Abstract
Feline retroviruses are rarely reported in lynx species. Twenty-one Iberian lynx (Lynx pardinus) blood and tissue samples collected from Doñana National Park and Los Villares (Sierra Morena) in southern Spain during 1993–2003 were analyzed by polymerase chain reaction to amplify nucleic acids from feline retroviruses. Six samples were positive for Feline leukemia virus (FeLV), but no samples tested positive for Feline immunodeficiency virus. The BLAST analysis indicated that 5 of the 6 sequences were closely related to FeLV strain Rickard subgroup A, whereas 1 sequence was identical to FeLV. To the authors' knowledge, this is the first report of FeLV in the endangered Iberian lynx.
Keywords
Retroviral diseases are debilitating processes that can affect felines. In its natural host, the domestic cat, the infection by Feline leukemia virus (FeLV; family Retroviridae, subfamily Orthoretrovirinae, genus Gammaretro-virus) usually occurs through the oronasal route. After infection, some cats eliminate the virus and produce neutralizing antibodies, whereas others remain latently infected or become persistently infected and eventually succumb to neoplastic or nonneoplastic diseases induced by the virus. 6 Illness in affected cats usually results from either the direct effects of the virus on the bone marrow (bone marrow suppression, myeloproliferative disease) or lymphoid tissue (lymphoma) or, more often, the effects of viral immunosuppression, resulting in life-threatening secondary infections. The virus is shed mostly through saliva and is very labile in the environment. 6
Feline immunodeficiency virus (FIV; family Retroviridae, subfamily Orthoretrovirinae, genus Lentivirus) is very similar to Human immunodeficiency virus 1 (HIV-1). It is shed mainly in saliva and is transmitted mostly through bite wounds, but it is also transmitted in utero or through colostrum and milk. Feline immunodeficiency virus induces a progressive immune dysfunction. The clinical course of immunodeficiency associated with FIV, which develops for months or even years, is divided into the same 5 stages as acquired immunodeficiency syndrome (AIDS). Animals progress from a period of nonspecific signs to a long asymptomatic phase when CD4+ cells decrease to a point in which the cat acquires secondary and opportunistic infections that lead to its death. 17
