Abstract
Brief introduction
Chondrosarcoma accounts for nearly one-third of adult musculoskeletal malignant tumors and its incidence ranks the second only to osteosarcoma.1–3 Histologically, chondrosarcomas are characterized by non‐osteoid cartilage matrix produced by neoplastic cells, including primary chondrosarcomas derived from sporadic mutations and secondary chondrosarcomas originating from malignant transformations of benign cartilaginous lesions, such as osteochondromas or enchondromas.4,5 The histological grade is one of the most powerful prognostic factors for overall survival (OS), metastasis and recurrence. Lower-grade tumors with less cellularity, rich cartilage matrix and poor metastatic behavior are prone to favorable outcomes after surgical curettage or resection, with survival rates between 88.5% and 95.8% at 5 years.3,6,7 On the other hand, metastatic relapse was observed in 70% of higher-grade cases, and advanced cases exhibited a higher grade of malignancy, with the median OS reported as poor as 18 months.6,8,9 Patients with higher-grade chondrosarcomas, especially in advanced clinical settings, might be insensitive or even resistant to conventional chemotherapy.4,8,10 At present, few prognostic studies have focused on the efficacy of chemotherapy for higher-grade tumors.
Tumor size, stage, pathological subtype, age, location and margin status are among the key prognostic indicators that have been identified to influence the survival of patients with chondrosarcoma.1,2,11,12 However, the clinical profiles of patients have changed due to the evolution of treatment regimes in recent years, which affects the reliability of previously published studies.2,4 On the other hand, owing to the low morbidity and scarcity of several pathological types, most investigations were subjected to single-institution series with limited sample size, which makes clinical decision-making and survival prediction difficult.11,13–15 The large multi‐institutional datasets provided by the Surveillance, Epidemiology, and End Results (SEER) database ensure statistical power and population‐level representation of the rarity such as chondrosarcoma. 16 The analysis of the SEER database will surely be a practical way to investigate the characteristics of chondrosarcoma in a modern context. The SEER database has been kept up-to-date with new cases enrolled over the past decade. To date, no study has comprehensively evaluated the prognosis of high-grade chondrosarcoma via assessing the latest edition of the SEER database.
The Cox proportional hazards (CoxPH) model is widely accepted for integrating and measuring significant clinical factors of patients when evaluating the odds of occurrence of events.1,13,17 However, most studies using the CoxPH model merely focused on using linearity assumptions rather than translating nonlinear variables into predictive models for real-world practice. Additionally, the conventional statistical model does not consider the fact that the effect of predictor variables on individual patients changes at different time points, which is far from providing comprehensive insight into the patient’s long-term outcomes. 11 Computational approaches such as machine learning and deep learning have been reported in long-term survival prediction of chondrosarcoma.18,19 Machine learning enables recognizing complex combinations of predictors from the existing huge amounts of data and performing model improvement over time. 20 The limitations of single machine learning models are obvious in model interpretability and flexibility due to simplification of the event when dealing with multi-dimensional information. Moreover, these models lack stability due to bias related to data collection or model selection.20,21 Compared to single machine learning models, ensemble learning model exhibits improved discriminative performance and enhanced data handling capabilities. The performance improvement of ensemble learning model has been reported to reach 25%. 22 On the other hand, deep learning tends to overfit when dealing with relatively small amounts of data. The average performance of ensemble learning has been reported to be approximately 8% greater than that of deep learning models.21,23,24 Overall, ensemble learning takes non-proportionalities, multicollinearity and nonlinearity of datasets into account, which produces better calibration and discriminative performance by combining results from multiple models. 20 The application of ensemble learning methods in therapy decisions and prognostic prediction for patients with chondrosarcoma has not received much attention in recent studies.
The primary objective of this study was to develop the ensemble learning algorithm with selected prognostic features which outperforms the Cox regression and single computational models in survival prediction and analysis of benefit of chemotherapy of high-grade chondrosarcoma based on SEER data. The secondary objective was to implement the optimal algorithm for identifying specific patient groups most likely to benefit from chemotherapy and guidance in chemotherapy strategies for high-grade chondrosarcoma.
Materials and methods
Patient selection and data collection
A flowchart of the detailed study process was shown in Figure 1. The data of patients with diagnosis of chondrosarcoma from January 2000 to December 2019 were obtained from the Surveillance, Epidemiology, and End Results (SEER) database. The International Classification of Diseases for Oncology, 3rd edition (ICD-O-3), was used to identify patients with chondrosarcoma. Inclusion criteria were all the diagnoses of chondrosarcoma as per ICD-O-3 definition, including those with ICD-O-3 code 9220/3 (Chondrosarcoma, NOS), 9221/3 (Juxtacortical chondrosarcoma), 9231/3 (Myxoid chondrosarcoma), 9240/3 (Mesenchymal chondrosarcoma), 9242/3 (Clear cell chondrosarcoma), and 9243/3 (Dedifferentiated chondrosarcoma). Patients with missing or ambiguous information, a secondary tumor at diagnosis, or a primary site of tumor other than the bone or joints were excluded from the study. In total, 1931 patients were included. The histological grade of the SEER database consists of four categories, with Grades I, II, III, and IV refer to well differentiated, moderately differentiated, poorly differentiated, and undifferentiated lesions, respectively.16,25 Tumor grade was classified as either high (Grade III or IV, 468 patients) or low (Grade I or II, 1463 patients) according to the commonly recognized standards in the clinical and academic fields.1,18,26,27 The patients with low-grade chondrosarcoma (n = 1463) were further excluded and 468 patients with high-grade chondrosarcoma were finally included in this study. Ethical approval was not sought for this study as the studies using the SEER database, including this one, were exempt from institutional review board approval. Informed consent was not applicable because the data from the SEER database were anonymous, and the study was an observational one. Study profile and analysis pipeline.
