Abstract
The polymorphisms of invasion suppressor gene CDH1 and DNA mismatch repair gene Exo1 have been reported to play critical role in the development, tumorigenesis, and progression of several kinds of cancers including prostate cancer. This study was designed to analyze the contribution of single-nucleotide polymorphisms of the CDH1 (-160C/A) and Exo1 (K589E) to prostate cancer susceptibility in Bangladeshi population. The study included 100 prostate cancer cases and age-matched 100 healthy controls. Polymerase chain reaction-restriction fragment length polymorphism analysis was used to determine the genetic polymorphisms. A significant association was found between CDH1 -160C/A (rs16260) and Exo1 (rs1047840, K589E) polymorphisms and prostate cancer risk. In case of CDH1 -160C/A polymorphism, the frequencies of the three genotypes C/C,C/A, and A/A were 45%, 48%, and 7% in cases and 63%, 32%, and 5% in controls, respectively. The heterozygote C/A genotype and combined C/A + A/A genotypes showed 2.10-fold (odds ratio = 2.1000, 95% confidence interval = 1.2956–4.0905, p = 0.013) and 2.08-fold (odds ratio = 2.0811, 95% confidence interval = 1.1820–3.6641, p = 0.011) increased risk of prostate cancer, respectively, when compared with homozygous C/C genotypes. The variant A allele also was associated with increased risk of prostate cancer (odds ratio = 1.6901, 95% confidence interval = 1.0740–2.6597, p = 0.0233). In case of Exo1 (K589E) polymorphism, G/A heterozygote, A/A homozygote, and combined G/A + A/A genotypes were found to be associated with 2.30-, 4.85-, and 3.04-fold higher risk of prostate cancer, respectively (odds ratio = 2.3021, 95% confidence interval = 2.956–4.0905, p = 0.0031; odds ratio = 4.8462, 95% confidence interval = 1.0198–23.0284, p = 0.0291; OR = 3.0362, 95% confidence interval = 1.7054–5.4053, p = 0.0001, respectively). The “A” allele showed significant association with increased susceptibility (2.29-fold) to prostate cancer (odds ratio = 2.2955, 95% confidence interval = 1.4529–3.6270, p = 0.0004). Our results suggest that CDH1 -160C/A and Exo1 K589E polymorphisms are associated with increased susceptibility to prostate cancer in Bangladeshi population.
Introduction
Prostate cancer (PCa) mostly occurs to the men at an older age. It is the most commonly occurring non-cutaneous malignancy in males in developing countries. 1 American Cancer Society has estimated prostate cancer new cases and deaths in 2017 to be 161,360 and 26,730, respectively.2,3 In the present world, PCa is the third-leading cause of cancer-related deaths among males. 3 In Bangladesh, there are around 1.3–1.5 million prostate cancer patients among whom only 0.2 million patients are diagnosed each year (Strategy, 2009–15; Global Burden National Cancer Control of Disease Study, 2013). Prostate cancer might not have any symptoms in early stage. It means men having prostate cancer may not realize that they have it until they experience more difficulties in urination and sexual functions in the later stage. Unfortunately, incidence of mortality is seen to the majority men who are not diagnosed at early stage. However, since 1990, the average mortality rate from prostate cancer is increasing by 1.5% a year (Global Burden of Disease Study (ghdx.healthdata.org) as of 2013; refreshed July 2016). So, an effective screening method for prostate cancer is still required to minimize the cancer-related mortality in Bangladesh.
In a cohort study in Scandinavian countries, Lichtenstein et al. 4 demonstrated large effect of heritability in prostate cancer. That group indicated that 42% of the variation in PCa risk might be attributed to genetics. Alterations in several genes like mismatch repair genes (such as MLH1, MSH2, MSH6, and PMS2), ELAC2/HPC2 (17p11), MSR1 (8p22), RNase L/HPC1, and HOXB13 have been found to exert moderate-to-high risk for encountering PCa.5,6 Another study showed that 30% of estimated heritability of PCa can be attributed to single-nucleotide polymorphism (SNP). 7 SNP analyses have been found to be useful for identifying men with European ancestry to be at higher risk of prostate cancer. 8 Association of genetic SNPs of different genes and susceptibility of several cancers occurring around the world has been studied extensively.
