in 18 of the 300 cats, including 4 cats infected with “Ca. M. haemominutum” and M. haemofelis, 9 cats infected with “Ca. M. haemominutum” and “Ca. M. turicensis”, 3 cats infected with M. haemofelis and “Ca. M. turicensis”, and 2 cats infected with all 3 species. When dried whole-blood smears were analyzed, mixed infections were detected in 11 of the 300 cats, including 2 cats infected with “Ca. M. haemominutum” and M. haemofelis, 6 cats infected with “Ca. M. haemominutum” and “Ca. M. turicensis”, 1 cat infected with M. haemofelis and “Ca. M. turicensis”, and 2 cats infected with all 3 species. No statistical difference was found among the sensitivities of real-time PCR on dried whole-blood smears as opposed to liquid whole blood for each of the hemoplasma species.
When real-time PCR was applied to serial dilutions of 3 known positive liquid whole blood and corresponding dried-blood smears, for each of the 3 species, hemoplasmas could no longer be detected in dried-blood smears at dilutions of whole blood higher than 10-1. In contrast, hemoplasmas could be detected in dilutions of liquid whole blood of 10-3 for the sample positive for M. haemofelis, 10-3 for the sample positive for “Ca. M. haemominutum”, and 10∼4 for the sample positive for Ca. M. turicensis”. Because the initial DNA load varied between the 3 samples, and the volume of blood used to make each blood smear was not consistent (as would be the case in a clinical situation), it was not possible to compare the results for each of the 3 hemoplasma species.
The current study showed that the diagnostic sensitivity of real-time PCR for feline hemoplasmas was lower when applied to dried whole-blood smears compared with that applied to liquid whole blood. The lower volume of blood from which DNA was extracted is most likely the major reason for the reduction of sensitivity of the assays. The degree of reduction of sensitivity appeared to be dependent on the hemoplasma species present, with the loss of sensitivity being greatest for detection of “Ca. M. turicensis” and least for detection of M. haemofelis, although the numbers of positive samples were not sufficient to demonstrate statistical significance when using a 2-sided chi-square analysis. This likely reflects the higher organism burdens that have been often associated with infection with M. haemofelis. 7,14,16 In contrast, “Ca. M. turicensis” has been associated with very low organism burdens and consistent failure to detect the organism when using cytologic evaluation of stained blood smears. Given that M. haemofelis is most likely to be associated with hemolytic disease, 2,9 the low sensitivities noted for detection of “Ca. M. haemominutum” and “Ca. M. turicensis” may not necessarily be detrimental for the purpose of diagnosis of clinical hemoplasmosis.
Eukaryotic 18S rRNA was used as the housekeeping gene target for the dried-blood samples in the current study, because the GAPDH assay used for liquid whole blood generated high Ct values with low blood volumes analyzed. In contrast, stronger signals resulted from amplification of the 18S rRNA gene, perhaps reflecting a more abundant copy number of this gene (data not shown).
In the present study, application of real-time PCR to dried-blood smears was also useful to identify the infecting species when organisms were visible after cytologic evaluation. Negative results when organisms were visible occurred in 3 of the 8 samples; in 2 of these 3 samples, real-time PCR was predicted to yield negative results because of the presence of a variant strain of “Ca. M. haemominutum” not expected to be detected when using the real-time PCR assay for “Ca. M. haemominutum”. 8,9 Polymerase chain reaction assays on liquid whole blood have been shown to be much more sensitive for the diagnosis of hemoplasmosis than cytology, 10,13 and this was also true for dried whole-blood smears. As expected, the number of mixed infections detected was reduced when dried-blood smears were analyzed as opposed to liquid whole blood. This was often because of failure to detect “Ca. M. turicensis”, which occurred in 5 of the 7 samples in which mixed infections were detected when using liquid whole-blood samples.
The loss of sensitivity for detection of each pathogen when using dried whole-blood smears was related to the Ct value for liquid whole-blood samples, which, in turn, is a crude measure of the number of organisms present in the sample. It was not always possible to predict whether a dried blood smear would yield negative results when using real-time PCR based on the Ct value for the liquid whole-blood sample. The authors chose to present Ct values in the current study rather than hemoplasma copy numbers, because variation in DNA extraction efficiency can result in variation in the calculated load. Normalization to an internal control has been used to overcome this problem, but when performed on whole blood, feline GAPDH copy numbers are dependent on nucleated cell numbers, which may vary from cat to cat.
Positive results for “Ca. M. haemominutum” were noted in 10 dried whole-blood smears when the results of real-time PCR on the corresponding liquid whole-blood samples were negative. The reasons for these results were unclear. Although possible, aerosol contamination during PCR seemed unlikely given consistently negative results for negative-control samples, the use of a robotic extractor, and inclusion of uracil N-glycosylase to destroy amplicons before initiating PCR. Contaminations during blood-smear preparation or transport, slide scraping, or sample loading errors are other possible explanations for discrepant results. The majority of these samples had high Ct values (>35). The other possibility is that these results represented true positives that were undetected when using real-time PCR on liquid whole blood. Positive PCR results for feline GAPDH in the liquid whole-blood samples made false negatives in these samples because of PCR inhibitors an unlikely explanation. Pipetting errors during addition of template to the PCR master mix could conceivably have led to false negatives, but the results of conventional PCR, which was performed in a previous study on these samples, were also negative. 8
In conclusion, application of real-time PCR to dried whole-blood smears from cats can be used to detect hemoplasma infections, although sensitivity is reduced compared with the use of liquid whole blood. This may potentially be useful for retrospective analysis, when larger volumes of liquid whole blood are not available, or after commencement of empiric antimicrobial therapy, with assuming dried blood smears were prepared and stored before starting therapy. Although dried blood smears were stored at 4°C in the present study, results are likely to be similar for slides stored at room temperature because of the high degree of stability of DNA in air-dried blood at various conditions of relative humidity and temperature. 16
Acknowledgements. The authors would like to thank Nicole Drazenovich for technical assistance and Dr. Christian Leutenegger for assistance with real-time assays. This project was funded, in part, by a generous donation from Tom and Penny McCormack, and a grant from the Students in Advanced Research (STAR) program. The project was supported in part by IDEXX Laboratories.
Footnotes
a.
Corbett Robotics Pty Ltd, Sydney, New South Wales, Australia.
b.
Applied Biosystems Inc., Foster City, CA.
c.
GraphPad Prism version 4.00, GraphPad Software Inc., San Diego, CA.