Data preprocessing and feature selection
The ordinal features were encoded as ordinal numeric values. Binary categorical features were coded as 0 or 1. Dummy encoding was used to deal with categorical features. Kaplan-Meier analyses were used for evaluating overall survival (OS) preliminarily, with the log-rank test used for determining the statistical difference between the estimated survival curves of different chondrosarcoma grades. CoxPH models and random survival forests (RSF) were applied to select the potential features that were associated with overall survival benefit in patients with high-grade chondrosarcoma for further model training. The concordance index (c-index) was used to evaluate the predictive power of the Cox regression model. The permutation importance is a method used to evaluate the contribution of each feature to the predictive power of a model. In this study, the RSF model combined with the permutation importance method was used to evaluate the importance of clinical characteristics. Among the five features with the smallest ranked value of c-index according to cox model, those with an RSF mean importance of no more than 0 were considered as lowly correlated features and were merged into reference features.
Model design and development
Ensemble learning algorithm was used in model design and development for prognostic prediction of high-grade chondrosarcoma in this study. The primary predictive outcome was overall survival (OS). The subjects were separated into training (70%, n = 327) and test (30%, n = 141) sets. For comparing ensemble learning model with single computational models, survival support vector machine regression models with different kernels (polynomial and radial basis function kernels) were developed and trained. These single models were chosen due to their ability to predict survival time quantitatively based on prior work on dealing with nonlinearity of clinical features. Moreover, a multivariate CoxPH model was constructed for comparison. To reduce the influence of potential confounding variables, the RSF model retrained on the training set was used as a feature scaler. The permutation importance of the obtained features was weighted to each feature by linear transformation. The model training involved a weighted summation of survival support vector machines trained with different kernel functions. To find the best configuration for our proposed model, hyperparameter tuning was conducted through 1000 iterations of random search and cross-validation on the training dataset. After the training was completed, the models were weighted and integrated to obtain the final model.
Model evaluation
The models were assessed for discrimination, calibration, and overall performance. The concordance index (c-index) based on the inverse probability of censoring weights was selected as the measurement of discrimination. Its value ranged from 0.5 to 1.0, with 1.0 indicating perfect discrimination. Receiver operating characteristic (ROC) curves and area under the curve (AUC) values at each time point (from the 12th to the 120th month, every 6 months) were obtained for evaluating the time-dependent specificities and sensitivities of the models. The survival data predicted by the ensemble learning model with the real data from the test set were presented case by case via a scatter plot, respectively.
Model interpretation
For prognostic evaluation of the effect of chemotherapy, we included different prognostic features in the ensemble learning model for survival prediction. The virtual patients for prediction were generated in each group according to different age groups (10, 20, 30, 40, 50, 60, 70, or 80 years old) with specific prognostic factors selected. We adjusted the imbalanced numbers of the patients in each age group with specific factors selected, which enables balanced and comparable sample size for matched age-stratified survival analysis. Some clinical features with fewer patients in real datasets could therefore be evaluated. Among these virtual patients with an estimated survival of more than 36 months in each group, the numbers of those who received chemotherapy to those who did not receive chemotherapy were counted to determine the ratio. In addition, χ2 tests (n > 40) and Fisher exact test (n ≤ 40) were performed in each group to compare the number of the virtual patients who received chemotherapy and those who did not receive chemotherapy, with specific prognostic factor selected and the estimated survival more than or no more than 36 months.
For further identification of the effect of chemotherapy with specific prognostic factors selected, the virtual patients in each group were further divided into subgroup A, B and C. Subgroup A contained those with longer predicted survival time if receiving chemotherapy regardless of tumor size. Subgroup B contained those with specific tumor size intervals which could be referred to for choosing chemotherapy or not to get longer survival time. Subgroup C contained those with shorter predicted survival time if receiving chemotherapy, regardless of tumor size. We counted the patients in each subgroup as a percentage of the total number of patients. Among the different age groups with specific prognostic factor selected, if the percentage of subgroup A differed from that of subgroup C by more than 25%, the patients in this group were considered to benefit from chemotherapy.
Statistical analysis
Percentages and frequencies were used to characterize categorical variables. Medians and ranges were used to characterize continuous variables. Clinical characteristics were compared using a Wilcoxon rank sum test for continuous variables and Fisher exact test for categorical variables. Log-rank test was performed to evaluate the statistical significance of the differences between the survival curves. The 36-month survival time was used as the standard for judging the pros and cons of chemotherapy in different age groups, with specific prognostic factor selected. χ2 testing (n > 40) and Fisher exact test (n ≤ 40) were used to examine the differences between the number of the patients who received chemotherapy and those who did not receive chemotherapy, with specific prognostic factor selected and estimated survival more than or no more than 36 months. p < .05 was considered statistically significant.
Python packages
The python environment was based on version 3.7. Numpy and Pandas were used to construct the basic data structure. Scikit-learn, lifelines and Scikit-survival were used to train and validate the survival-related machine models. All the images in this article were plotted with matplotlib. Supporting information shows further details of the code [see Supplemental Material].
Results
Demographic and clinical features of high-grade chondrosarcoma patients for prognostic evaluation based on SEER data
Demographic and clinical characteristics of the patients with high- and low-grade chondrosarcoma.
