Abstract
Jaagsiekte sheep retrovirus (JSRV) causes ovine pulmonary adenocarcinoma. JSRV can be transmitted via infected colostrum or milk, which contain somatic cells (SCs) harboring JSRV provirus. Nevertheless, the cell types involved in this form of transmission and the involvement of the mammary gland remain unknown. We separated adherent cells (macrophages and monocytes) by plastic adherence, and lymphocytes (CD4+ and CD8+ T cells, and B cells) by flow cytometry, from SCs in milk samples from 12 naturally infected, PCR blood test JSRV–positive, subclinical ewes. These cell populations were tested by PCR to detect JSRV provirus. The ewes were euthanized, and mammary gland samples were analyzed immunohistochemically to detect JSRV surface protein. We did not detect JSRV provirus in any milk lymphocyte population, but milk adherent cells were positive in 3 of 12 sheep, suggesting a potential major role of this population in the lactogenic transmission of JSRV. Immunohistochemistry did not reveal positive results in mammary epithelial cells, pointing to a lack of participation of the mammary gland in the biological cycle of JSRV and reducing the probability of excretion of free viral particles in colostrum or milk.
Jaagsiekte sheep retrovirus (JSRV; Retroviridae, Betaretrovirus) causes ovine pulmonary adenocarcinoma (OPA, jaagsiekte, sheep pulmonary adenomatosis), 15 a contagious lung neoplasia of sheep that occurs worldwide. 19 JSRV induces adenocarcinoma of type II pneumocytes and club cells of the lung; the clinical consequences are chronic respiratory disease, weight loss, and death. 5
JSRV has very high activity in the cells in which the tumor develops, but the provirus can also be detected in white blood cells, lymph nodes, and other lymphoid organs prior to the onset of OPA lesions.2,4,6,9,11,14 In addition, JSRV proviral DNA can be detected in somatic cells (SCs) from colostrum and milk (C/M) in sheep in OPA-affected flocks.3,10 It has also been reported that lambs artificially fed C/M from JSRV-infected ewes became blood-infected at a very early age, 10 and that JSRV can reach Peyer patches and mesenteric lymph nodes of newborn lambs nursed by infected mothers. 3 These data demonstrate that JSRV can be transmitted to lambs through the maternal route via infected C/M, but other details of this means of transmission for JSRV are not known. The lactogenic route is an important transmission route for other exogenous retroviruses infecting animals, such as visna-maedi virus (VMV) and caprine arthritis encephalitis virus, referred to as small ruminant lentiviruses present in both sheep and goats, and mouse mammary tumor virus (MMTV). These viruses replicate actively in mammary epithelial cells and are excreted in the C/M as cell-associated and/or free viral particles, infecting the newborn.1,16,17
To expand the understanding of JSRV transmission through the lactogenic route, we tested for JSRV proviral DNA in the main populations of mononuclear leukocytes isolated from milk obtained from naturally JSRV-infected ewes. In addition, we tested the mammary gland of infected ewes by immunohistochemistry (IHC) to detect cells containing JSRV surface protein in this tissue. The experimental protocol was approved by the Ethics Committee for Animal Experiments from the University of Zaragoza (reference PI 01/07).
We studied 12 Rasa Aragonesa ewes (5–7 y old), positive by JSRV-PCR blood test but subclinical. Samples of C/M from these ewes had been JSRV-PCR tested through a lactation period in a previous study. 3 Eight of these 12 ewes were PCR-positive in at least one of the C/M samples analyzed (group A in the current study); the remaining 4 ewes were PCR-negative in all samples (group B). A new gestation was induced in these 12 animals after estrus synchronization, and milk samples were taken in the subsequent lactation. We collected 5 milk samples (200 mL) from each ewe in both groups on days 4, 7, 10, 13, and 16 after parturition. SCs from these samples were obtained by several steps of centrifugation and washing with PBS (MilliporeSigma), as described previously. 8 The total number of cells was determined by counting microscopically, and an aliquot of 5 × 106 cells was stored at −80°C for PCR analysis. The remaining SCs were used for cell isolation procedures.
To obtain a milk leukocytic mononuclear cell-enriched fraction, 35–50 × 106 SCs resuspended in 1 mL of PBS were loaded on 5 mL of Lymphoprep (Axis-Shield) and centrifuged at 400 × g for 15 min at 20°C. Cells at the interface were washed twice and resuspended in PBS. After counting, these cells were utilized for the subsequent isolation procedures.