E-cadherin (CDH1) gene encodes CDH1, a class of type-I transmembrane glycoproteins expressed in epithelial cells. E-cadherin plays important role in calcium-dependant cell–cell adhesion.9–11 Reduced expression of CDH1 gene has been found to be involved in the dysfunction of cell–cell adhesion system, leading to cancer invasion and metastasis. This is why CDH1 gene is considered as an important invasion suppressor gene. 12 Polymorphic variants of CDH1 gene have been reported in several populations. The -160C/A (rs16260) SNP, located at the upstream position of transcriptional site of the CDH1, has been shown to play a significant role in transcriptional activity and controls the development of catenin-containing complexes.13,14 A C→A transition at -160 imparts decreased transcriptional activity. 15 Compared with C allele, A allele has 68% decreased transcription factor binding capacity. 16 And thus presence of A allele has been implicated to increase the risk of many types of cancers in different ethnicities.16–18 The CDH1 (-160C/A) SNP has been reported to be associated with prostate cancer in Swedish people in 2004 19 and with other cancers such as colorectal cancer in the United Kingdom in 2009. 20
The gene Exo1 (exonuclease 1), located at chromosome 1q42 to q43, belongs to the RAD2 nuclease family and encodes 846-amino acid protein.21–23 Exo1 gene product functions in DNA replication, repair, and recombination. 24 Exo1 goes to the MMR (mismatch repair) system, which is one of the foremost DNA-repair pathways and confirms genomic stabilities, and also curbs recombination of DNA and cell cycle arrest in human.25–27 In addition, Exo1 has been shown exert mutation prevention activity. 28 SNPs of DNA repair genes have been reported to be associated with several types of human cancers counting prostate, lung, oral, and breast cancer.29–35 Among a number of SNPs of Exo1 (A1419G, C908G, A238G, C498T, K589E, G670E, C723R, L757P, and C3114T)36,37 studied, K589E SNP has been found to have positive association with susceptibility to lung, breast, and oral cancer in different ethnicities.32–34,37,38
Incidence of prostate cancer has been shown to vary based on ethnicity. To the best of our knowledge, no study regarding this association has been conducted in Bangladeshi population. Here we propose the association of CDH1 -160C/A and Exo1 K589E SNPs with prostate cancer susceptibility in Bangladeshi population. Thus, this study aims to reveal a better understanding of prostate cancer risk associated with these two polymorphisms in Bangladeshi male population counting some risk factors including age, weight, body mass index (BMI),and so on.
Materials and methods
Study population
One hundred adult prostate cancer patients and 100 unrelated gender- and age-matched healthy controls were recruited from same geographical area. Each volunteer (both patient and healthy controls) was informed about the experimental procedures and the aim of the study before signing the consent form. However, the volunteers were free to withdraw from the study anytime. This experiment has been conducted for about 1 year at Molecular Biology & Genetics Lab of Pharmacy Discipline, Khulna University in accordance with the declaration of Helsinki, and its subsequent revisions. 39 Samples were collected from the Ahsania Mission Cancer and General Hospital, Dhaka Medical College Hospital, Dhaka, and Bangabandhu Sheikh Mujib Medical University, Dhaka, Bangladesh. The experimental protocol has been approved by the ethical committees of the respective hospitals.
Venous blood collection and genomic DNA isolation
The volunteers were included in the experiment after proper counseling, explanation, and clarification of their queries. From both patients and control individuals, about 3 mL of venous blood was collected in a sterile vacuum tube containing EDTA-Na2 (ethylenediaminetetraacetic acid disodium) as anticoagulant. The blood samples were preserved at 4°C until the DNA isolation process. Genomic DNA from the blood sample was isolated following the DALY’s method. 40 The purity and quantity of the isolated DNA was then estimated by spectrophotometric assay using an ultraviolet (UV)-spectrophotometer (UV Prove v2.1). Absorbance quotient (OD260/OD280) value indicated the estimation of purity. Absorbance quotient value between 1.8 and 2.0 was considered to be good and purified DNA.