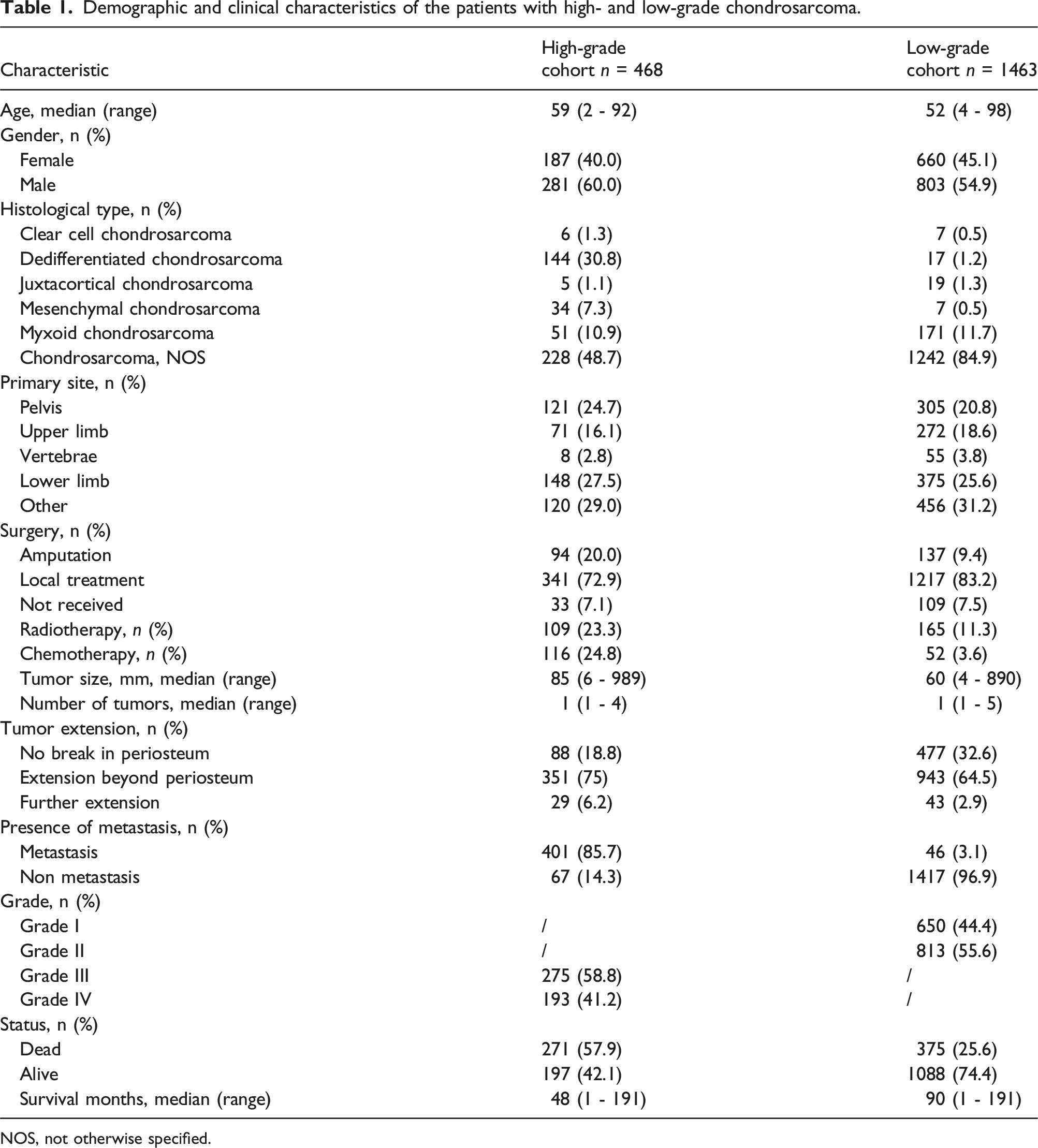
NOS, not otherwise specified.

Kaplan–Meier survival curves for the patients with different grade of chondrosarcoma. Log-rank test was performed to compare the difference between each pair of grade levels. (a) The difference between the 4 grades was significant. (b) Grade I and II were classified as low-grade chondrosarcoma. Grades III, and IV were classified as high-grade chondrosarcoma. The difference between the two categories was significant.
Cox Proportional Hazards Analysis of the effect of various characteristics on overall survival.

A p-value of <0.05 was considered statistically significant. HR, hazard ratio. CI, confidence interval. C-index, c-index on cox model of each variable. Ref, reference category. NOS, not otherwise specified.
Feature importance of various characteristics estimated by Random Survival Forest (RSF).

Will ensemble learning algorithms outperform the cox regression and single computational models in prognostic evaluation of high-grade chondrosarcoma?
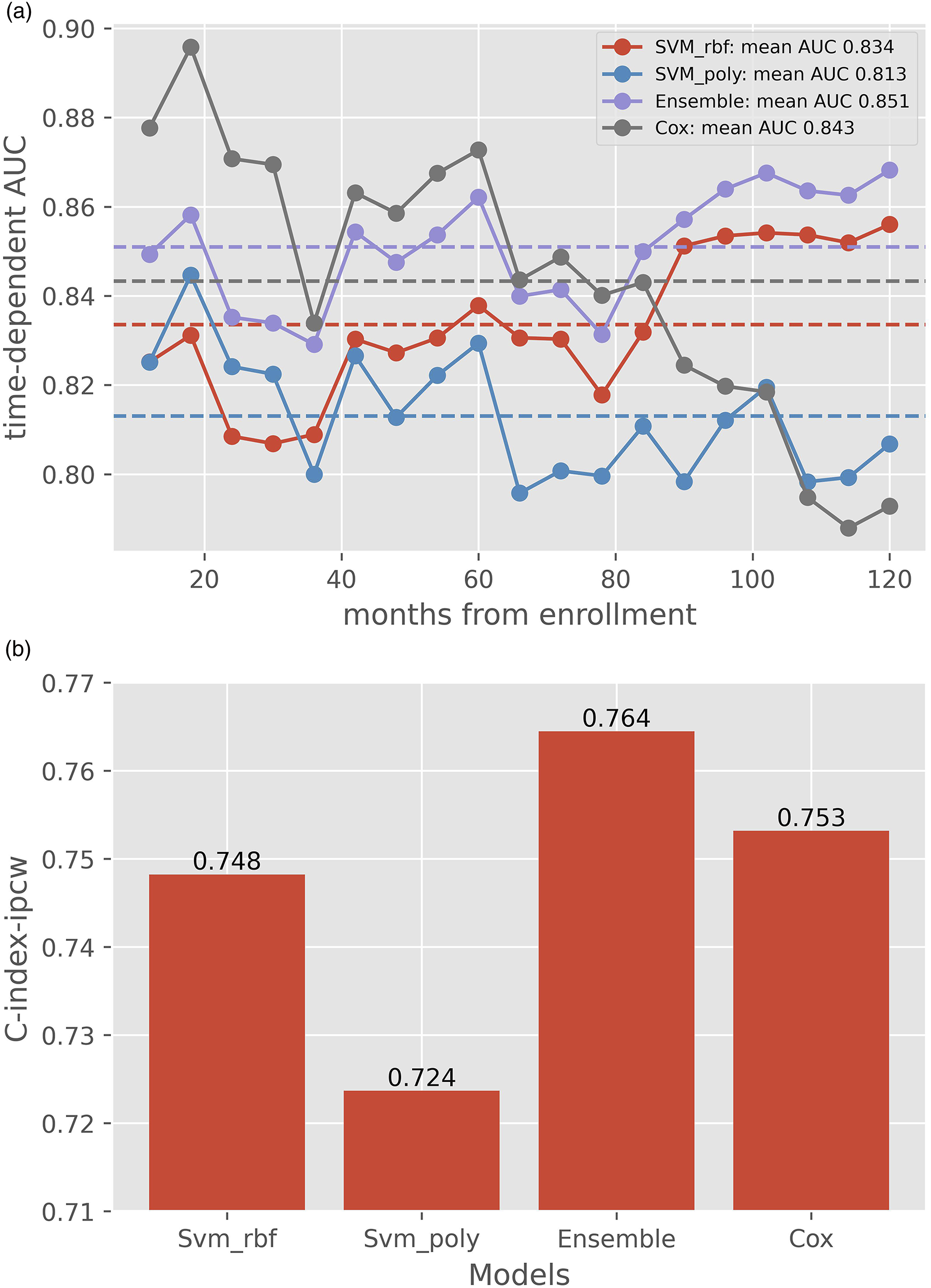
The scatter plot shows the survival data predicted case by case by each learning model and the real survival data from the test set, with the Pearson correlation coefficient of 0.640. (Figure 3). In Figure 4(A), AUC values at each time point are presented as the broken line. In general, AUC values of the ensemble learning model were above 0.83 at different follow-up times and were superior to those of the other two models, which showed that the ensemble learning model had far better accuracy than the other two machine learning models (Svm_rbf, support vector machine with radial basis function kernel, and Svm_poly, support vector machine with polynomial kernel) for survival prediction. Moreover, cox proportional hazard model seemed to perform better than the other three models at first few years but was lately surpassed by ensemble learning model after approximately 