Plastic adherence was used to purify the adherent cell population (macrophages and monocytes), as described previously. 11 Briefly, an aliquot of 5 × 106 cells/mL was cultured in Roswell Park Memorial Institute (RPMI) medium (MilliporeSigma) supplemented with 10% fetal bovine serum (MilliporeSigma) and antibiotics (amphotericin B 1.25 µg/mL, streptomycin 100 µg/mL, penicillin 100 IU/mL; MilliporeSigma). Cultures were maintained overnight at 37°C in a humidified atmosphere of 5% CO2. Non-adherent cells were removed by washing with RPMI medium, and adherent cells were digested with lysis buffer 18 and stored at −80°C for PCR analysis.
Viable CD4+ T cells, CD8+ T cells, and B cells were isolated by flow cytometry using the following fluorescein isothiocyanate (FITC)-labeled mouse monoclonal antibodies: anti-ovine CD4 (MCA2213F) and CD8 (MCA2216F), and anti-bovine CD21 (MCA1424F), respectively (Bio-Rad). Fresh aliquots of cell suspensions (106 cells in 100 µL of PBS) were incubated with 10 µL of specific antibody for 15 min at 4°C in darkness. Cells were washed, resuspended in 500 µL of PBS with 2 mM EDTA, and nonviable cells were marked by incubation with 0.5 µL of Sytox Blue (Thermo Fisher) for 5 min at room temperature. Labeled cells were analyzed (FACSAria flow cytometer; BD Biosciences), using labeled blood leukocytes from the same animal and unlabeled milk cells as positive and negative controls, respectively. Variable numbers of samples from each animal were analyzed in order to collect at least 2–3 × 105 cells for each population. Cells were stored at −80°C for PCR analysis.
PCR analysis for the detection of JSRV proviral DNA was performed with the SC aliquots and each of the separated cell populations. Total DNA was extracted by a modified phenol–chloroform method. 18 DNA purity and concentration were estimated by optical density at 260 and 280 nm, and DNA was tested by U3 hemi-nested (U3-hn) PCR. 14 Three replicates (500 ng of test DNA each) were prepared for each SC aliquot, and 6 replicates (100 ng each) for the different cell populations. Appropriate positive and negative controls were used at each step. Amplification products were analyzed by electrophoresis in 2% (w/v) agarose gels with 0.5 mg/mL of ethidium bromide. Samples were considered positive when amplification was found in at least one of the replicates. Negative samples were tested by glyceraldehyde-3-phosphatase dehydrogenase (GAP-DH) PCR 14 to verify DNA quality and evaluate possible inhibitors of PCR.
None of the ewes had clinical signs of OPA throughout the study. At the end of the experimental period, all of them were euthanized; 7 ewes were lactating and 5 were in the dry period. The lungs were examined thoroughly postmortem, and samples were taken from any gross lesion suggestive of OPA plus 7 samples representing all lung lobes together with 4 samples from each mammary gland. Tissue samples were processed routinely for histologic examination and, to detect JSRV proteins, sections were evaluated by immunohistochemical procedures described previously. 7 A mouse monoclonal antibody against JSRV surface protein (encoded by the env gene) was utilized as primary antiserum. This antibody has been tested in natural OPA tumors with very high sensitivity and specificity. 20 In each assay, sections from naturally occurring confirmed cases of OPA were used as positive control tissue, and all of the tissues were IHC-positive. These controls were validated previously by PCR 14 and IHC. 7
Although the 8 ewes in group A had tested JSRV-positive by PCR testing of SCs from C/M in the previous lactation, positive results were detected in only 3 of the 8 ewes in this lactation. However, in the previous study, 3 more samples (n = 8) had been collected from each sheep throughout a lactation period and analyzed by PCR. None of the ewes was positive in more than one sample of milk SCs, and none of the samples was positive in more than one replicate (Table 1). The lack of repeatability of positive results over time resembles the inconsistent JSRV detection obtained when peripheral blood mononuclear cells (PBMCs) are analyzed by PCR in infected asymptomatic sheep.2,4,6 Considering the high sensitivity of the U3-hn PCR, which can detect a single copy of the JSRV template in a background of 500 ng of sheep genomic DNA, 14 and the absence of inhibition of GAP-DH PCR in the SC samples that tested JSRV-negative, these results are in agreement with those reported previously. 3 Our results also reinforce the hypothesis of a low level of JSRV infection in SCs from milk secretions, as in the case of PBMCs, 11 reducing the chance of virus detection in this lactation given that we tested a smaller number of samples. None of the samples of SCs from group B resulted positive (Table 1).
