Abstract
Management of prion diseases in livestock would benefit greatly from availability of a validated blood test. A promising immunocapillary electrophoresis technique (also known as capillary electrophoresis fluoroimmunoassay) to detect abnormal prion protein in blood from live sheep is evaluated here. Capillary electrophoresis fluoroimmunoassay was applied to analysis of extracted blood from scrapie-exposed sheep (n = 87; 347 samples) at various stages of incubation, and to control sheep (n = 194; 489 samples). Overall, test values for the control and test populations were not significantly different, and a similar proportion of control (7%) and test (10%) sheep were classified as positive. Over 2−3 month intervals from birth until clinical disease, test specificity and sensitivity ranged from 66.7% to 100% and 0% to 66.7%, respectively, indicating poor diagnostic performance at all stages of pathogenesis. In routine application, in its present form, the capillary electrophoresis fluoroimmunoassay procedure proved to be insufficiently robust for use as a blood test for scrapie diagnosis.
Keywords
Development of a blood test for transmissible spongiform encephalopathies (TSEs) remains a high priority in order to minimize the possibility of iatrogenic transmission through blood transfusion, to further understanding of pathogenesis, and to facilitate preclinical diagnosis and management of disease in both humans and livestock. Blood from TSE-affected subjects can be infectious, and infectivity may be present before clinical signs of disease are evident. This has been demonstrated, for sheep infected with bovine spongiform encephalopathy (BSE) or with scrapie, by blood transfusion into sheep. 5,13 Similarly, iatrogenic transmission by blood transfusion remains the probable cause of recent cases of variant Creutzfeldt-Jacob disease in humans. 18,21 Assessing infectivity or associated diagnostic markers remains problematic, however, because there are no validated blood-based assays for TSEs.
The presence in brain and other tissues of the pathological form of prion protein, PrPSc, is closely associated with infectivity and further, PrPSc represents the only validated diagnostic marker of TSEs. 2,16 A major challenge in developing a blood PrPSc test is that concentrations are likely to be several orders of magnitude less than found in brain.4−11 Current biochemical tests based on the analysis of PrPSc are only validated for CNS 1,26 and, to some extent, for nonneural tissues. 10,25 A number of alternative, promising procedures have been proposed recently and are at various stages of validation; these include PrPSc enrichment using prion-aptamers or RNA-ligand based adsorbents 9,23 and application of PrP amplification, 6 immuno-PCR, 3,8 or conformationally responsive labeled palindromic PrP peptides 12 to enable highly sensitive detection of PrPSc. Although some of these principles have been applied to analysis of PrPSc in blood, to our knowledge, their use for TSE diagnosis in livestock remains to be reported in the literature. The potential for BSE diagnosis using non-PrPSc-related surrogate biomarkers has recently been demonstrated by determination of biochemical signatures derived from midinfrared spectroscopy of dried blood serum films and data analysis using diagnostic pattern-recognition algorithms. 20
Immunocapillary electrophoresis (ICE, or capillary electrophoresis fluoroimmunoassay) is capable of very low detection limits, and has been demonstrated to offer much potential for blood PrPSc detection. 24 Immunocapillary electrophoresis is essentially a fluoroimmunoassay (FIA) which, following completion of the competitive immunoassay stage, uses capillary electrophoresis (CE) to separate free fluorescently labeled analyte from antibody-bound fluorescently labeled analyte, and laser induced fluorescence (LIF) detection to quantify label in the free and bound fractions as they flow through the capillary. This enables determination of the degree of competition between the label and analyte for limited antibody binding sites and thus assessment of the relative quantity of PrPSc in the sample. In recent limited studies, ICE analysis of blood from sheep enabled correct prediction of disease in 93%-100% of sheep (depending on the PrP genotype; ARQ/ ARQ, ARQ/VRQ, and VRQ/VRQ) 7−12 months after exposure to natural sources of scrapie infectivity: this indicated a hematogenous phase of disease well in advance of clinical symptoms. 17 The PrP genotype is an important determinant of scrapie pathogenesis because the time to onset of clinical disease is variable in sheep, being influenced by polymorphisms of the PrP gene at codons 136, 154, and 171, which are linked strongly to susceptibility (V at 136 and/or Q at 171) and resistance (the ARR allele) to classical scrapie 14 ; alleles are signified here using the single letter amino acid code. Although blood may be infectious during preclinical stages of disease, 15 understanding of the relationship between the PrP genotype, infectivity, and the presence of detectable PrPSc in blood, is very limited and clearly warrants more detailed investigation.
