Abstract
Epstein–Barr virus is an etiologic agent of several malignancies. In this study, we explored the association of Epstein–Barr virus–encoded RNA1 and Epstein–Barr virus latent membrane protein 1 co-expression with osteosarcoma. Epstein–Barr virus–encoded RNA1 expression in tumor cells was quantified using reverse transcriptase polymerase chain reaction and in situ hybridization and Epstein–Barr virus latent membrane protein 1 expression was measured using immunohistochemistry staining. There was a statistically significant association between Epstein–Barr virus latent membrane protein 1 and Epstein–Barr virus–encoded RNA1 co-expression and characteristics of osteosarcoma such as nodal stage (p < 0.04), metastasis (p < 0.04), Ki67 index (p < 0.03), and tumor stage (p < 0.05). Co-expression of Epstein–Barr virus–encoded RNA1 and Epstein–Barr virus latent membrane protein 1 in tumors correlated with advanced osteosarcoma and indicated the aggressiveness of bone sarcoma.
Keywords
Introduction
Though there is no clearly defined etiology in most cases of bone sarcoma, some associated or predisposing factors have been identified. 1 In many cases, an association between viral infection and sarcoma has been reported, for example, between HIV/human herpes virus 8 and Kaposi’s sarcoma and between Epstein–Barr virus (EBV) and smooth muscle tumors. EBV, which belongs to the subfamily Gammaherpesvirinae, has been identified as the etiologic agent of several malignancies that are dependent on the expression of pro-angiogenic factors. EBV is a ubiquitous γ1 herpes virus that infects approximately 95% of the global population. EBV is associated with several malignancies, including B-cell lymphoma, T-cell lymphoma, natural killer cell lymphoma, post-transplant lymphoproliferative disease (PTLD), endemic Burkitt’s lymphoma, a subset of Hodgkin’s lymphomas, epithelial malignancies, nasopharyngeal carcinoma, and gastric carcinoma. 2
EBV induces different latency programs in infected cells through a viral episome. EBV transforms resting B-cells into immortalized lymphoblastoid cell lines that express several viral genes, including all six Epstein–Barr nuclear antigens (EBNAs), latent membrane protein (LMP) 1, LMP2A, and 2B, and EBV-encoded RNAs (EBERs) 1 and 2. This type of latency, known as latency III, promotes the proliferation of infected B-cells, allowing the replication of not only the host genome but also the viral episome. Latency III is associated with PTLD and diffuses large B-cell lymphomas. 3 Latency II is a less immunogenic latency program that involves expression of EBNA1, LMP1, LMP2A and 2B, and EBERs. Latency II is seen in EBV-induced epithelial cancers, nasopharyngeal cancers, and gastric carcinomas. Memory B-cells are the reservoir of EBV, which induces two distinct latency programs (latency 0 and latency I) in these cells. In latency 0, EBERs are expressed. Burkitt’s lymphoma shows a latency I expression profile. A specific latency pattern is not required for the induction of cell transformation by EBV because malignant cells show different gene expression profiles. 3 Several previous studies have explored the role of EBV proteins and noncoding RNAs in inducing pro-angiogenic processes that enhance tumorigenesis.4–8 LMP1 is a major oncogenic protein of EBV and is expressed in most EBV-associated cancers. 9 LMP1, being a constitutively active member of the tumor necrosis factor receptor family, can activate signaling pathways including the NF-κB, PI3K/Akt, MAPK/ERK, JAK/STAT, and c-Jun N-terminal kinase pathways.10–13 Activation of these pathways modulates apoptosis, cell migration, cell cycle progression, and angiogenesis. LMP1 activates multiple signaling pathways to induce vascular endothelial growth factor (VEGF), which is associated with poor prognosis and more aggressive disease. 14 Furthermore, LMP1 induces several growth factor receptors, such as pro-angiogenic factors and epidermal growth factor receptors, which play a role in the malignant transformation of tumor cells and contribute to the promotion of angiogenesis.15–17 LMP1 also induces the expression of pro-migratory factors such as Twist, which increases cell migration, facilitates angiogenesis, and drives epithelial–mesenchymal transition (EMT). 18 EBERs are noncoding RNAs that are expressed in latently EBV-infected cells and are useful for the detection of EBV-infected tissues. 19 Previous studies have demonstrated that EBERs are involved in preventing apoptosis while promoting cell proliferation, tumor growth, and possible angiogenesis. Therefore, EBERs contribute to oncogenesis.20,21