Spain is the only country in Europe in which colonies of Iberian lynx (Lynx pardinus) still remain. 5 However, the population is declining rapidly, and the species is considered critically endangered (International Union for Conservation of Nature and Natural Resources: 2007, IUCN Red List of Threatened SpeciesTM. Available at: www.iucnredlist.org. Accessed on November 13, 2007). It is limited to 2 isolated and very small metapopulations in Doñana National Park and Sierra Morena (southern Spain), with a total estimated population of no more than 160 individuals. 5 This population is considered highly endogamic, which could decrease the genomic potential of the lynx. In addition, the lynx are relatively close to human dwellings and, thus, are endangered by proximity to man, domestic and feral cats (Felis catus), dogs (Canis familiaris), and European wildcats (Felis silvestris), which might even share territories with the lynx. These species could transmit infections to the lynx, as has been described for other wild felids that live close to domestic cats. 7 Although intensive biological research has been carried out on this species during the last 20 years, peer-reviewed reports on diseases that affect wild Iberian lynx are very limited. 2,8,11
The presence of FIV-related lentiviruses has been described in a large variety of wild felids throughout the world. 19 In contrast, few reports indicate that nondomestic felids, both free-ranging and in zoological settings or privately owned, are susceptible to FeLV infection. 16,18 In many of the cases, domestic and feral cats were suspected to be the FeLV source. 7 In general, FeLV epizootics in wild felids are less likely than in zoological clusters because of felid's solitary and territorial behavior, with the exception of African lions (Panthera leo). 14 However, a relatively high prevalence of FeLV antigenemia has been reported in several populations of wildcats from Europe and the Middle East (Luaces I, Lutz H, Barreiro B, et al.: 2002, Feline leukemia virus, immunodeficiency virus, coronavirus and panleukopenia virus seroprevalence in free-living and captive wildcats (Felis silvestris) from Spain, In: Proc European Association of Zoo and Wildlife Veterinarians and European Wildlife Disease Association combined meeting, ed. Erken AHM, Dorrestein GM, pp. 223–225. Heidelberg, Germany), 3,4,10 where it could increase mortality in this species. 4
