Abstract
Prion protein (PrP) is encoded by the
Scrapie is a transmissible spongiform encephalopathy (TSE) that affects sheep and goats. Like all TSEs, it is characterized by spongiform lesions and accumulation in the central nervous system of an abnormal isoform (PrPSc) of the normal host-encoded cellular prion protein (PrPC). 18 To date, PrPSc represents the only diagnostic marker, and the majority of testing procedures are based on its immunological detection after the proteolysis of endogenous PrPC. Although scrapie in the ovine population has been known for more than 2 centuries, 9 it was reported in goats only in the early 1940s in France 8 followed by reports from other countries, including Italy. 7 Since the implementation of an active TSE surveillance system in Europe in 2002, widespread testing for TSE has lead to an increase in the detection of scrapie in both sheep and goats.
Susceptibility to classical scrapie in sheep is known to be linked to polymorphisms of the PrP encoding gene (
Genetic studies have shown that the caprine
Several monoclonal antibodies (mAbs) targeting different PrP regions have been developed for scrapie diagnosis but none of them, even when used for scrapie diagnosis in goats, have been developed specifically for PrP detection in this species. It should be expected that some antibodies, particularly those that target epitopes at polymorphic sites, will not be universally suitable for the detection of PrP in this species. To evaluate the capacity of some mAbs to bind PrP of goats bearing different
To further investigate this reactivity pattern, Western blot analyses were performed on the brain tissue of 30 scrapie-negative samples with the 3 different genotypes at codon 222: 6 goats homozygous for lysine (222K/K), 12 heterozygous goats (222Q/K), and 12 wild-type goats (222Q/Q). Prion protein extraction from each sample was carried out as follows: 10% (w/v) homogenates of brain tissue were prepared in lysis buffer (10% N-lauroylsarcosine diluted in Tris-buffered saline [TBS], pH 7.4) and clarified by centrifugation at 22,000 RCF for 20 min. a A volume of 10 µl of a 100 mM phenylmethylsulphonyl fluoride b solution was then added to 1 ml of each supernatant; the samples were centrifuged at 215,000 RCF for 1 hr. c The pellets were dissolved in Laemmli buffer and subjected to sodium dodecyl sulfate–polyacrylamide gel electrophoresis on a 12% handmade mini-gel (acrylamide/bisacrylamide ratio of 37.5:1) and then transferred onto polyvinylidene difluoride membranes. d Blots were blocked by Tris-buffered saline–bovine serum albumin 5% and incubated with mAb F99/97.6.1 e (4 µg/ml) recognizing PrP residues 220–225. 16 Immunodetection was carried out with an alkaline phosphatase–conjugated goat anti-mouse immunoglobulin G revealed by a chemiluminescent substrate. The images of the blots were captured and analyzed using commercial software. f
In order to make the 30 samples comparable, the amount of tissue equivalent (mg/ml) to be loaded for each sample was previously calculated by normalizing the signal. Briefly, after PrP extraction, 8 serial dilutions (1:2–1:256) were prepared from the pellet of each sample. Blots were revealed with mAb P4 g (0.1 µg/ml), which recognizes PrP residues 93–99 and therefore is not affected by the 222 polymorphism. The PrP signals were quantified by densitometry. Dilutions with similar optical densities were selected from each sample, and the tissue equivalents (mg/ml) to be used as the starting amount for the next analysis were then calculated. After signal normalization, each sample was analyzed in triplicate, again making 8 serial dilutions (1:2–1:256). Scrapie-negative ovine brain was included as a control for each blot. Furthermore, the samples were always revealed in parallel with mAb P4 to check that signal normalization had been correctly made.
No PrPC signal was detected by mAb F99/97.6.1 in any of the goat samples with genotype 222K/K, differently from mAb P4 (Fig. 1A, 1B). The PrPC signals of the 222Q/K goats were revealed with an apparently lower sensitivity (1 dilution) than those obtained from the 222Q/Q samples (Fig. 2A, 2C). Instead, the PrPC signals were detected by mAb P4 with similar sensitivity in all the analyzed samples, independently of their polymorphisms (Fig. 2B, 2D, 2F). The apparent lower sensitivity of mAb F99/97.6.1 in the 222Q/K samples compared to the 222Q/Q samples suggested that mAb F99/97.6.1 could discriminate heterozygous from wild-type animals. In order to verify this hypothesis, the 12 Q/K and the 12 Q/Q samples were again analyzed by Western blot as described above, this time loading them undiluted and revealing the PrPC signals both with mAbs F99/97.6.1 and SAF84
h
(1 µg/ml), which targets PrP residues 163–173 and shows a sensitivity similar to mAb F99/97.6.1, as shown in a preliminary study (data not shown). The possibility of recognizing the different genotypes of the samples was then verified by calculating the ratio between the signal intensity given by mAb F99/97.6.1 and that given by mAb SAF84. Again, the samples were analyzed in triplicate and the optical densities given by mAbs F99/97.6.1 and SAF84 were measured by commercial software.