7 years. In Figure 4(A), time-dependent mean AUC values of each model are presented as the dotted line (0.851, 0.843, 0.834 and 0.813 for ensemble learning, CoxPH, Svm_rbf and Svm_poly model, respectively). The C-indexes, which indicates the performance of various machine learning models, were 0.764 for ensemble learning model, 0.748 for Svm_rbf model, 0.724 for Svm_poly model, and 0.753 for CoxPH model, respectively (Figure 4(B)). The ensemble learning model had better performance metrics than the other models. Survival data predicted by the ensemble model and the real data of the test set. The scatter plot was generated for comparing the survival time predicted by the ensemble model with the real data of the test set one by one regardless of the last follow-up status. Performance assessment of ensemble model, CoxPH model and the single models of Svm_rbf and Svm_poly (a) The area under the curve (AUC) values of the receiver operating characteristics (ROC) curve at each time point and its mean value. AUC values of ROC curve at each time point (from the 12th to the 120th month) were presented as the broken lines. Time-dependent mean AUC value of each model was presented as the dotted line. The ensemble learning model predicted the risk of survival status far better than the other two single machine models alone. At the same time, Cox model performed better at first but was surpassed in reverse after approximately 7 years. (b) C-index of the ensemble learning model compared with those of the Cox model and the single model of Svm_rbf and Svm_poly. Svm_rbf, support vector machine with radial basis function kernel; Svm_poly, support vector machine with polynomial kernel; Ensemble, ensemble learning; CoxPH, Cox proportional hazards.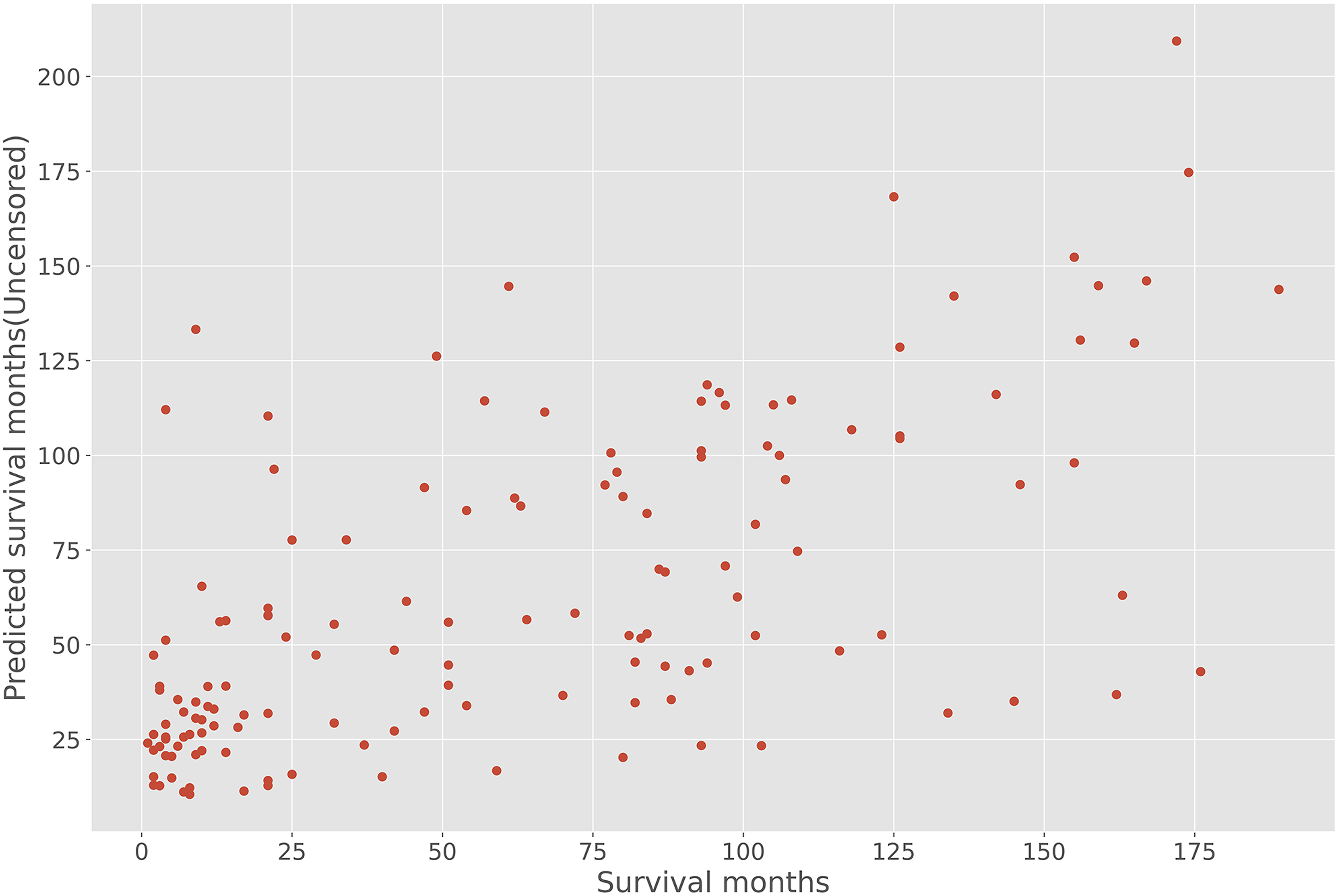
Among distinct age groups with specific factors, what kind of patients with high-grade chondrosarcoma is expected to benefit most from chemotherapy?