Angiogenesis is one of the hallmarks of cancer, and the ability of a tumor to hijack this process for its benefit facilitates tumor growth and enhances its ability to metastasize. 22 EBV is linked to several malignancies; importantly, EBV is known to have a pro-angiogenic activity that enhances tumorigenesis. Moreover, it is already known that several viral factors of EBV, in particular, play a direct and substantial role in modulating pro-angiogenic processes; therefore, many clinical trials have been conducted using angiogenesis inhibitors. Both latent and lytic proteins are involved in angiogenesis, supporting the hypothesis that paracrine signaling is vital for viral-induced malignancies. However, in the context of EBV tumorigenesis, the latent protein LMP1 appears to be the major pro-angiogenic factor. 21 However, stable EBV infection associated with the expression of latent EBV genes might transform mesenchymal cells into cancerous cells through multiple pathways. Furthermore, we described that a latency III expression profile (LMP1 and EBERs) of EBV infection might induce bone sarcoma development, which might contribute to EBV-induced osteosarcoma (OS). To the best of our knowledge, this is the first study to investigate the possible role of EBV in OS development.
Methods
Participants
A total of 48 primary OS samples were included in this retrospective cohort study. Formalin-fixed and paraffin-embedded (FFPE) tumor resection specimens, obtained between May 2012 and December 2018, were provided by two pathology laboratories of the Kermanshah and Iran Universities of Medical Sciences, Iran. The study protocol was approved by the Ethics Committee of the Kermanshah University of Medical Sciences (Imam Reza Hospital, KUMS.REC. 91103). Written informed consent was obtained from all patients and their guardians (parents).
Sampling and data collection
We identified 48 acceptable primary OS. Of those, 15 derived from new cases, while 17 and 16 from patients who were identified with recurrence and metastases, respectively. For this study, we used the samples of primary tumor, collected at first diagnosis.
The inclusion criteria were availability for the patients associated with optimal surgical specimens for pathological assessment and immunohistochemical (IHC) study. Immunocompromised patients, patients with chronic diseases, patients with other malignancy, patients exposed to neoadjuvant chemotherapy, radiation therapy, and hormone therapy before surgery, patients with incomplete data, and unsuitable pathology paraffin blocks were excluded. A total of 58 FFPE samples were included in this study; the case group consisted of 48 OS samples and the control group consisted of 10 non-cancerous bone tumor samples. The diagnosis of OS was confirmed by histopathology patterns and IHC study (positive for SATB2, S100 protein, SMA, CD99, and osteocalcin). All specimens were re-analyzed independently by two pathologists. Demographic and clinicopathologic data, including age, sex, tumor size, histological type, tumor grade, and tumor stage, were obtained from the medical records of the patients. All patients were Iranian, with a mean age of 17.43 ± 4.12 years. Non-cancerous bone tumor specimens were collected from non-pathologic fractures and classified as healthy controls. All tests were performed by the same trained laboratory personnel.
Reverse transcriptase polymerase chain reaction (RT-PCR) for EBER1
RNA preparation
The FFPE tissues were sectioned (10 μm) and treated with 1-mL xylene in sterile 1.5 mL tubes to remove paraffin from the sectioned specimens. Ethanol was added to remove xylene, followed by the addition of homogenizing buffer (75 μL), 10% sodium dodecyl sulfate (SDS, 15 μL), and proteinase K (9 μL) and overnight incubation at 55 C. The tissues were then treated with 1 mL TRIzol reagent and total RNA was extracted following the manufacturer’s instructions. Briefly, 200 μL of chloroform was added to the samples and centrifuged at 14,000 r/min for 12 min. An equal volume of isopropanol was added to the aqueous phase of the samples in a fresh tube to precipitate RNA, incubated for 10 min at 37 C, and then centrifuged again at 14,000 r/min for 12 min. The precipitated RNA was washed with 75% ethanol in diethylpyrocarbonate (DEPC) water and then dissolved in DEPC water.