PCR genotyping assays
Polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) assay was used for genotyping of DNA found from both healthy volunteers (control) and cancer patients (case) because of its reliability and affordability. Figure 1 represents the schematic of the -160C/A SNP in CDH1 gene (A) and the K589E SNP in Exo1 gene (B) with respective restriction enzyme sites and the resulting fragments on digestion. PCR-RFLP method includes PCR amplification followed by digestion of the amplified PCR product with a restriction enzyme (REase). Desired genomic target region containing the specific SNP of interest has been amplified using primer-directed polycyclic chain reaction following Saiki et al.’s 41 method with required modification. Genomic DNA isolated from whole blood was used as PCR template. Each PCR mixture contained of about 50–70 ng template DNA, 0.25 µL Taq polymerase, 2.5 µL 10× standard reaction buffer, 0.625 µL dNTPs (2.5 mM), 1.25 µL of each primer (10 μM), and 18.125 µL water. For CDH1, gene thermal cycle was programmed as follows: initial denaturation step for 5 min at 95 C, followed by 30 cycles of 40 s at 95 C as denaturation, 30 s at 66 C as annealing, 50 s at 72 C as elongation, and final elongation for 10 min at 72 C. However, for Exo1, gene amplification condition was as follows: initial denaturation at 95°C for 3 min followed by 35 cycles with denaturing at 95°C for 30 s, annealing at 55°C for 30 s and extension at 72°C for 30 s, and final extension at 72°C for 10 min.

Schematic of the -160C/A SNP in CDH1 gene (a) and the K589E SNP in Exo1 gene (b) with restriction sites of each enzyme and the resulting fragments.
The amplified PCR products of both genes (CDH1, 448 bp; Exo1, 306 bp) were then confirmed by gel electrophoresis in 2% agarose gel (Figures 2(a) and 3(a)). The primer sequences for human CDH1 were as follows: forward, 5′-CGCCCCGACTTGTCTCTCTAC-3′; reverse, 5′-GGCCACAGCCAATCAGCA-3′, and for Exo1 gene were forward, 5′-GACACAGATGTAGCACGTAA-3′; reverse, 5′-CTGCGACACATCAGACATAT-3′.

(a) PCR product of CDH1 (448 bp), Lanes 1 to 7, Lane-M contains molecular ruler. (b) Restriction enzyme (HincII) digestion fragments of CDH1 (Lane 1 to 7) as viewed in 3% agarose gel. Lane-M, molecular ruler. Lane-1, 3, 4, 5, and 6, normal homozygote (448 bp), Lane-7, heterozygote (448, 368, 80 bp), and Lane-2, mutant homozygote (368, 80 bp).

(a) PCR product of Exo1 (306 bp), Lanes 1 to 5, Lane-M contains molecular ruler. (b) Restriction enzyme (Msel) digestion fragments of Exo1 (Lanes 1–7) as viewed in 2% agarose gel. Lane-M, molecular ruler; Lane-1, 2, 4, 5, and 6, normal homozygote (306 bp), Lane-7, heterozygote (306, 196, 110 bp), and Lane-3, mutant homozygote (196, 110 bp).
RFLP assays
Restriction digestion of the PCR product will determine the presence or absence of the desired SNPs at the restriction site. RFLP assay confirms the frequencies of desired SNP within a specified sample cohort. After getting the specific PCR products, they were digested with restriction endonucleases. PCR products for CDH1 gene and Exo1 gene were digested with 2 units of HincII (Thermo Fisher Scientific, USA) at 37°C for overnight with Msel (Tru1l) (Thermo Fisher Scientific, USA) at 65°C for 16 h, respectively, following the manufacturers’ standard protocol. The digested products of both genes were then evaluated by electrophoresis through 3% agarose gel (Figures 2(b) and 3(b)). Homozygous A/A genotype of CDH1 was cut by HincII yielding two fragments with 368 and 80 bp, C/C genotype produced a single 448 bp band because of absence of cutting site, whereas heterozygous genotype resulted all three bands. Similarly, on MseI digestion of Exo1 PCR products, homozygous A/A allele yielded 196 and 110 bp fragments, homozygous G/G genotype yielded single 306 bp, whereas heterozygosity yielded all three fragments. We checked our initial results by regenotyping randomly selected 20 samples (10% of our total samples) by the previously followed PCR-RFLP method.