f
In order to facilitate the normality and the homoskedasticity of the distribution of the ratio, a logarithmic transformation of the F99/SAF84 ratio (
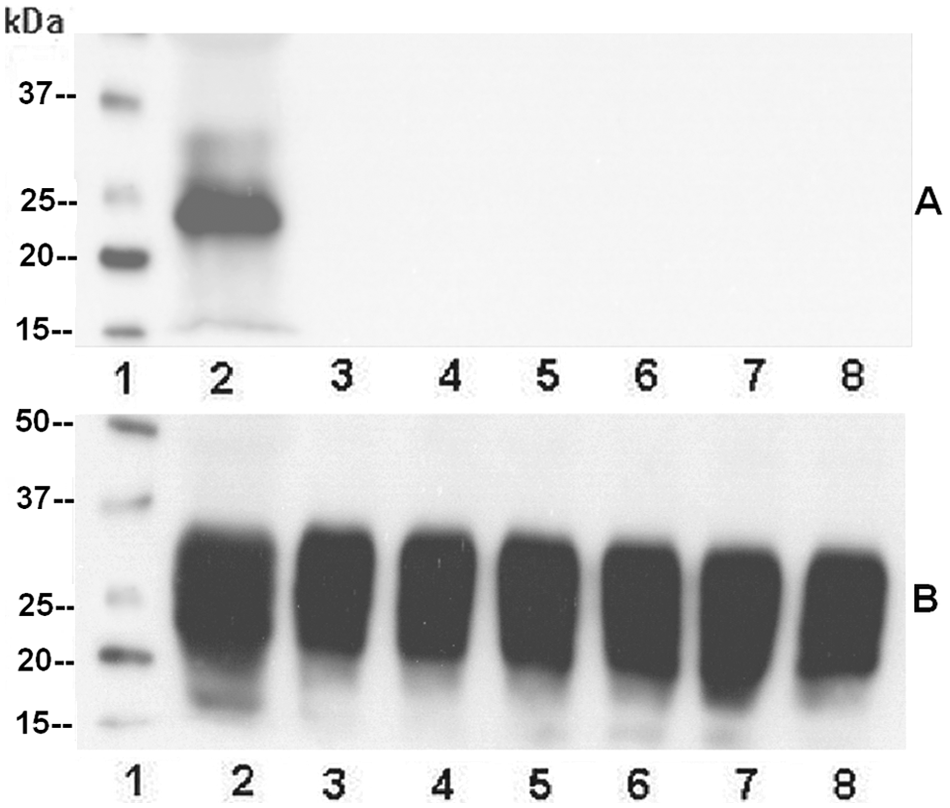
Western blot analysis of six 222K/K samples analyzed by monoclonal antibodies F99/97.6.1 (
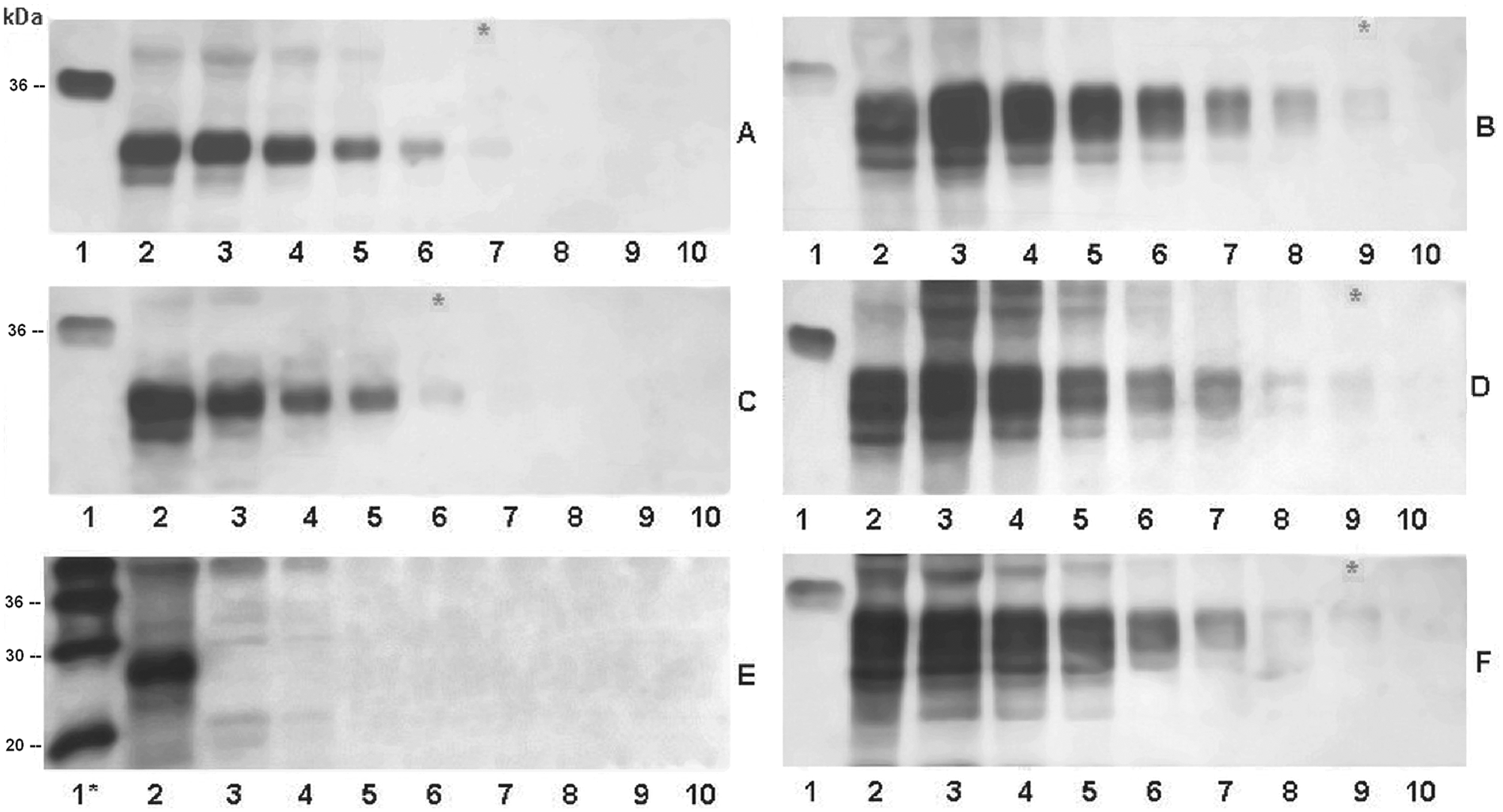
Western blot analysis of 3 samples, representative of each genotype at codon 222, analyzed by monoclonal antibodies F99/97.6.1 (left column) and P4 (right column).
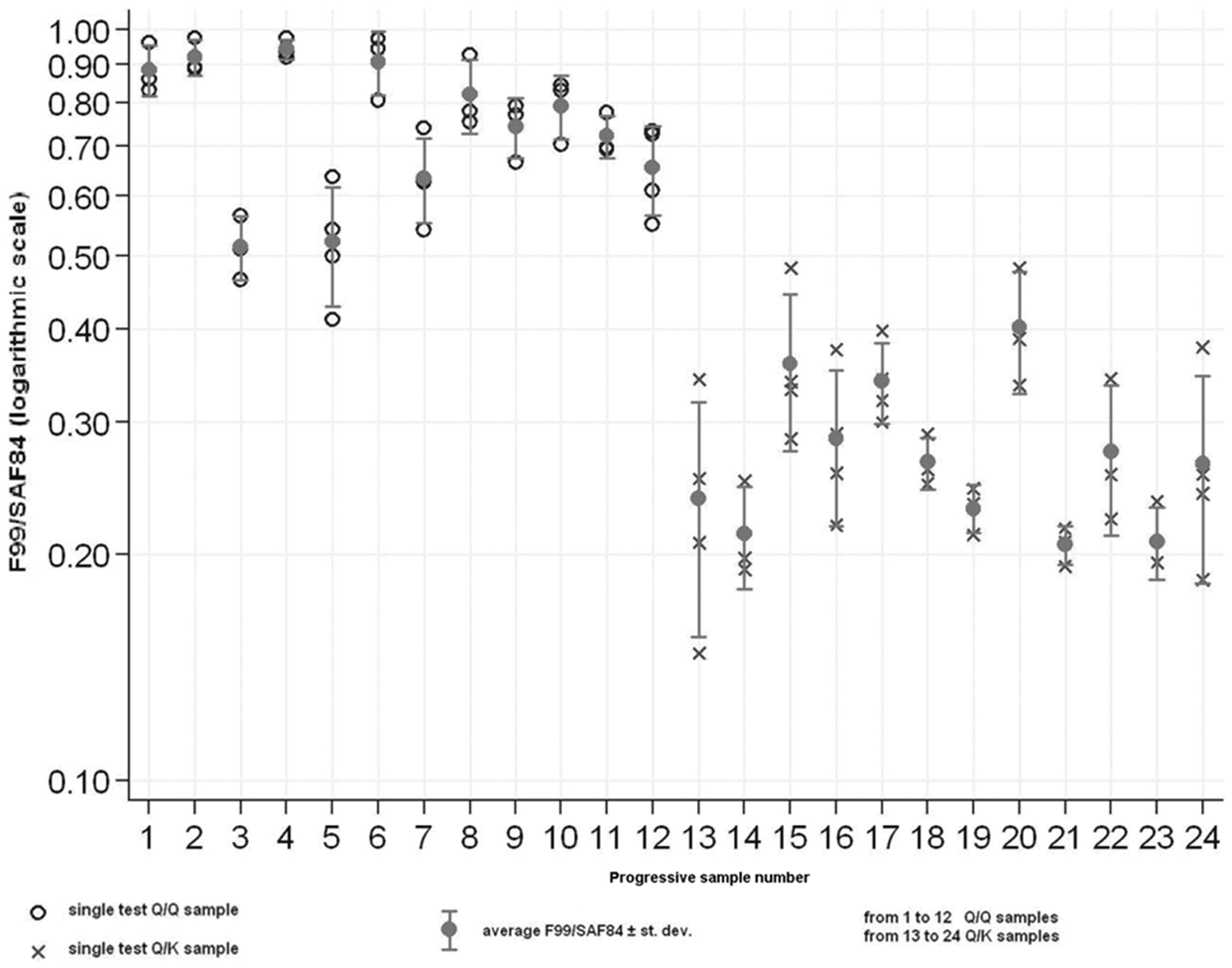
Scatter graph of F99/97.6.1/SAF84 ratio of 222Q/Q (samples 1–12, empty circles) and 222Q/K (samples 13–24, crosses), represented in logarithmic scale. Filled circles represent the average F99/97.6.1/SAF84 ratio for each sample, and standard deviation is reported.
The results obtained in the current study revealed an influence of the substitution of glutamine to lysine at codon 222 on the capacity of mAb F99/97.6.1 to bind the goat PrP. This mAb, generated by immunization with a synthetic peptide comprising the residues 217–231 of the ovine PrP,
16
recognizes a very short epitope on the PrP (220QY