The aims of the current study were, first, to perform a retrospective study of retroviruses on Iberian lynx samples that originated from the Wildlife Rehabilitation Centers Acebuche (Doñana National Park, southern Spain) and Los Villares (Sierra Morena, southern Spain) and, second, to further characterize the viruses to correlate them with the origin of infection. To achieve these goals, 25 blood, serum, or tissue samples (liver and lymph nodes; Table 1), obtained aseptically from all dead and live Iberian lynx during health checks or postmortem examinations from 1993 to 2003, were analyzed. Samples were preserved for different investigations in Tris-sodium dodecyl sulfate–ethylenediamine tetra-acetic acid (Tris-SDS-EDTA) buffer specifically designed for DNA studies or were frozen at −70°C. The presence of FeLV p27 antigen and antibodies against FIV p24 was determined in serum or plasma samples with a commercial kit. a
Origin of the samples and polymerase chain reaction (PCR) and serology results from lynx included in the study.*
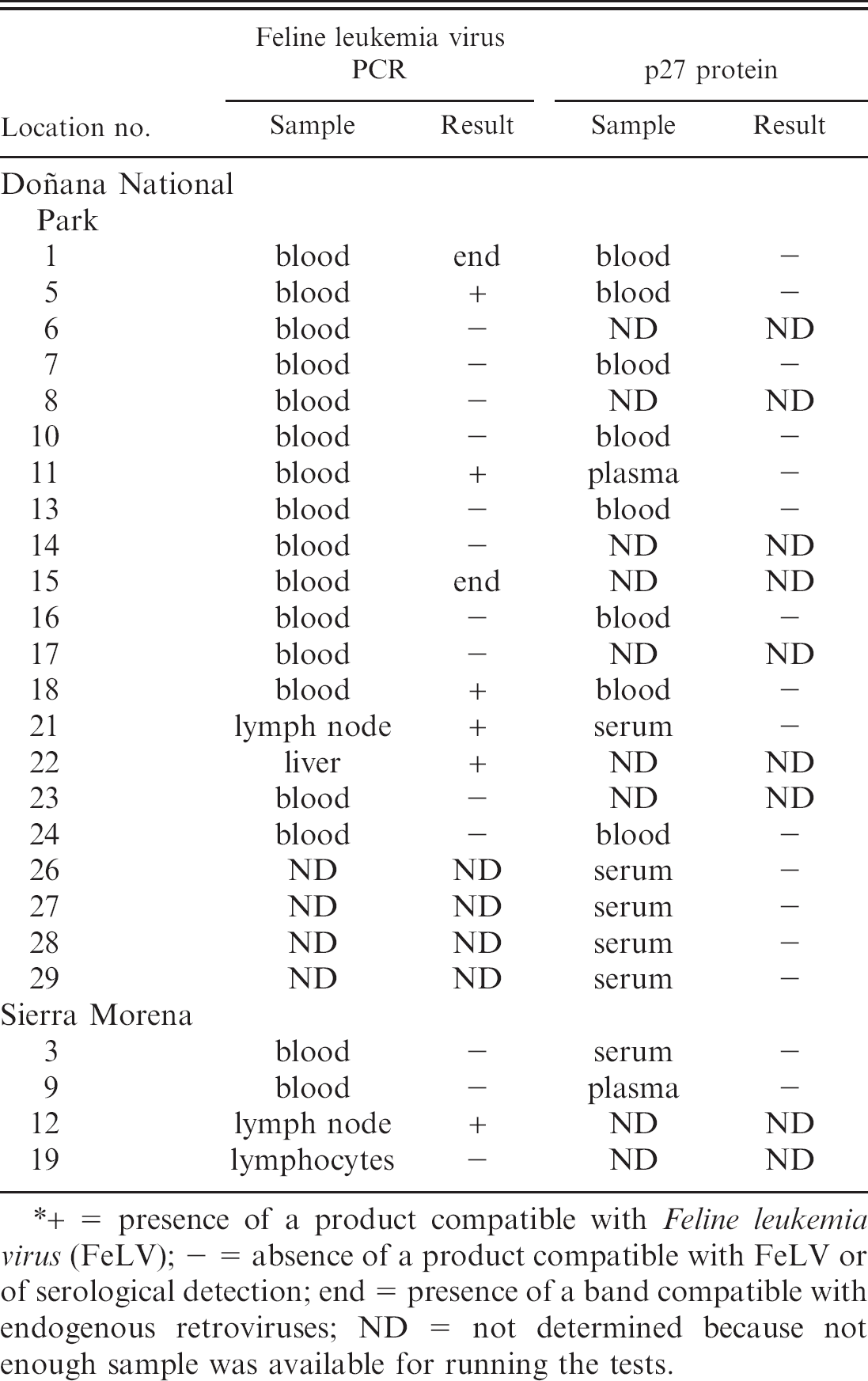
+ = presence of a product compatible with Feline leukemia virus (FeLV); - = absence of a product compatible with FeLV or of serological detection; end = presence of a band compatible with endogenous retroviruses; ND = not determined because not enough sample was available for running the tests.
DNA was extracted from lynx blood and tissue samples with the use of a commercial kit b according to the manufacturer's instructions. DNA extracted from 2 feline lymphocytic cell lines persistently infected with FeLV (FL74-UCD [ATCC CRL-8012]) or with FIV (Ho6) were used as positive controls, and DNA from the monocytic U-937 (ATCC CRL-1593) was a negative control. The virus was detected by a double-nested polymerase chain reaction (PCR) designed by the authors, 1 which amplifies a region in the pol gene of FeLV and FIV. As a control of the correct DNA extraction and quality, the gene for glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was also amplified with primers GAPDH-F and GAPDH-R. 9
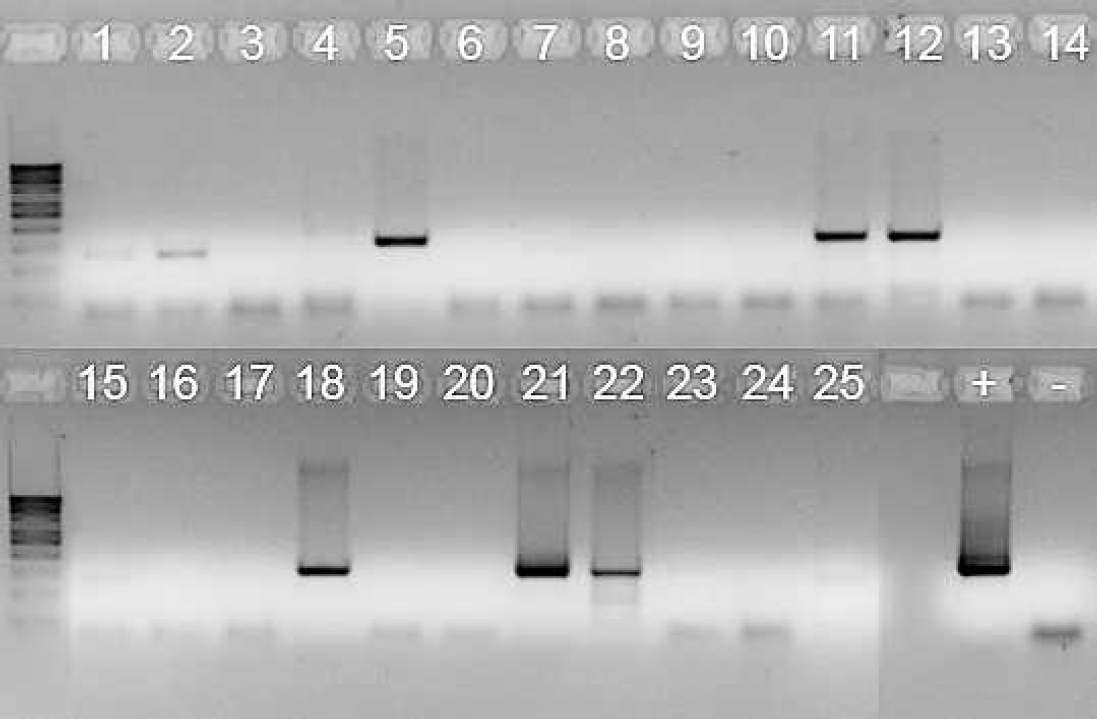
Result of the polymerase chain reaction. The lane numbers correspond to the numbers assigned to the lynx. The darker bands of about 300 base pairs (bp) were excised from the gel and sequenced. The lighter smaller band corresponds to endogenous sequences. DNA from FL74 was used as positive control (+) and from U937 as negative control (-). The DNA ladder shown is 100 bp.
The PCR products were purified with the use of a commercial gel extraction kit c and were sequenced by the DNA Sequencing Service at the Biological Research Center (CIB-CSIC, Madrid, Spain) with the sequencer 3730 DNA Analyzer. d Sequences were analyzed with the BLAST 2.0.10. algorithm (http://www.ncbi.nlm.nih.gov/BLAST/Blast.cgi) and were compared with each other and with different sequences by ClustalW (http://www.ebi.ac.uk/clustalw/). Products sequenced included the positive bands obtained in the lynx samples and those that corresponded to the positive FeLV control (FL74-UCD). The lynx sequences have been deposited in GenBank with the following accession numbers: EF418004, EF418005, EF418006, EF418007, EF418008, and EF418009.