Statistical analysis
Genotype as well as allelic frequencies was expressed in percentage. Before performing disease risk analysis, the distribution of observed and expected frequencies of the genotypes in our study population in Hardy–Weinberg equilibrium was analyzed using Pearson’s chi-square test. The distribution of genotype frequencies was compared between cases and healthy controls using the chi-square test. Multiple logistic regressions were used to estimate crude odds ratio (OR) and their 95% confidence intervals (CIs) using the statistical software package MEDCAL version 13.3.3. For both genes, the homozygous and heterozygous carriers of the polymorphisms were regarded as polymorphic genotypes. IBMSPSS (Statistical Package for the Social Sciences) Statistics 25 software was applied for multivariate logistic regression analyses to estimate the association between genotype combinations and risk of prostate cancer by measuring the crude ORs and 95% CIs. Fisher exact probability test (two-tailed) was performed when the number of samples was very low. Two-sided statistical test of significance was conducted and a p < 0.05 was regarded as statistically significant.
Results
Characteristics and features of the subject
This is a case–control study conducted on 200 individuals consisting of 100 prostate cancer patients and 100 healthy controls for both genes analyzed. The mean ages were 65.60 years (range 63–72 years) for cases, whereas 65.69 years (range 62–72 years) for controls. It was predicted that the incidence of prostate cancer was higher among individuals ≥65 years of age (59%) compared with those <65 years of age (41%). Table 1 represents the distribution of selected demographic variables such as age, weight, and BMI between prostate cancer cases and controls. With respect to these parameters, we did not find any statistically significant difference between cases and controls used in this study.
Distribution of selected demographic variables among prostate cancer patients (cases) and controls.

The purity of the extracted genomic DNA material was determined by spectrometric measurement of OD260/OD280 ratios, which were found to be within the range of 1.6–1.9. The PCR products were checked by gel electrophoresis in 2% agarose gel to see whether they were in their expected lengths. PCR product of CDH1 gene was 448 bp (Figure 2(a)) and that of Exo1 gene was 306 bp (Figure 3(a)). After confirmation of the size of the PCR products, restriction enzyme digestions were carried out using HincII for CDH1 and Msel (Tru1l) for Exo1. Figure 1(b) shows the genotypes resulted from CDH1 (-160C/A) named as normal homozygote (C/C; 448 bp), heterozygote (C/A; 448, 368, and 80 bp), and variant homozygote (A/A; 368 and 80 bp). Similarly, Figure 3(b) shows the digested fragments of Exo1 (K589E) termed as normal homozygote (G/G; 306 bp), heterozygote (G/A; 306, 196, and 110 bp), and variant homozygote (A/A; 196 and 110 bp).
Correlation of different genotypes with prostate cancer
Impact of CDH1 (-160C/A) polymorphism on prostate cancer risk
In this experiment, the association and impact of CDH1 (-160C/A) have been studied with the prostate cancer susceptibility. Genomic and allelic frequencies and distribution of both genes have been analyzed and summarized in this study (Tables 2 and 3). The frequencies of the three genotypes C/C, C/A, and A/A of CDH1 (-160C/A) were found to be 45%, 48%, and 7% for cancer cases and 63%, 32%, and 5% for healthy controls, respectively. The distribution of the above genotypes did not deviate significantly (p > 0.05) from the Hardy–Weinberg equilibrium. The frequency of heterozygote (C/A) genotype and heterozygote plus variant homozygote (C/A + A/A) was significantly (p < 0.05) increased in prostate cancer cases as compared with controls (48% vs 32% and 55% vs 37%, respectively). In analysis of the genotypes of the CDH1 (-160C/A), heterozygote (C/A) genotype showed to be significantly associated with 2.1 times increased risk of prostate cancer (OR = 2.1, 95% CI = 1.1657–3.7830, p = 0.0135) when compared with the wild-type normal homozygote (C/C).