RT-PCR procedure
Extracted RNA was quantified using an infinite M200PRO microplate reader (TECAN) and cDNA was synthesized using 2 μg of the extracted RNA. Tenfold-diluted cDNA was used for PCR. The thermal cycling conditions were as follows: denaturation at 95 C for 5 min; followed by 30 cycles of amplification at 90 C for 30 s, 50 C for 30 s, and 70 C for 30 s; and a final extension at 72 C for 5 min. The amplified product was resolved on a 2% agarose gel, stained with ethidium bromide (0.5 μg/mL) for 15 min, and photographed using a GelDoc Image Analyzer (Bio-Rad Hercules, CA, USA). The sequences of the primers were as follows: 5′-GATCCAAACTTTAGTTTTAG-3′ (EBER1 forward) and 5′-GCGAACCGTAACTCTATAC-3′ (EBER1 reverse). 23
In situ hybridization (ISH) for EBER1
ISH detects EBER1, which is expressed during the latent phase of EBV infection, on histological sections. EBER expression was detected using a 20-bp double digoxigenin-labeled (5′-end) oligonucleotide probe (antisense), 5′-CTACAGCCACACACGTCTCC-3′, designed by Primer 5.0 software (GeneBank gi|16326314|AB065135.1, human herpes virus 4 genes for EBER1 small RNA) and synthesized by Takara Biotech Co. (Dalian, China). ISH performed according to the protocol of Boster Biotechnology Co. (Wuhan, China). The FFPE sections were mounted on 3-aminopropyltriethoxysilane (APES)-treated glass slides, deparaffinized, hydrated, predigested with pepsin (3%) for 5–10 min, and hybridized with 2 ng/μL of a probe for 14–16 h. The slides were washed sequentially with 2×, 0.5×, and 0.2× silver sulfadiazine Crème (SSD) for 15 min at 37 C, blocked with bovine serum albumin for 30 min at 37 C, treated with biotinylated anti-digoxigenin rabbit antibodies for 60 min at 37 C, and washed four times with 0.5 M phosphate-buffered saline (PBS) for 5 min. Next, streptavidin–biotin complex (SABC) was added to the slides and incubated at 37 C for 20 min followed by the addition of biotin-peroxidase and incubation at 37 C for 20 min. The sections were then washed four times with 0.5-M PBS for 5 min, dyed with diaminobenzidine (DAB) for 10 min, and counter-stained with hematoxylin for 8 min. 24 Two tonsil sections from a case of EBV-positive infectious mononucleosis were used as positive controls. Two slides prepared without the probe were used as negative controls. A dark brown nuclear stain indicated positive EBER1-ISH (Figure 1).

(a)–(d) Histopathological images of Epstein–Barr virus-encoded small RNA1 (EBER1) tumor cells of osteosarcoma (OS), stained with (a) hematoxylin and eosin (H&E) and (b) EBER1 detection by in situ hybridization (ISH) and of EBER1-negative OS, stained with (c) H&E and (d) EBER1 detection by ISH. (Each panel represents 40× objective of sections and 50 μm magnification bar.)
IHC staining of LMP1
IHC was performed on 6 μm sections of the tissue microarray (TMA). All sections were transferred to an adhesive-coated slide system (Instrumedics, Hackensack, NJ, USA), deparaffinized, and processed using an automated Ventana Benchmark staining system (Ventana Medical Systems, Tucson, AZ, USA). For IHC staining of LMP1, 1:100 diluted anti-LMP1 (Dako, Carpinteria, CA, USA) was used. Immunobinding of anti-LMP1 was visualized using biotin-labeled secondary antibodies and SABC with DAB as the chromogenic substrate (DAB chromogenic kit, ScyTek). The sections were counter-stained with hematoxylin. EBV-positive Hodgkin’s lymphoma sections, which are known to be immunoreactive for LMP1, were used as positive controls. For negative control, the primary antibody was replaced with an antibody diluent. 25 LMP1 expression was localized and quantitated by microscopic evaluation of a cytoplasmic or membranous pattern of staining. LMP1 expression was considered positive when more than 10% of the neoplastic cells showed a cytoplasmic or membranous pattern of LMP1 staining deemed to be positive (Figure 2).