Serological tests were not run on 9 of the samples, either because only tissue was available or because the amount of blood received was not enough for both serology and PCR. Similarly, PCR was not done in 4 lynx from which only serum was received. The FeLV p27 antigen and FIV p24 antibody tests were negative for all 16 lynx samples tested. However, a product compatible with FeLV was observed in 6 of the 21 lynx DNA samples (3 of 18 blood samples, 2 of 2 lymph node samples, and 1 of 1 liver sample) analyzed by nested PCR (Fig. 1; Table 1). A band corresponding to FIV was not detected in any of the animals. The design of this PCR technique allows the detection of endogenous retroviral sequences, distinguishable from FeLV, to which they are closely related because of their smaller size. In domestic cats, endogenous sequences should be present in all DNA samples, and its detection is considered an internal control of the reaction. 1 Bands compatible with endogenous sequences were observed in DNA from 2 of the nonpositive lynx, but the detection of a band that corresponded to GAPDH in all 21 samples (results not shown) confirmed the good quality of the DNA for the assay and the absence of PCR inhibitors.

Nucleotide sequences of the amplified bands. “FeLV1” sequence corresponds to GenBank accession AF052723; all others were sequenced in this study. Mutations are shown in bold letters and shaded.
The lack of correlation between serology and PCR results was published previously. 7,15 In this study, an explanation for this discrepancy could be that the viruses generated in lynx have no cross-reactions with those from domestic cats, and the reagents used in the commercial kits do not detect them. This raises a doubt on the usefulness of serological techniques developed for domestic cats to detect retrovirus in wild felids. The PCR designed by the authors was chosen to determine the infection by FIV, FeLV, or both because it is sensitive and specific 1 and is successful in amplifying DNA sequences from small blood samples, as was the case with the lynx. The target sequence is in the pol gene of the viral genome, a region that is considered highly conserved 1 and thus permits the detection of FeLV and FIV genomes in isolates otherwise divergent.