PCR detection of jaagsiekte sheep retrovirus (JSRV) in ovine milk cell populations, and immunohistochemical detection of JSRV in mammary gland and lung tissues.
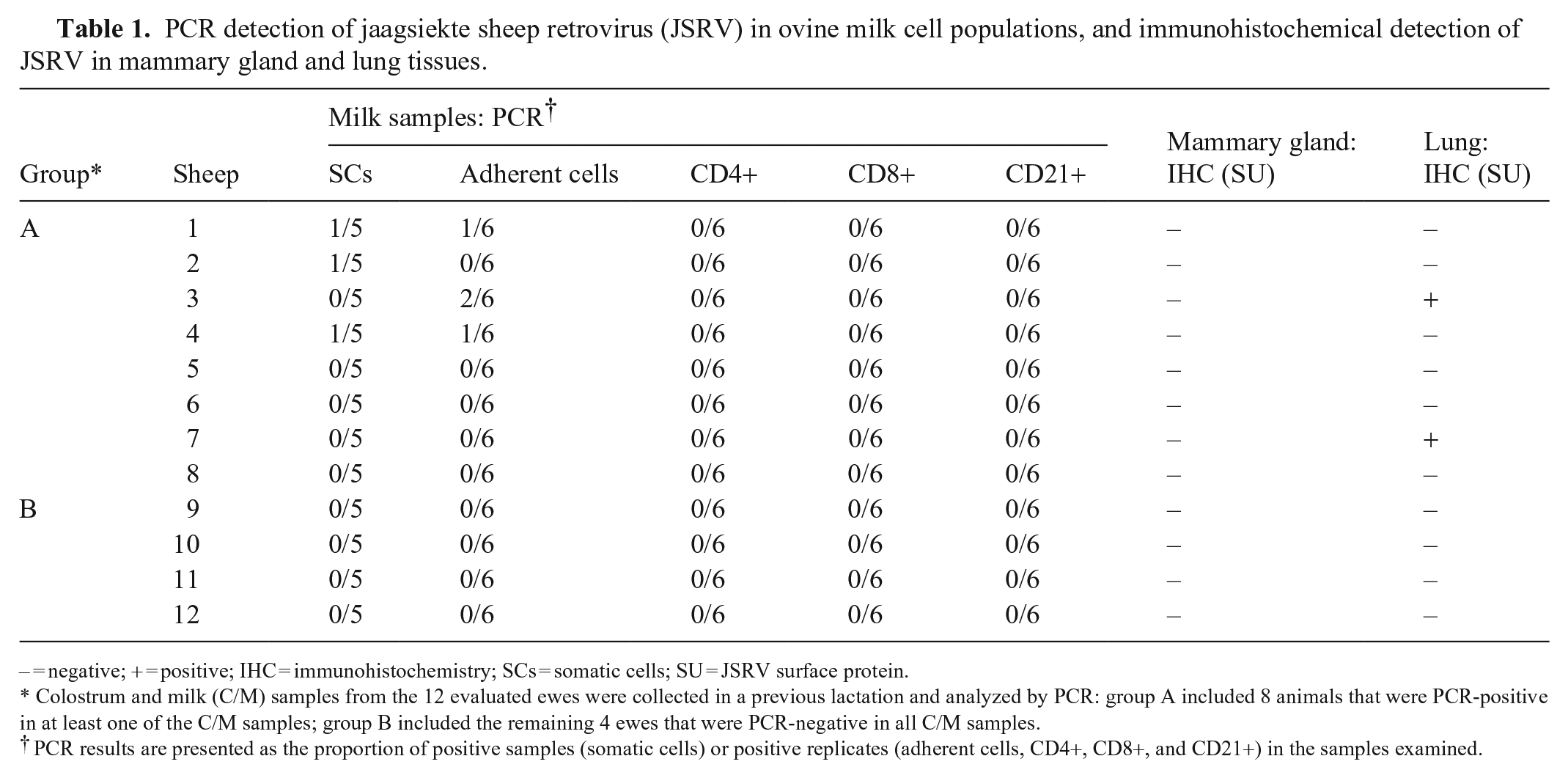
– = negative; + = positive; IHC = immunohistochemistry; SCs = somatic cells; SU = JSRV surface protein.
Colostrum and milk (C/M) samples from the 12 evaluated ewes were collected in a previous lactation and analyzed by PCR: group A included 8 animals that were PCR-positive in at least one of the C/M samples; group B included the remaining 4 ewes that were PCR-negative in all C/M samples.
PCR results are presented as the proportion of positive samples (somatic cells) or positive replicates (adherent cells, CD4+, CD8+, and CD21+) in the samples examined.