The absence of binding between mAb F99/97.6.1 and the 222K PrP variant also explains the lower reactivity obtained in the heterozygous goats compared to the wild-type animals. The statistical analysis carried out on the values of PrPC signals obtained by the ratio F99/SAF84 allowed to outline statistically significant differences between 222Q/Q and 222Q/K samples and, by these criteria, it enabled the authors to distinguish the different genotypes at codon 222. Because the 222K allele could be a target for genetic selection in goats, the differential reactivity of mAb F99/97.6.1 could be exploited to set up a genotyping test (i.e., an ELISA method that would identify scrapie-resistant animals), as has been done using mAb 211 for codon 171 in sheep. 5
A study based on the capacity of mAb SAF84 to bind ovine PrP, in connection to 171Q-containing allelotypes, allowed the investigation of the ARR allele contribution to scrapie prion formation in naturally infected heterozygous 171Q/R animals. 13 In previous experimental studies, 222Q/K genotypes have been demonstrated to increase resistance to classical scrapie, at least prolonging the incubation period,2,22 but some 222 heterozygous scrapie-positive goats have been reported in field studies3,6,11: investigations on the contribution to PrPSc formation given by the 222K allele could be carried out using mAb F99/97.6.1. Finally, the demonstrated different reactivity of mAb F99/97.6.1 could also be a useful biochemical approach to identify any false-positive scrapie cases in goats with genotype 222K/K, deriving from cross-contamination with wild-type positive samples. Indeed, such an event cannot be completely excluded during contemporary sampling of numerous animals in TSE outbreaks.
Footnotes
a.
Ultracentrifuge Optima TLX, Rotor TLA 55; Beckman Coulter Inc., Fullerton, CA.
b.
Sigma-Aldrich, St. Louis, MO.
c.
Ultracentrifuge Optima TLX, Rotor TLA 110; Beckman Coulter Inc., Fullerton, CA.
d.
Millipore Corp., Billerica, MA.
e.
VMRD Inc., Pullman, WA.
f.
UVI Prochemi software, Uvitech, Cambridge, UK.
g.
R-Biopharm, Darmstadt, Germany.
h.
Cayman Chemical Co., Ann Arbor, MI.
Declaration of conflicting interests
The author(s) declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grant GoatBSE–Specific Targeted Research Project (FOOD-CT-2006-36353).