The ratio of the patients who receive chemotherapy or not with survival over 36 months.
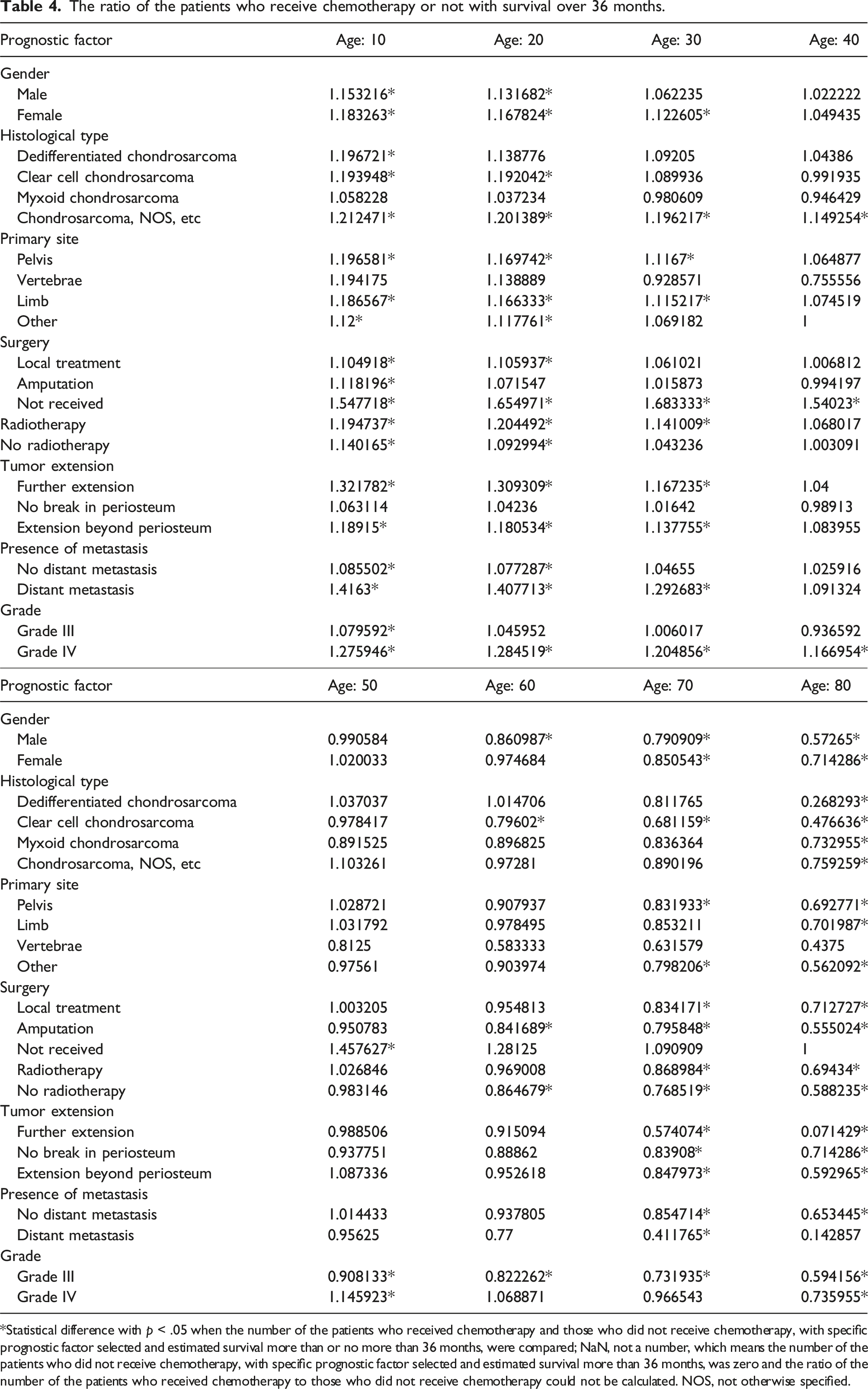
*Statistical difference with p < .05 when the number of the patients who received chemotherapy and those who did not receive chemotherapy, with specific prognostic factor selected and estimated survival more than or no more than 36 months, were compared; NaN, not a number, which means the number of the patients who did not receive chemotherapy, with specific prognostic factor selected and estimated survival more than 36 months, was zero and the ratio of the number of the patients who received chemotherapy to those who did not receive chemotherapy could not be calculated. NOS, not otherwise specified.
The 2 × 2 contingency table design for χ2 or Fisher exact test in each group in Table 4.