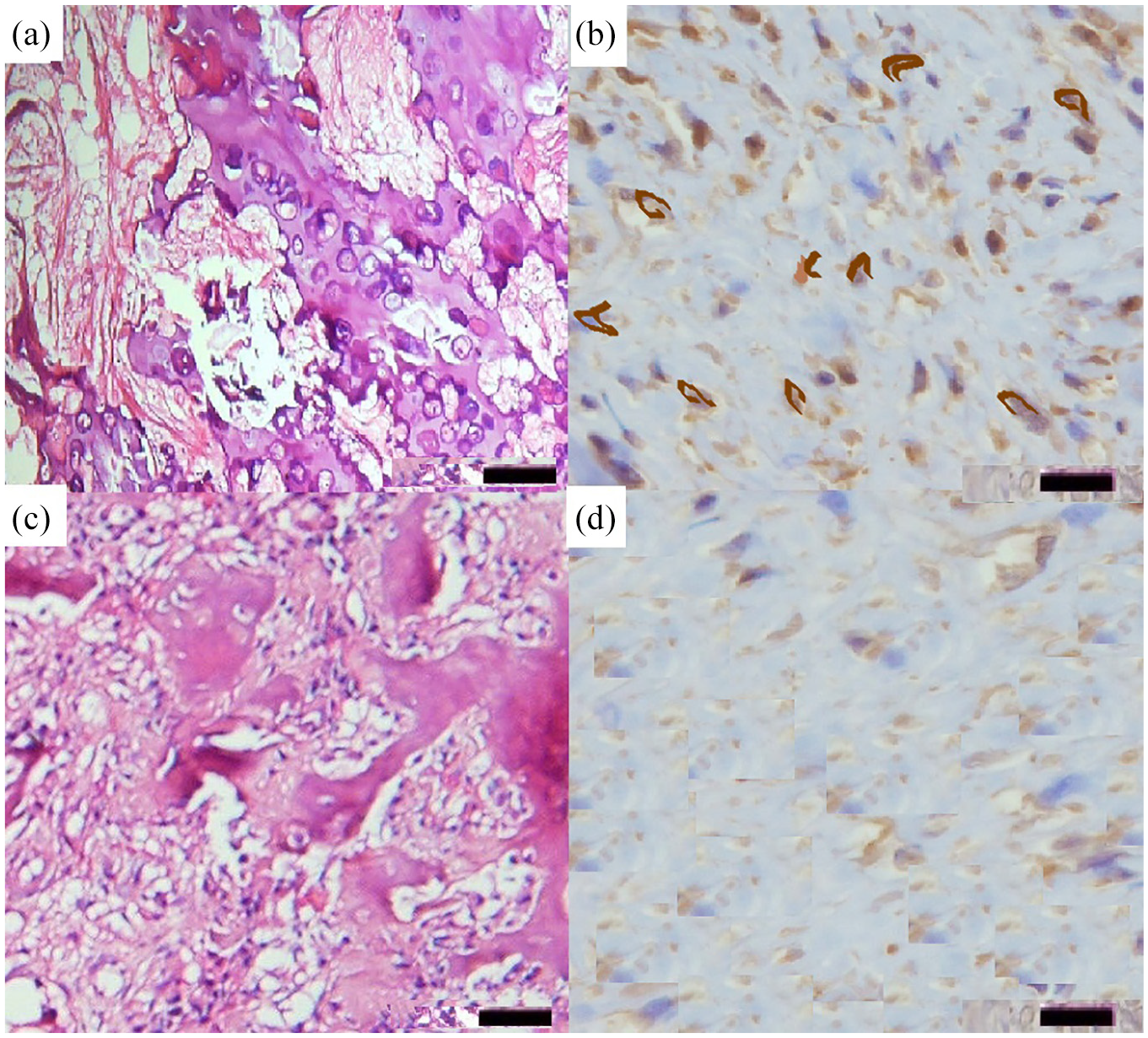
(a)-(d) Histopathological images of Epstein–Barr virus latent membrane protein 1 (LMP1) tumor cells of osteosarcoma (OS), stained with (a) hematoxylin and eosin (H&E) and (b) LMP1 detection by Immunohistochemistry (IHC) and of LMP1-negative OS, stained with (c) H&E and (d) LMP1 detection by IHC. (Each panel represents 40× objective of sections and 50 μm magnification bar.)