The PCR evaluation of the main populations of milk mononuclear leukocytes revealed JSRV proviral DNA only in adherent cells. We found JSRV-positive milk adherent cells in 3 of the 8 animals in group A; SCs from 2 of these ewes were also positive (Table 1). However, it was not possible to detect JSRV proviral DNA in CD4+ and CD8+ T cells or CD21+ B cells analyzed from this group. All samples from all populations of mononuclear leukocytes in group B were negative (Table 1). The low number of positive replicates detected in adherent cells (1 of 6 or 2 of 6), together with the absence of inhibition of GAP-DH PCR in those samples that tested JSRV-negative, could indicate a low frequency of infection in this milk cell population. Considering that 100 ng of DNA correspond to ~15,000 cells 11 and that 6 replicates of 100 ng were analyzed in each ewe, it can be estimated that at least 1 of 90,000 milk adherent cells were infected by JSRV in those sheep with PCR-positive results. A previous study revealed that proviral burden was higher in the adherent cells (macrophages or monocytes) than in lymphocytes (CD4+ T cells, CD8+ T cells, and B cells) isolated from mediastinal lymph nodes of sheep with OPA. 11 This phenomenon could also occur in milk secretions, and the low proviral load may be the cause of the lack of detection of JSRV in milk lymphocyte populations. In addition, the higher proviral burden in the adherent cells would explain the positive results obtained in this population in a sheep whose SCs tested negative (Table 1).
Immunohistochemical analysis of the mammary gland was negative in all of the animals regardless of their physiologic state, which would explain the absence of PCR-positive results in this tissue reported previously. 9 Given that we did not find labeled epithelial cells, it seems unlikely that there was active replication of JSRV in this type of cells, in contrast with observations in the mammary gland from animals infected with VMV or MMTV. The mammary gland is involved in the biological cycle of these viruses, which actively replicate in infected mammary epithelial cells and are excreted in the C/M as cell-associated and/or free viral particles and transmitted to offspring during lactation.1,16,17 Our results indicate that the mammary gland does not play such a role in the case of JSRV, and therefore, excretion of free viral particles in C/M seems unlikely. According to our findings and taking into account that mammary secretions contain leukocytes, including macrophages and lymphocytes that can reach the mammary gland from the bloodstream,12,13 JSRV-infected SCs found in C/M could have originated as peripheral blood leukocytes, in which the provirus can be detected in JSRV-positive sheep.4,9
None of the 12 studied sheep had clinical signs of OPA; nevertheless, small nodules suggestive of OPA (1–2 cm diameter) were detected in 2 of them and were confirmed as OPA lesions by histologic and immunohistochemical examination (Table 1).
We evaluated a limited number of sheep and milk samples, but we believe that our findings can be used as a solid base for future investigations. Our results show that JSRV can be found in the SCs of naturally JSRV-infected ewes, although OPA tumors may not have developed (Table 1), and are in agreement with several studies that have found JSRV provirus in lymphoid tissues and PBMCs prior to the onset of OPA lesions.2,4,9 Our previous research demonstrated that JSRV-infected cells present in C/M (possibly lymphoreticular cells) can cross the intestinal barrier of newborn lambs, reach the mesenteric lymph nodes, and enter into circulation. 3 Our current study contributes to the knowledge about this route of transmission, which should be taken into account in apparently healthy animals for the control of the disease. We demonstrated that milk from naturally JSRV-infected ewes can contain adherent cells, interpreted as mainly macrophages or monocytes, harboring proviral DNA in their genome, and suggest a major role of this cell type in the lactogenic transmission of the virus. However, the participation of milk lymphocytes cannot be ruled out, and more studies are needed to ascertain whether milk T and B cells can be infected with JSRV. In addition, our data suggest a lack of participation of the mammary gland in the biological cycle of JSRV that should be further investigated. This event would reduce the probability of excretion of free viral particles in C/M and reinforce the relevance of lymphoreticular cell involvement in the transmission of JSRV infection from ewes to offspring.
Footnotes
Acknowledgements
We are grateful to Dr. Dusty Miller for generously providing the antiserum against JSRV surface protein, and to M. Carmen Carreras for technical support in the isolation of milk leukocyte populations by flow cytometry.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was supported by a grant from the Spanish Ministerio de Ciencia e Innovación (AGL2008-00803), the Department of Science, Technology and University of the Government of Aragon, and the European Social Fund. Marta Borobia was a predoctoral research fellow under the Gobierno de Aragón programme (B103/09).