The BLAST analysis of the nucleotide sequences obtained from the PCR amplicons in this study indicated that amplified sequences had 98% homology with the FeLV strain Rickard subgroup A (GenBank accession AF052723). In the ClustalW comparison among the sequences from the lynx, the positive control and AF052723 showed variability within some of the nucleotides (Fig. 2; Table 2). As deduced from the pol open reading frame from AF052723, mutations in the nucleotides from the lynx sequences at positions 2901 and 2907 involved a change in the encoded amino acids, specifically at residue 263, where methionine is substituted by isoleucine, and in 265, where threonine is substituted by methionine, assuming that there are no deletions upstream of the DNA fragment amplified (Fig. 3). This could alter the functionality of the reverse transcriptase enzyme (which corresponds to the amplified region) and, thus, the pathogenicity of the virus.
Point mutations observed between the sequences from the lynx and Feline leukemia virus (FeLV) strain Rickard subgroup.*
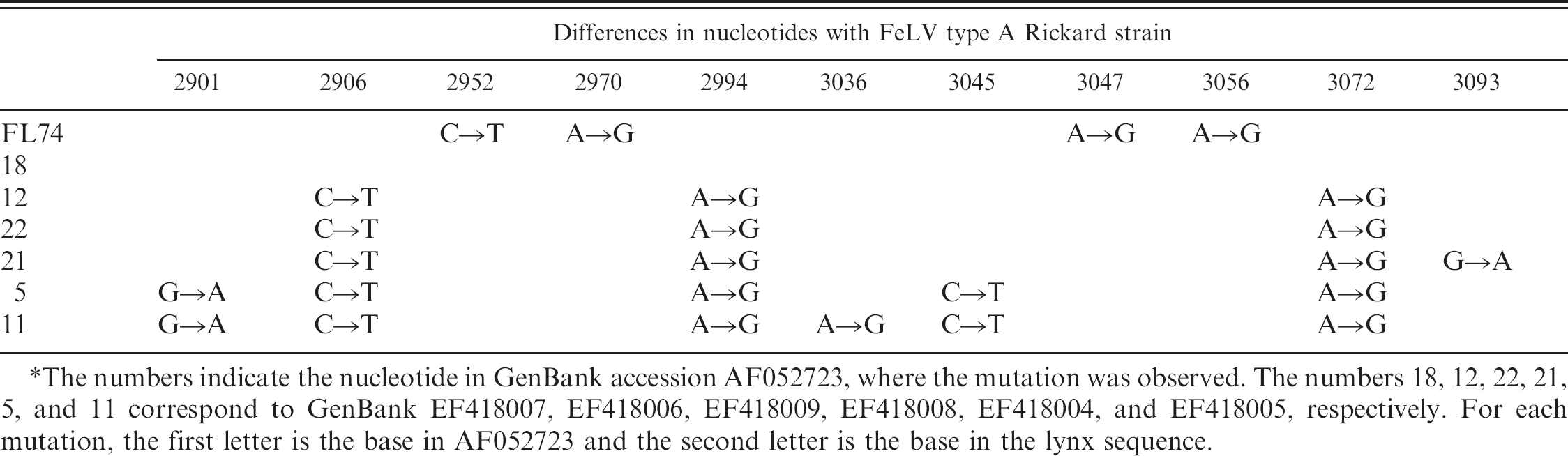

The numbers indicate the nucleotide in GenBank accession AF052723, where the mutation was observed. The numbers 18, 12, 22, 21, 5, and 11 correspond to GenBank EF418007, EF418006, EF418009, EF418008, EF418004, and EF418005, respectively. For each mutation, the first letter is the base in AF052723 and the second letter is the base in the lynx sequence.
To confirm that the PCR products obtained from these lynx were not a laboratory contamination, the control FeLV used in the laboratory was sequenced. As shown in Table 2, FL74 had mutations in nucleotides that were not altered in the lynx sequences when compared with AF052723; thus, it was considered that the animals had been infected in vivo. Some of the mutations were very common in the lynx samples.