Given the importance of early TSE diagnosis and the lack of a validated blood test, the present study was undertaken to evaluate the merits of ICE for sensitive detection of PrPSc in blood of scrapie-infected sheep of selected genotypes. The aim was to investigate in detail the temporal occurrence of PrPSc with disease progression and thus to establish the potential of the test for presymptomatic scrapie diagnosis.
In this study, ICE was applied to blood PrPSc analysis to establish test performance in a routine laboratory environment. The test-sheep population was of various breeds (Swaledale, Welsh Mountain, and Dorset) and PrP genotypes, which had been exposed to natural infection from birth. Blood (the buffy-coat fraction) was either derived from recently archived material (sheep born in 2000) or collected prospectively from sheep selected specifically for this investigation (born in 2002, 2003, and 2004). The animals used were largely ARQ/ARQ, ARQ/VRQ, and VRQ/VRQ PrP genotypes (Table 1). Test animals were maintained from birth at the Veterinary Laboratories Agency-Weybridge, in a flock with a high incidence of scrapie, until disease was diagnosed. Control samples (Table 1) were collected from sheep that were direct progeny of scrapie-free New Zealand stock (New Zealand is believed to be free from scrapie) and maintained and bred on a closed, secure, scrapie-free farm. A single group of control samples, from ARQ/VRQ sheep born in 2000 and sampled at age 25−36 months (Table 1), were derived from a U.K. flock (closed to imported stock since 1972) with no evidence of scrapie, either historical or currently, as defined by clinical criteria or at cull, by postmortem examination. Sheep displaying clinical signs of scrapie were euthanized and subjected to postmortem (p-m) examination and scrapie status confirmed by standard methods (brain pathology, immunohistochemistry, and Western-blot analysis). 26
Sample collection, processing, storage, and analysis was essentially as described elsewhere 17 ; only the solid phase extraction (SPE) process differed in that a robotic processor was used. Briefly, the buffy-coat fraction from blood samples (20 ml) was washed twice and stored frozen (−80°C) until required (day 1). The buffy-coat preparations were lysed by 2 freeze-thaw cycles, incubated with DNase I, solubilized in sarcosyl (10%), and subjected to proteinase K (PK) digestion; the residual PK resistant PrPSc was extracted into hexafluoro-2-propanol (HFP), and the organic layer aspirated off and dried down overnight by centrifugal evaporation (day 2). The residue was redissolved in formic acid (0.05 M) and HFP, the PrPSc content further purified by hydrophilic interaction SPE (PolyHY-DROxYETHYL A(tm) as adsorbent) and the resulting eluate dried by centrifugal evaporation (45°C) overnight (day 3). The residue was redissolved in assay buffer (10 μl) and an aliquot (5 μl) analyzed by FIA, incubating overnight at 2−8°C (day 4). The FIA used a polyclonal antibody (AFF5) raised in rabbits against a synthetic peptide corresponding to a C-terminal amino acid sequence of ovine PrP (residues 223−237). The label was synthesized from the same peptide, with fluorescein isothiocyanate coupled to the N-terminus. Following the FIA incubation stage, free and antibody-bound label were separated by free-zone CE (day 5), and their relative proportions determined from the height of electropherogram peaks. Normalized test values (NTVs) were then calculated from the fraction of label bound for the test or control sample analysis, as a proportion of that bound for the reference negative control sample, as described. 17 Normalized test values enabled samples to be categorized according to a predetermined cutoff value, 24 which defined the outcome of the test as positive (NTV < 0.7) or negative (NTV ≥ 0.7).
Scrapie testing of various sheep PrP genotypes by ICE; assay sensitivity and specificity in relation to age and duration of exposure.