Genotype frequencies of CDH1 (-160C/A) and Exo1 (K589E) gene polymorphisms in relation to prostate cancer cases and controls.
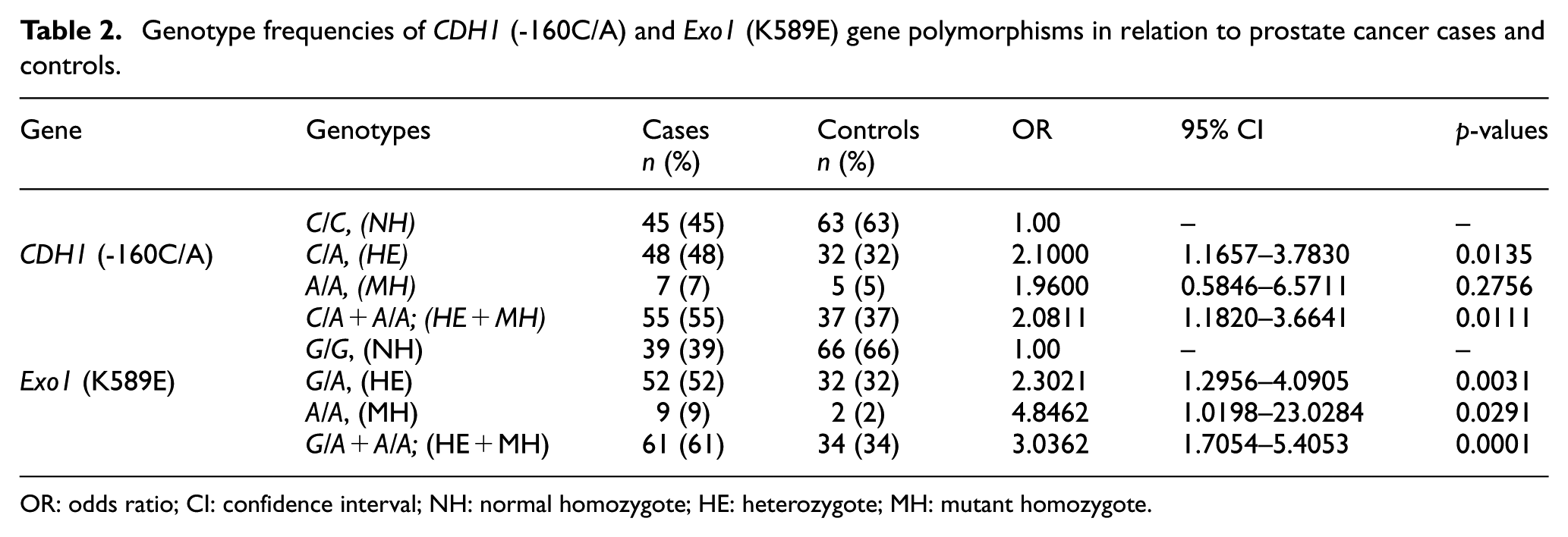
OR: odds ratio; CI: confidence interval; NH: normal homozygote; HE: heterozygote; MH: mutant homozygote.
Allele frequencies of CDH1 (-160C/A) and Exo1 (K589E) polymorphism in relation to PC cases and controls.

OR: odds ratio; CI: confidence interval.
We also compared the normal homozygote (C/C) genotype with heterozygote plus variant homozygote (C/A + A/A) genotype, which showed 2.08-fold significantly increased risk of prostate cancer (OR = 2.0811, 95% CI = 1.1820–3.6641, p = 0.0111) (Table 2). However, we did not get any statistically significant association with variant homozygote (A/A).
Allelic frequencies of CDH1 (-160C/A) among the prostate cancer cases and controls and their association with prostate cancer susceptibly are shown in Table 3. Normal (C) and variant (A) allele contents were found to be 69.0% and 31.0% in prostate cancer cases whereas 79.0% and 21.0% in controls, respectively. Distribution of A allele was significantly increased in prostate cancer cases compared with controls. Variant (A) allele has been found to be significantly associated with 1.69-fold increased susceptibility of prostate cancer (OR = 1.6901, 95% CI = 1.0740–2.6597, p = 0.0233) (Table 3). Moreover, the allelic frequencies found in other studies in different ethnic groups and regions are also shown in Table 4.
Comparison of our study results with different ethnic groups for both CDH1 (-160C/A) and Exo1 (K589E).

Impact of Exo1 (K589E) polymorphism on prostate cancer risk
The association of Exo1 (K589E) gene with prostate cancer susceptibility has also been studied (Tables 2 and 3). The genotypic distributions did not deviate significantly (p>0.05) from the Hardy–Weinberg equilibrium. The frequencies of the normal homozygote (G/G), heterozygote (G/A), and variant homozygote (A/A) genotypes were found to be 39%, 52%, and 9% for cases and 66%, 32%, and 2% for controls, respectively. The frequencies of G/A, A/A, and G/A + A/A genotypes were significantly increased in prostate cancer cases when compared with controls. Compared with wild-type normal homozygote (G/G), both heterozygote (G/A) and variant homozygote (A/A) showed 2.3-fold (OR = 2.3021, 95% CI = 1.2956–4.0905, p = 0.0031) and 4.85-fold (OR = 4.8462, 95% CI = 1.0198–23.0284 p = 0.0291) significant increases in prostate cancer susceptibility, respectively. In keeping likeness, the distribution of heterozygote plus mutant homozygote (G/A + A/A) together significantly increases 3.04 times the prostate cancer risk (OR = 3.0362, 95% CI = 1.7054–5.4053, p = 0.0001) in relation to controls.
Table 3 listed the allele frequency of Exo1 (K589E) for both cases and controls where normal (G) and mutant (A) allele frequencies were 65.0% and 35.0% for cases and 82.0% and 18.0% for controls, respectively. According to our findings, variant (A) allele was observed to be positively associated with prostate cancer susceptibility (OR = 2.2955, 95% CI = 1.4529–3.6270, p = 0.0004). We have also compared the allelic distribution of CDH1 (-160C/A) and Exo1 (K589E polymorphisms in our study population with that observed in different regions in different studies by other investigators (Table 4).
Combined genotypic effects of CDH1 (-160C/A) and Exo1 (K589E) polymorphisms on prostate cancer risk
The CDH1 (-160C/A) and Exo1 (K589E) SNPs are located on two different chromosomes namely 16q22.1 and 1q42 to q43, respectively. These two loci assort independently and are said to be unlinked as they are found on different chromosomes. However, we found these SNPs to be associated with prostate cancer risk when considered in individualistic way. Then we performed analysis to check whether the combined effects of the CDH1 (-160C/A) and Exo1 (K589E) SNPs influence the risk of prostate cancer in our samples (Table 5). Our results revealed that the combination of some variants increases the susceptibility to prostate cancer development when compared with the two wild-type genotypes. CDH1 (-160C/A)–Exo1 (Any A) (OR = 9.410; 95% CI = 3.371–26.265; p < 0.0001) genotypic interactions have been shown to cause the highest increase in prostate cancer risk followed by CDH1 (-160Any A)–Exo1(Any A) (OR = 8.726;95% CI = 3.365–22.627; p < 0.0001) and CDH1 (-160C/A)–Exo1(G/A) (OR = 8.324; 95% CI = 2.954–23.452; p = 0.0001) genotypic interactions. In addition, the combination of the CDH1 (-160C/C)–Exo1 (A/A) genotypes was associated with more than sevenfold increase in prostate cancer susceptibility (OR = 7.60; 95% CI = 1.377–41.947; p = 0.016). The other two genotypic combinations CDH1 (-160A/A)–Exo1(G/A) and CDH1 (-160A/A)–Exo1 (A/A) genotypes led to an increase with ORs of 6.333 (OR = 6.333; 95% CI = 1.106–36.276; p = 0.036).