Furthermore, survival time of the virtual patients in subgroup A (Figure 5(A)), B (Figure 5(B)) and C (Figure 5(C)) with different tumor sizes ranging from 40 to 120 mm was analyzed by the ensemble learning model. In each age group with specific prognostic factors selected, we counted the number in subgroup A, B and C as a percentage of the total (Table 6). We noted that among the aforementioned factors that were identified to contribute to increased benefit from chemotherapy in Table 4, the factors leading to more than 25% difference in the percentage of the patients in subgroup A and C in Table 6 were: Dedifferentiated chondrosarcoma, amputation, local treatment, no distant metastasis, or grade III with the age no more than 10; Male, clear cell, primary site other, or no radiotherapy with the age no more than 20; Female, primary site at pelvis, primary site at limb, radiotherapy, extension beyond periosteum, further extension, or distant metastasis with the age no more than 30; Chondrosarcoma, NOS, etc (including mesenchymal, juxtacortical and classical chondrosarcoma) with the age no more than 40; No surgery received or grade IV with the age no more than 50. Under these factors, the benefit from chemotherapy was definite. Besides, tumor extension, no surgery received and grade IV tumor led to more than 40% differences in the percentage of patient number between subgroup A and C among all age groups, which indicates the decisive effect of these factors on evaluating chemotherapy benefit. Relationships between predicted survival time and tumor size of the patients in each subgroup (a) Subgroup A included the patients whose survival time was longer if they received chemotherapy, regardless of tumor size. (b) Subgroup B included the patients with specific tumor size intervals which can be referred to for choosing chemotherapy or not to obtain longer survival time. (c) Subgroup C included the patients with shorter predicted survival time if receiving chemotherapy, regardless of tumor size. The proportion of patients with extra or no benefit from chemotherapy in different age groups. aThe percentage of the patients with longer estimated survival if receiving chemotherapy in different age groups with selected prognostic factor, regardless of tumor size. bThe percentage of the patients with specific tumor size intervals in different age groups under selected prognostic factor, which can be referred to for choosing chemotherapy or not to get longer survival time. cThe percentage of the patients with shorter estimated survival if receiving chemotherapy in different age groups with selected prognostic factor, regardless of tumor size.
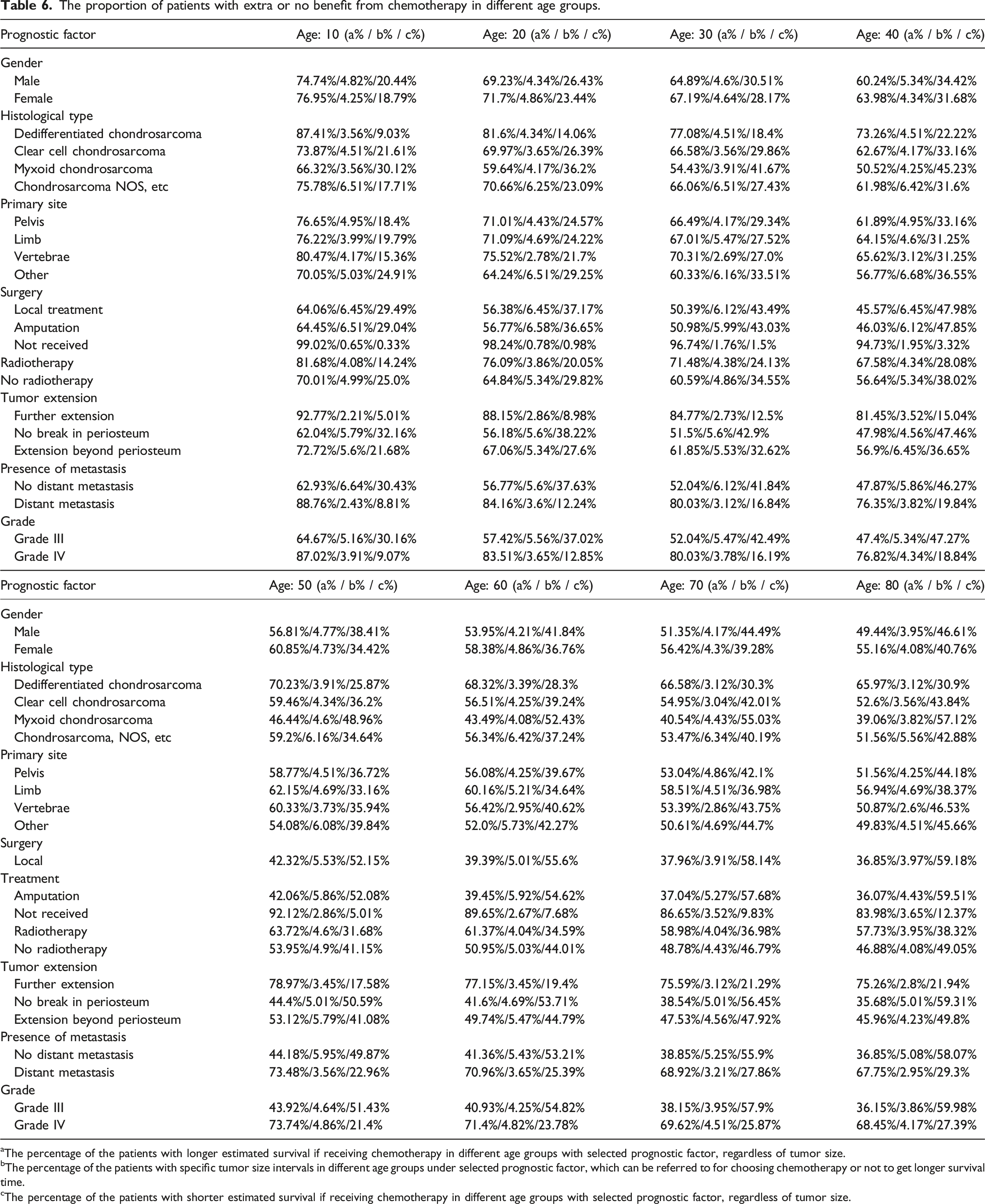
Specific factors with adverse prognostic outcomes after chemotherapy in different age groups were also identified. These factors were: Grade III with the age over 50; Amputation with the age over 60; Local surgical treatment, no break in periosteum, extension beyond periosteum, or no distant metastasis with the age over 70; myxoid chondrosarcoma or no radiotherapy with the age over 80. These factors led to statistical difference of χ2 or Fisher exact test with the ratio <1 in Table 4 and were also associated with opposite difference in the percentage of the patients in subgroup A and C (subgroup A% < subgroup C%) in Table 6.