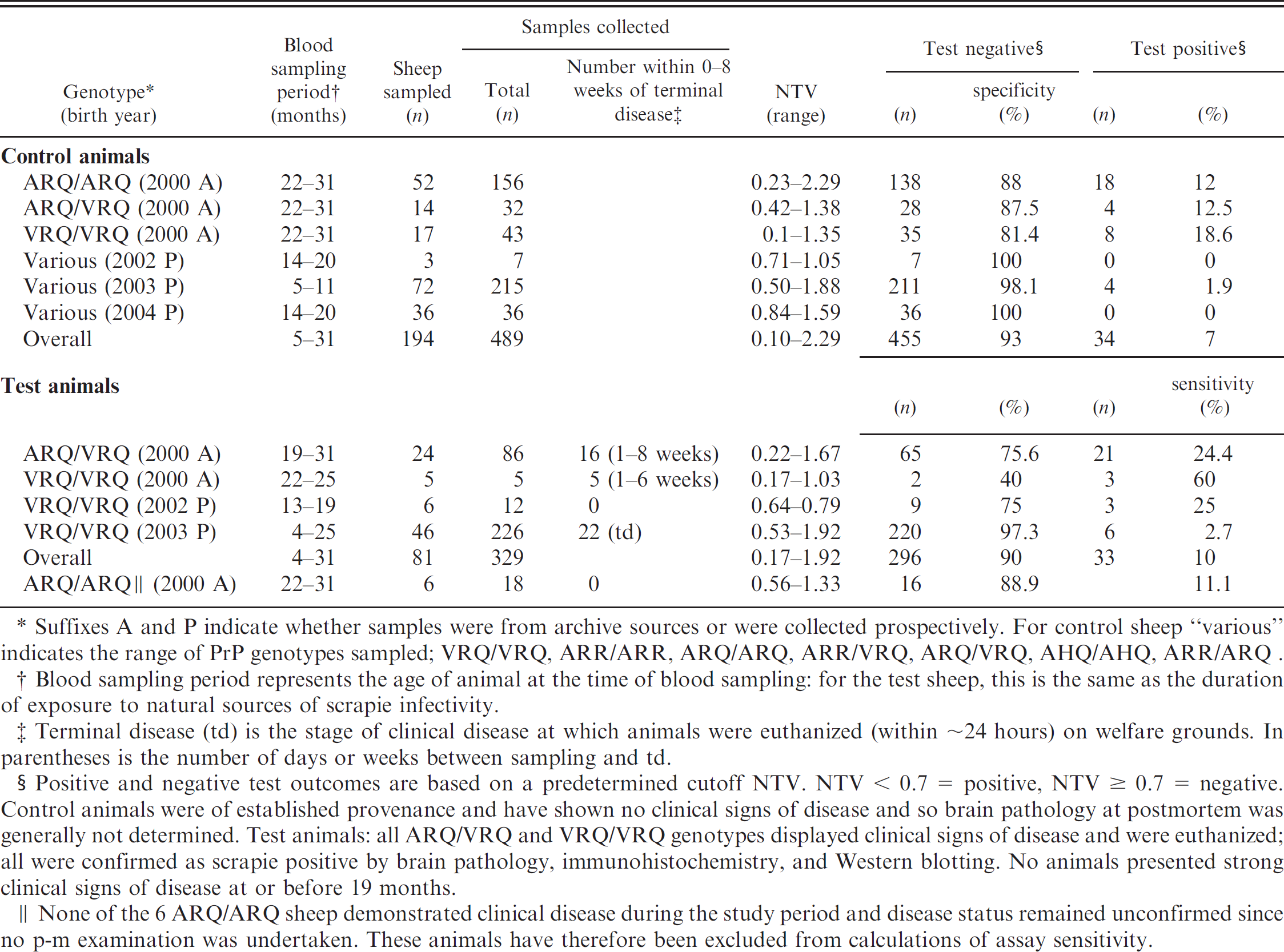
Suffixes A and P indicate whether samples were from archive sources or were collected prospectively. For control sheep “various” indicates the range of PrP genotypes sampled; VRQ/VRQ, ARR/ARR, ARQ/ARQ, ARR/VRQ, ARQ/VRQ, AHQ/AHQ, ARR/ARQ.
Blood sampling period represents the age of animal at the time of blood sampling: for the test sheep, this is the same as the duration of exposure to natural sources of scrapie infectivity.
Terminal disease (td) is the stage of clinical disease at which animals were euthanized (within ∼24 hours) on welfare grounds. In parentheses is the number of days or weeks between sampling and td.