Statistical analysis
All statistical analyses were performed using SPSS 16.0 statistical package software (SPSS, Chicago, IL, USA) or GraphPad Prism (Graph Pad Software, Inc., San Diego, CA). All data were qualitative and presented as numbers or percentages; data were compared using the chi-square test and Fisher’s exact test. A p-value of 0.05 was considered indicative of a statistically significant difference. Overall survival (O-S) was defined as the time from the date of initial diagnosis to the date of death from any cause or last follow-up. Disease free survival (DFS) was defined as the time from the date of curative resection to the date when recurrence or metastasis was confirmed, death from any cause, or last follow-up. Progressive free survival (PFS) was defined as the time from the date of palliative treatment to the date when disease progression was confirmed, death from any cause, or last follow-up. O-S, DFS, and PFS were calculated using the Kaplan–Meier (K–M) method, and the 95% confidence intervals (CIs) were reported. DFS is defined as the period between the primary surgical treatment and the date of relapse.
Results
Among the 48 OS patients evaluated, 33 (69%) were boys, and 15 (31%) were girls (ratio of boys to girls was 2.2:1). The mean age of the subjects was 17.43 ± 4.12 years (range: 7.5–25 years). Histologically, subjects diagnosed with OS were at different stages of the disease. According to World Health Organization guidelines, 15 (31%), 17 (36%), and 16 (33%) of the biopsies were classified as stages I, II, and III, respectively. The clinicopathologic characteristics of EBV were analyzed using three different laboratory methods (Table 1).
The clinicopathologic characteristics of EBV, using three different laboratory methods (RT-PCR).

EBER: EBV-encoded RNAs; ISH: in situ hybridization; LMP: latent membrane protein; IHC: immunohistochemistry; RT-PCR: reverse transcriptase PCR.
EBER1 was detected by RT-PCR in 9 out of 48 (18.8%) OS specimens but never detected in the control group (p < 0.14). Using ISH, EBER1 expression was detected in 6 out of 48 (12.5%) OS specimens, while it was not detected in the control group (p < 0.09). LMP1 expression was detected by IHC staining in 7 out of 48 (14.6%) OS specimens and 2 out of 10 control samples (p < 0.88); among the 7 LMP1-positive samples, 6 were EBER1-positive. Therefore, 6 out of 48 (12.5%) OS cases showed co-expression of EBER1 and LMP1 in tumor cells (p < 0.02; Figure 3).

A strong correlation between LMP1 and EBER1 expression in osteosarcoma was found in our study.