Discussion
To the best of our knowledge, this study represents the first attempt to combine ensemble learning methods with multiple prognostic factors to predict survival and evaluate the efficacy of chemotherapy. In this study, the statistical difference among the estimated survival curves of the patients with different grades of chondrosarcoma further validated the poor prognosis of high-grade chondrosarcoma. CoxPH analysis and the RSF method were used to identify potential variables associated with the prognosis of high-grade chondrosarcoma. Survival support vector machine with different kernel methods were utilized for training purposes. We successfully developed an ensemble-learning based model for survival prediction in patients with high-grade chondrosarcoma.
Ensemble learning algorithms outperform CoxPH and single computational models in prognostic evaluation of high-grade chondrosarcoma
Computational approaches such as machine learning have made significant contributions to predicting metastasis, drug response, survival and recurrence rate in the field of clinical oncology, especially for chondrosarcoma. However, the limitations of single machine learning models are evident.18,19 In our study, the ensemble learning model outperformed the other models when dealing with large samples and multiple variables, followed by support vector machines with polynomial kernel, support vector machine with radial basis function kernel, and CoxPH. A 1000-repeated random search with cross-validation for hyperparameter tuning was conducted to obtain the best model configuration with stability and accuracy.
Previous studies have focused mainly on nomogram models based on CoxPH analysis of the SEER data of various bone sarcomas. The c-indexes for model evaluation in these studies were lower than those in the studies of machine learning models (including this study).28–31 Moreover, the c-index of the ensemble model in our study was significantly better than those reported for The American Joint Committee on Cancer (AJCC) staging system.28,32,33 Thio et al. developed the machine learning -based Skeletal Oncology Research Group (SORG) algorithm and analyzed the SEER data to predict the 5-year survival of the patients who were surgically treated. 18 The c-index of moderately and poorly differentiated tumors (sorting of tumor grade) reached 0.74 27. The commonly used c-index for discriminative performance was considered inappropriate when predicting the risk for the time-to-event result, due to a higher c-index of mis-specified model under a defined time interval.32,34 Thus, time-dependent AUC was also utilized for describing time-dependent specificities and sensitivities at all time points. The AUC value of the ensemble model in this study was over 0.85 (highly reliable), which is higher than those of the reported nomograms from the SEER database.30–32,35 In addition, the ensemble model demonstrated no inferior AUC value when compared to various computational models. 36 Sung et al. used neural network machine learning algorithms to analyze the role and outcomes of surgical resection and radiation therapy in spino-pelvic chondrosarcoma with the mean AUC reached 0.84 19. The SORG algorithm model showed good discriminative ability and overall performance when applied in the SEER derivation cohort and external validation cohort, with the AUC for 5-year survival around 0.85.18,27
The model in our study demonstrated superiority in terms of c-index and mean AUC over those of previous studies. Moreover, our study model reflected a larger and more recent collection of patients from numerous centers available via the SEER database. There was significant superiority in our performance metrics favoring the ensemble learning model over the other three single algorithms for survival prediction. Therefore, by using various optimized algorithms to fit time-to-event data, the ensemble learning model becomes more accurate and flexible when handling complex and nonlinear data.
Identification of distinct age groups with specific factors of chondrosarcoma for tailored use of chemotherapy
A previous study comparing the tumor features revealed significant differences in median survival between chondrosarcoma subtypes, with the highest median survival in the juxtacortical subtype (97 months), followed by clear cell (79 months), myxoid (60 months), and mesenchymal subtypes (33.5 months), and the lowest in dedifferentiated subtype (11 months). 11 The rate of metastasis emerged as the only prognostic variable for decreased long-term survival and differed significantly with 2.1% in juxtacortical, 5.7 % in clear cell, 7.6% in myxoid, 10.6% in mesenchymal, and 19.8% in dedifferentiated subtype. 11 As the median OS for advanced chondrosarcoma with a higher grade of malignancy was reported as poor as 18 months and the survival rate at 5 years was very low,6,8,9 the time point for survival prediction would be appropriate between 18 and 60 months. Although the performance superiority of the ensemble learning model over traditional models in real datasets appeared after 36 months in this study, we adjusted the imbalanced number of virtual patients in several groups in Table 4 for reaching comparable sample size. Some clinical features with fewer patients in real datasets could be examined individually, considering that these patients have shorter survival times. Therefore, the survival prediction based on the ensemble learning model was judged based on a 36-month period in this study.
Distant disease is the dominant mode of treatment failure in dedifferentiated subtype.3,4 Whether chemotherapy should be used for dedifferentiated chondrosarcoma has been one of the controversies in the medical field. Italiano et al. reported that conventional chemotherapy had relatively better efficacy for patients with advanced mesenchymal and dedifferentiated chondrosarcoma than for those with other advanced subtypes, although the benefit was limited. 8 In this study, the benefit of chemotherapy over no chemotherapy for dedifferentiated chondrosarcoma was significant only when patients were as young as 10 years old (Tables 4 and 6). Such benefit was limited with no statistical significance in other age groups (from 10 to 50 years old). Consistent with the finding of previous studies, our study confirmed the efficacy of age-adapted chemotherapy for treating dedifferentiated chondrosarcoma with aggressive entity.