Positive and negative test outcomes are based on a predetermined cutoff NTV. NTV < 0.7 = positive, NTV ≥ 0.7 = negative. Control animals were of established provenance and have shown no clinical signs of disease and so brain pathology at postmortem was generally not determined. Test animals: all ARQ/VRQ and VRQ/VRQ genotypes displayed clinical signs of disease and were euthanized; all were confirmed as scrapie positive by brain pathology, immunohistochemistry, and Western blotting. No animals presented strong clinical signs of disease at or before 19 months.
None of the 6 ARQ/ARQ sheep demonstrated clinical disease during the study period and disease status remained unconfirmed since no p-m examination was undertaken. These animals have therefore been excluded from calculations of assay sensitivity.
The performance of the ICE procedure was assessed continuously by reference to sample blanks (SB) and quality control (QC) samples assayed within each assay batch. These were prepared from pooled bulk control buffy-coat sample extracts, dissolved in a proportionate volume of assay buffer as described above, and stored frozen (-80°C) as single-use aliquots. The QC sample extracts were fortified with ovine recombinant PrP (ARQ allele, full length protein, 22 to 1 and 2 ng rPrP/ml) before storage. The SB samples were used to normalize test and control sample results 17 to provide NTVs; these gave a mean (±SD) ratio of bound:free label between all assays (n = 69) of 1.01 ± 0.23. Similarly, QC samples gave mean ±SD NTVs in = 32) of 0.86 ± 0.17 and 0.58 ± 0.12 at 1 and 2 ng rPrP/ml−1, respectively.
Application of the predetermined NTV cutoff criterion enabled assay performance to be assessed in terms of sensitivity (the number [%] of confirmed scrapie-positive animals that tested positive) and specificity (the number [%] of uninfected animals that tested negative). Specificity was evaluated from the analysis of 489 blood samples from 194 control animals; these gave a mean ±SD NTV of 1.00 ± 0.25. Thirty-four (7%) of these samples gave NTVs < 0.70 (Table 1), indicating an overall test specificity of 93% (range 66.7%-100%, depending on sample time point). No culled control animals showed clinical signs of disease or gave a positive p-m scrapie diagnosis based on standard validated confirmatory tests on brain. Assay specificities for archived samples from VRQ/VRQ (81.4%, n = 8), ARQ/ARQ (88%, n = 18), and ARQ/VRQ (87.5%, n = 4) were similar, whereas those for prospective samples gave a higher overall specificity (98.4%; 4 of 254 samples). The interval between sample collection and analysis varied from ∼2−24 months for prospectively collected material, and ranged up to ∼36 months for archived samples. It has not been established whether specificity was influenced by the duration of sample storage at −80°C because this was beyond the scope of this study. For all practical purposes, there were no significant differences between mean NTVs in relation to period of sampling or genotype, and this justified comparison between test (below) and control time windows, irrespective of genotype.
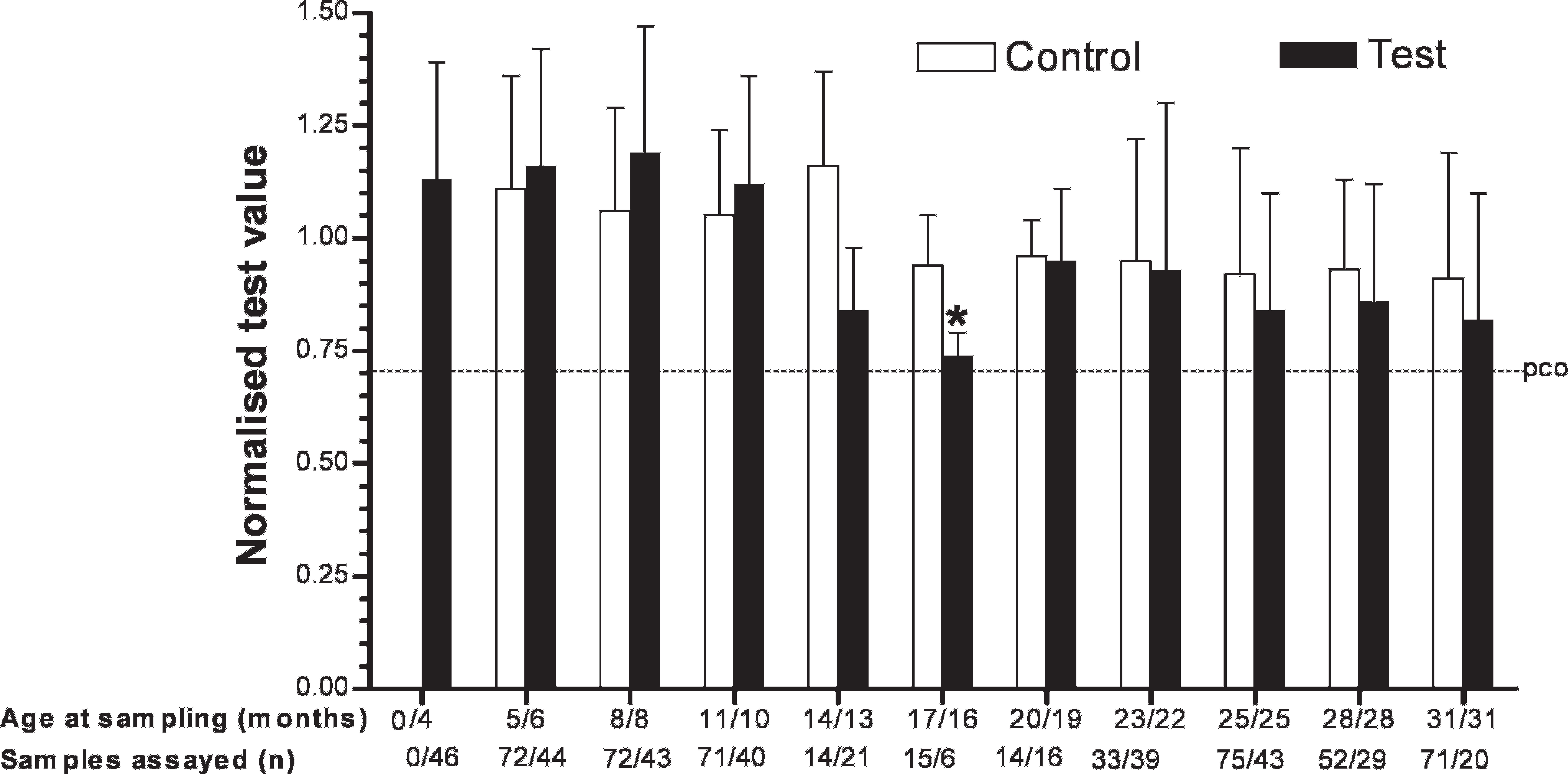
Normalized test values (mean ± SD) of control and test samples from sheep ranging from 4 to 31 months of age, or months of exposure to scrapie. There was a significant difference between NTVs from control and test samples at 17 and 16 months, respectively (P = 0.043). pco = predetermined cutoff (NTV = 0.7).
A total of 329 blood samples were analyzed from 81 test sheep of the VRQ/VRQ and ARQ/VRQ genotypes (Table 1). Thirty-three samples gave NTVs <0.7 (all animals confirmed positive), giving an overall test sensitivity of 10% (range 0%−66.7%, depending on sample time point). The diagnostic sensitivity for the VRQ/VRQ (n = 243) and ARQ/VRQ (n = 86) genotype populations were 4.9% (over all 3 groups) and 24.4%, respectively (Table 1). At best, diagnostic sensitivity of 50%-67% was observed for small sets of 2−3 samples (n = 7 in total) during the time windows 19, 22, and 25 months (VRQ/VRQ sheep born in 2000 and 2002). This outcome was not reproducible, however, because NTVs of a much larger sample population (n = 36) from sheep born in 2003 indicated zero sensitivity at age 19−25 months. Using the predefined cutoff-point criterion, a similar proportion of sheep were classified overall as positive in the control (7%) as in the scrapie exposed (10%) group over the study period (Table 1). A further 18 blood samples were tested from ARQ/ARQ sheep (n = 6), of which 2 gave a positive NTV (Table 1) but data were not included in calculations of test sensitivity because no clinical disease was observed during the study period and disease status remained unconfirmed (no p-m examination was undertaken).