Only 16.7% of the blood samples were positive by PCR, whereas a band corresponding to FeLV was detected in 100% of the tissue (lymph node or liver) samples. This finding could suggest that blood might not be the most adequate sample for PCR diagnosis. However, this cannot be confirmed because of the low number of samples and the absence of paired blood–tissue samples from the same lynx.

Derived amino acid sequences, assuming that no deletions or insertions were introduced in the DNA upstream of the sequenced fragment. The mutations are shown in bold letters and shaded. Conserved sequences are shown by a star and conservative variations by a colon; nonconservative variations have no associated symbol.
Published studies on wild lynx populations (including Iberian lynx, Eurasian lynx, and Bobcat) did not show the presence of antibodies against FeLV or FIV by different immunological techniques or FeLV p27 antigenemia. 12,13,15 In contrast, feline leukemia was diagnosed in captive Bobcat (Lynx rufus), 18 in which both FeLV antibodies and antigen were detected and were associated with clinical signs, which confirmed that these animals are susceptible to infection by FeLV.
Feline leukemia virus infection in the Iberian lynx could have come from a virus specific to the lynx populations or from infection from domestic or feral cats in the vicinities of the natural reserves. Although infrequent, viruses from domestic cats have been isolated from wild felids, which would mean that transmission is possible. The fact that Bobcats that acquired the infection had contact with domestic cats (a Bobcat with a nursing cat) 18 strengthens this hypothesis. In the case of the Iberian lynx, the rise in human population at the edge of its habitat increases the risk of contact with pathogens from both domestic and feral cats and other domestic animals. The high homology between the amplified sequences from lynx and from domestic cats suggests transmission from domestic cats.
It could happen that, after a period of contact between infected domestic or feral cats and lynx, the virus would cross the species barrier, and after adapting to the new host, it would be able to be transmitted between lynx. The solitary habits of the lynx greatly reduce the probability of intraspecies transmission in the wild. However, in some instances, close contact between lynx enhances the risk of contagion. These would include mating, stays in recovery centers, or mother–immature lynx relationships.
Certain mutations (i.e., in nucleotides 2906, 2994, and 3072) were prevalent in the lynx samples. This would support the possibility that the virus had been in the lynx population for quite some time. Other mutations besides the aforementioned 3 were also observed. Conceivably, the virus would acquire these mutations while it replicates in animals; thus, the more mutations, the longer the virus has been in the species. The DNA sequenced in one of the lynx samples (no. 18) was identical to AF052723, which could mean that it had acquired the infection directly from domestic cats.
To the authors' knowledge, this is the first report on the detection of FeLV provirus in Iberian lynx. Sequencing analyses suggest that FeLV isolated from Iberian lynx has a common origin with that of domestic cats and might have crossed the species barrier, reinforcing the risk that the Iberian lynx is exposed from its proximity to human dwellings.
Acknowledgements. The authors are indebted to David Bruhn for his editorial assistance. This work was supported with a grant of the Spanish Ministry of Science and Technology AGL2005–05248/GAN. V. M. Collado and G. Tejerizo are grantees of the Spanish Ministry of Science and Technology and of the Community of Madrid, respectively. The authors thank M. Horzinek (Utrecht, The Netherlands) for the generous gift of the 2 feline lymphocytic cell lines, and the Spanish Ministerio de Agricultura, Pesca y Alimentación, and the Junta de Andalucia for kindly providing the lynx samples.
Footnotes
a.
Snap Combo®, IDEXX Laboratories, Inc., Westbrook, ME.
b.
DNeasy Extraction Kit, Qiagen, Hilden, Germany.
c.
QIAquick® Gel Extraction Kit, Qiagen, Hilden, Germany.
d.
Applied Biosystems, Foster City, CA.