Among the six positive cases, two presented stage II (33.5%) and four presented stage III (66.5%) disease. The associations between EBER1 and LMP1 co-expression and histologic characteristics of OS specimens such as nodal involvement (p < 0.04), metastasis (p < 0.04), Ki67 reactivity (p < 0.03), and tumor stage (p < 0.05) were statistically significant. Although, there were no significant associations between EBER1 and LMP1 co-expression with two age groups (p < 0.27) and sex (p < 0.91), but patients who co-expressed EBER1 and LMP1 in tumor cells (EBV-positive) were elder male and showed worse clinical symptoms and poor prognosis as compared to patients who did not co-express EBER1 and LMP1 in tumor cells with co-expression. No significant association was found between EBER1 and LMP1 co-expression and tumor size (p < 0.53), tumor site (p < 0.89), bone tumor location (p < 0.75), histologic type (p < 0.91, grade (p < 0.42), and also vascular invasion (p < 0.57) of tumor. The mean follow-up period of surviving patients was 45 ± 12 months (range: 30–60 months). O-S (Figure 4(a)) and PFS (Figure 4(b)) were significantly worse for EBER1 and LMP1 positive patients compared to negative ones (O-S: 54% versus 85% at 5 years, p < 0.05; PFS: 41% versus 81% at 5 years, p < 0.04). Moreover, EBER1 and LMP1 positive patients with nodal involvement (Figure 5(a)), evidence of metastasis (Figure 5(b)), high Ki67 index in tumor (Figure 5(c)), and finally advanced stages of disease (stage II or III; Figure 5(d)) showed significantly shorter DFS as compared to EBER1 and LMP1 negative patients. These results suggest that co-expression of EBER1 and LMP1 in tumor cells might be associated with poor prognosis.
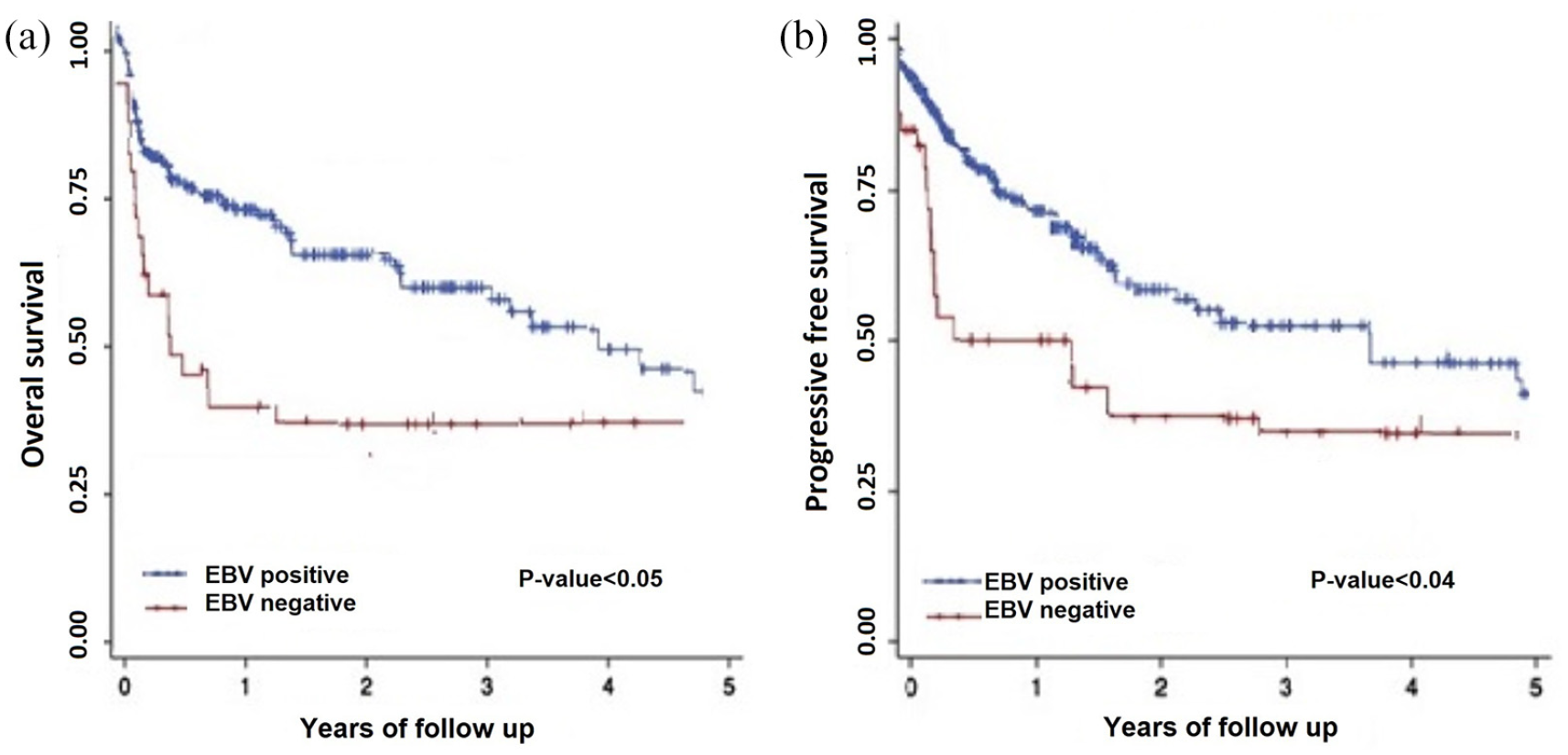
(a) and (b) Kaplan–Meier estimates of overall survival (O-S) and progression-free survival (PFS). The EBV-positive patients (n = 6) had significantly worse (a) O-S and (b) PFS than the negative ones (n = 42).