Assessing assay performance by summary determination of sensitivity over the subjects’ lifetime is clearly simplistic, because the various genotypes have different clinical susceptibilities and the timing of any hematogenous phase of disease may likewise vary. This was highlighted in a previous evaluation of ICE for scrapie diagnosis that reported 100% test sensitivity 9−13 months after exposure (14% in older sheep) to scrapie (VRQ/VRQ genotype). 17 In the present study, determination of the sensitivity at particular time windows was possible by reference to the comprehensively sampled VRQ/VRQ population. The testing of 238 samples from the prospectively sampled VRQ/VRQ sheep (Table 1) indicated a poor overall test sensitivity of 3.8% (2002, 25%, n = 12; 2003, 2.7%, n = 226), however. Furthermore, detailed comparison of data from each of the ∼3-month time windows failed to identify any consistent period during which ICE gave diagnostic sensitivity of practical significance (Table 1, Fig. 1). Using the pre-determined cutoff value, sensitivity ranged from 16.7%−50% and 0−17.7%, for 2002 and 2003 sheep, respectively, and examination of comparable windows for test and control sample population NTVs revealed no consistent significant differences (one-tailed unpaired Student's í-test). Thus although, for instance, control NTVs at 17 months were significantly different (P = 0.043) from those for test samples (n = 6, VRQ/VRQ, 2002) at 16 months (Fig. 1), the diagnostic value is questionable against the background of low test sensitivity (16.7%) at 16 months.
In summary, the diagnostic value of the ICE test in these studies proved to be poor. Recent studies using this approach reported detection of scrapie with high sensitivity (92.8%) in sheep of VRQ/VRQ and VRQ/ARQ genotypes in the age range (period of exposure) 7−12 months. 17 This is clearly at odds with the outcome of this larger and more comprehensive investigation, which has examined samples from VRQ/VRQ sheep from birth until clinical disease, but found no compelling evidence that routine application of the ICE procedure was effective for diagnosis at any stage of disease. The findings here are thus in keeping with the inconsistent diagnostic performance of ICE observed when applied to Creutzfeldt-Jacob disease in humans and chimpanzees: using the predetermined cutoff, the test proved to be 100% specific but incompletely (55%) sensitive, 19 and NTVs from infected and control groups could not be distinguished. 7,19 In the later studies, the favored explanation for poor performance was lack of reproducibility associated with the extraction procedure, perturbations of the immunoassay, or both. This is a reasonable generalized explanation for the evident lack of robustness, because the assay process is lengthy, involves multistage extraction procedures and a sophisticated CE based process. Together, these stages present more potential sources of variability than is generally the case for more widely used immunoassay formats. Furthermore, despite the promising outcome of the recent application of ICE to scrapie diagnosis, 17 the authors nevertheless pointed to a broad requirement to establish more robust conditions for most stages of the assay by way of refinement of extraction procedures, conditions for capillary electrophoresis, and by use of higher affinity antibodies (to improve the lower limit of detection). These needs are supported by the QC data presented here which, although representing variability of only the immunoassay and CE stages of the procedure, indicate between assay CVs for these samples of ∼20%; this is appreciable, given that these values exclude errors associated with the extraction process.
Whatever the potential merits of the ICE assay principle, it is clear that more reproducible and robust procedures are required of an antemortem diagnostic blood test for scrapie. Undoubtedly label detection by LIF lends exquisite sensitivity to the end-point determination in ICE, however, this represents only one factor contributing to overall assay sensitivity and in this application this potential is substantially defrayed in practice by other assay performance limitations. It remains to be established whether alternative extraction techniques and labeling technologies combined with more conventional immunoassays can better deliver the level of assay precision and sensitivity required to meet the substantial challenge of PrPSc detection in blood.
Acknowledgements. The authors wish to express their gratitude to the Department for Environment, Food and Rural Affairs (Defra), United Kingdom, for funding this work (Grant TS5970). Thanks to Drs. Danny Mathews and Jim Hope for constructive criticisms. The authors are indebted to Dr. Hugh Simmons and Sue Bellworthy and their staff for collection and provision of blood samples from control and test flocks, to Jemma Thorne and John Docherty for sample preparation, and to Robin Sayers for statistical advice. Many thanks to Dr. Pierre Sarradin for the generous supply of ovine recombinant PrP. Copyright ©2007 British Crown.