It is a well-established fact that the only definitive treatment for chondrosarcoma is wide resection. Conventional surgery for chondrosarcoma still subjects the limitation of treatment failure given the presence of distant metastasis and incomplete resection.14,17 Previous studies evaluating the efficacy of neoadjuvant or adjuvant chemotherapy for high-grade tumors have yielded controversial conclusions regarding improvements in progress-free survival and OS. Two separate studies from Cranmer et al. and Liu et al. both revealed no definitive improvement in survival associated with primary chemotherapy in treating high-grade tumors, such as dedifferentiated chondrosarcoma.1,37 Li et al. suggested that chemotherapy was even a risk factor for patient prognosis. 36 On the other hand, some studies demonstrated that aggressive administration of chemotherapy led to favorable results and contributed to the achievement of a surgical complete remission which was considered the crucial factor associated with prolonged OS.5,13,38,39 The specific clinicopathologic or treatment factors, such as primary disease arising in the context of osteochondroma or a large osteo-sarcomatous component, were found meaningful in identifying the patients who would gain benefit from neoadjuvant or adjuvant chemotherapy.13,39 In our study (Tables 4 and 6), the benefit of chemotherapy over no chemotherapy in the patient who underwent surgery was limited with the age no more than 10 years old. The variations in the results of perioperative chemotherapy on high-grade chondrosarcoma is likely caused by differences in the inclusion criteria (such as the inclusion of the cases with large tumor size or extensive metastasis when chemotherapy has to be chosen), the innate limitations of the retrospective or noncomparative studies and small sample size.11,13,14,17,36
Previous studies have demonstrated that the patients would survive longer after surgical resection, even with metastatic condition.15,36,40 In our study, we found that among the patients who underwent surgical treatment and had an estimated survival of more than 36 months, the number of those who received chemotherapy was greater when the age was no more than 50 years old. Thus, it is suggested that every effort, including perioperative chemotherapy, should be made to perform surgical resection for younger patients and prevent transformation into high-grade disease when feasible. Perioperative administration of chemotherapy in osteosarcoma, another high-risk bone malignancy, has shown encouraging outcomes and has been recommended by various management guidelines.4,10,13 Given the frequent existence of an osteo-sarcomatous component in high-grade chondrosarcoma and the osteosarcoma treatment protocols being proposed as a model for dedifferentiated chondrosarcoma treatment, it is reasonable to consider perioperative chemotherapy in other chondrosarcoma subtypes under careful consideration, or a clinical trial.1,9,10,39
Therapeutic plans based on calculated hazard ratio (HR) value of the classical CoxPH model are usually constant owing to its linearity assumption of variable fitting.9,14,41 Conventional cytotoxic agents have limited effects on advanced chondrosarcomas with higher tumor grades, whereas more active systemic therapies are considered probably meaningful in these cases stratified by age.2,3,41 Studies have identified age, tumor size and histological grade as prognostic factors which significantly associated with survival.2,7,12 However, no evidence-based prediction exists for the definite association between these factors and survival after receiving chemotherapy or not. We have generated estimated OS value for patients in the chemotherapy and non-chemotherapy group with various age distributions and tumor size, for reference in study design and statistical power calculations. In our study, grade IV tumor in younger patients (aged no more than 50 years) tended to benefit more from chemotherapy, while such benefits were not significant in older patients. Notably, chondrosarcoma is more common in older patients who may not tolerate more radical treatment regimens due to poor physical status.2,10 Understanding these treatment differences among different patient groups will positively affect risk‐benefit decisions for appropriate treatment recommendations. Furthermore, the model will also provide survival guidance and benchmarks for stratification in future clinical trials.
This study has several limitations. First, the inherent limitations of the SEER database should be mentioned when considering the potential confounding variables. Although the patients in this study have better reflected the actual diversity with complete and comprehensive incidence and survival data, most were classified as “Chondrosarcoma, NOS, etc.” (accounting for 48.7% and 84.9% of cases in the grade III/IV and grade I/II cohorts, respectively). SEER does not provide information about the chemotherapy administered, dose intensity, length of therapy, or other treatments (such as surgery and radiotherapy). 25 SEER does not provide a clear distinction between unknown treatment status and nonreceipt of chemotherapy. Consequently, patients who received chemotherapy but were classified in the no/unknown status group might inadvertently downplay the impact of chemotherapy treatment. Second, pathology results in the SEER database rely on the confirmation from participating institutions instead of the central review. It is possible that some cases may have been pathologically misclassified in the database, necessitating caution when interpreting the results. Third, the algorithm developed in this study requires validation with external datasets to assess its generalizability and reliability. Adequate validation with multi-institution cohorts is encouraged to improve the accuracy of the model.
Conclusion
The ensemble learning algorithm demonstrated outstanding performance for prognostic assessment of chemotherapy benefit in high-grade chondrosarcoma, particularly within specific age groups under specific factors. However, the findings must be viewed cautiously, given the substantial limitations of the SEER database and the lack of clinical validation of the prediction model. Future studies focusing on the identification of patient subsets that are more likely to benefit from chemotherapy are desired to improve the practicability of the prediction model.
Supplemental Material
Supplemental Material - Ensemble learning guided survival prediction and chemotherapy benefit analysis in high-grade chondrosarcoma: A study based on the surveillance, epidemiology, and end results (SEER) database
Supplemental Material for Ensemble learning guided survival prediction and chemotherapy benefit analysis in high-grade chondrosarcoma: A study based on the surveillance, epidemiology, and end results (SEER) database by Xu Zheng, Longqiang Shu, Shanyi Lin, Hanqiang Jin, Xiaoyu Wang and Ting Yuan in Journal of Orthopaedic Surgery
Footnotes
Authors’ contributions
XZ and XYW conceived and designed the study. XZ, LQS and XYW collected the data and contributed to image analysis. XZ, HQJ and XYW analyzed the data and drafted the manuscript. SYL, HQJ, XYW, and TY offered administrative, technical, and/or material support. All authors contributed to the manuscript revision and approved the final version of the manuscript. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Data Availability Statement
The datasets supporting the conclusions of this article are available in the Surveillance, Epidemiology, and End Results cancer registry (![]() ) and are also available from the corresponding author on reasonable request. All data generated or analyzed during this study are included in this published article and its Supplemental Material.
) and are also available from the corresponding author on reasonable request. All data generated or analyzed during this study are included in this published article and its Supplemental Material.
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