(a)-(d) Kaplan–Meier survival curves of disease-free survival: EBER1 and LMP1 positive patients with (a) nodal involvement, (b) metastasis, (c) high Ki67 index, and finally (d) the advanced stages (stages II or III) showed significantly shorter DFS as compared to negative ones.
Discussion
It is estimated that up to 18% of the cancers are associated with viral infections; and EBV contributes to 1% of all cancers. 26 In this study, we investigated whether EBV DNA is detectable in tumor resection specimens of patients with OS. Although EBV contributes to the oncogenesis of a considerable number of malignant tumors, its involvement in the pathogenesis of OS has not been detected in our study and the cause of OS is unknown. However, the predisposing factors for OS include age, male sex, and previous radiation therapy. 27 EBV is associated with a wide range of human cancers originating from epithelial cells, lymphocytes, and mesenchymal cells. The development of EBV-associated cancer is largely dependent on environmental and genetic factors. 28 Our findings demonstrate that EBER1 and LMP1 co-expression might be potential early biomarkers of OS in Iranian patients and are probably associated with tumor grade and stage. In contrast to our observation, Mizobuchiet et al. 29 found no significant association between esophageal carcinoma and EBV infection. 30 Consistently, two other studies revealed no association between EBV infection and esophageal carcinoma in Japanese and Northern Chinese patients. 31 Also, Torabizadeh et al. 32 observed no etiologic association between EBV infection and invasive ductal carcinoma of the female breast in Iranian patients. Moore et al. reported that EBV-related smooth muscle neoplasms have been associated with immune dysregulation and aggressive tumor behavior, most notably in immunocompromised/Immunosuppressed patients. While chemotherapy may slow down disease progression, resection, and improve the host immune status, providing the best opportunity for primary tumor control. 33 Recently, Salehzadeh et al. 34 demonstrated the presence of EBV in 30 Iranian patients with esophageal squamous cell carcinoma and its potential link with tumor size. These different results could be related to various factors such as environmental factors, the geographic distribution of EBV infections, genetic susceptibility, and diverse standardized detection methods. In agreement with our study, Moradi et al. reported an association of tumor size, node, metastasis status, and stage of esophageal cancer with EBV infection in 108 Iranian patients. Also, they observed shorter overall and progression-free survival in patients with EBV-associated esophageal cancer. 35 Gala et al. 36 investigated the role of EBV as an etiologic agent in various lymphomas. They studied 184 cases of lymphomas using a standard IHC panel and ISH for EBER1 expression. The prevalence of EBV-associated lymphoma was similar to that reported in Western literature. Our findings suggest that EBER1 and LMP1 co-expression may be a valuable biomarker in predicting the prognosis of patients with OS.
Limitations
By the small sample size and geographic limitation of the study population, the results still need to be supported by an enlarged sample size in other geographic regions.
In conclusion, our study showed that these patients with co-expression of EBER1 and LMP1 within the OS microenvironment presented advanced clinicopathologic features and tended to have shorter survival. Therefore, our study suggested opportunities for diagnosis of treatable stages of OS associated with EBER1 and LMP1 co-expression. These patients may benefit from immunotherapy to target EBV-infected cells.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