Interaction between CDH1 (-160C/A) and Exo1 (K589E) genotypes and prostate cancer risk.
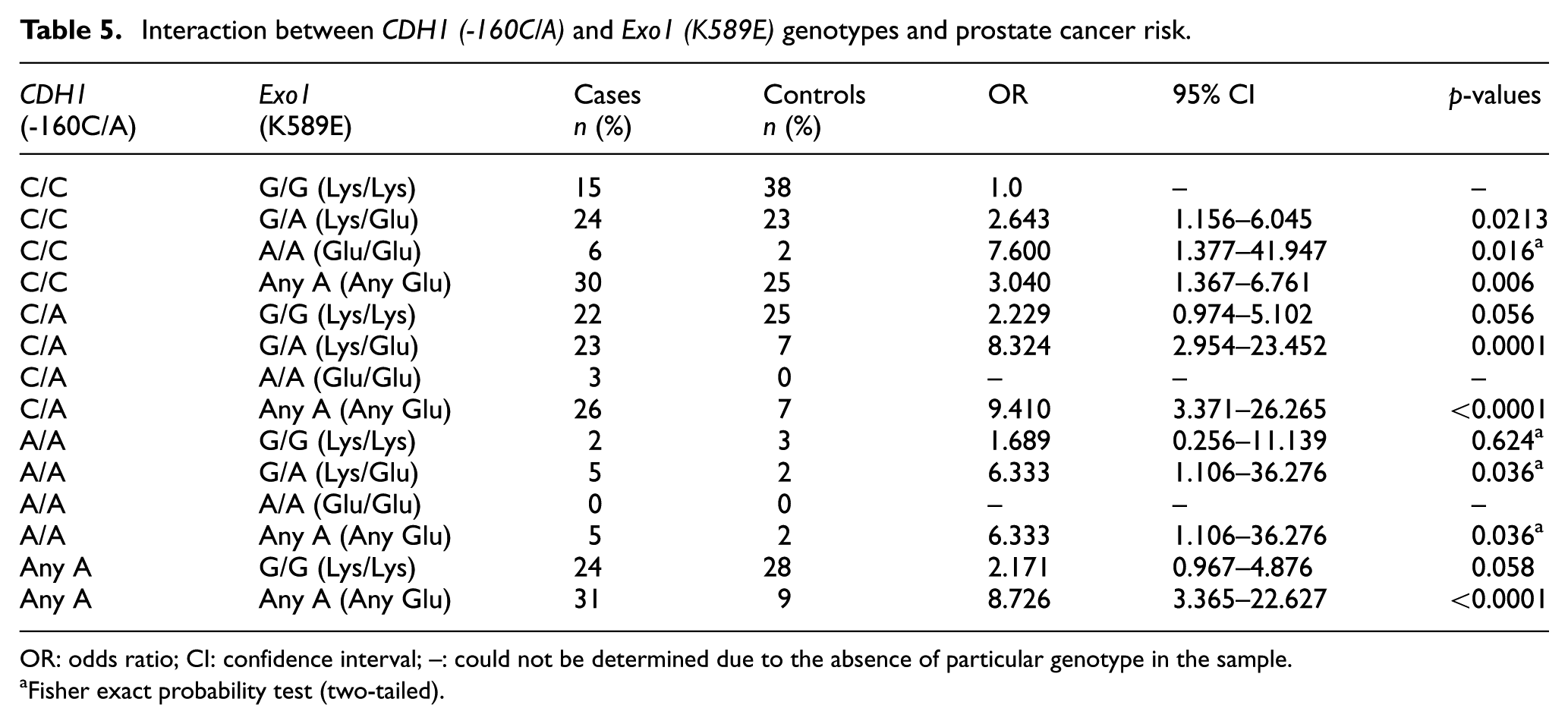
OR: odds ratio; CI: confidence interval; –: could not be determined due to the absence of particular genotype in the sample.
Fisher exact probability test (two-tailed).
Then we further analyzed the distribution of CDH1 (-160C/A) and Exo1 (K589E) polymorphisms with the prostate cancer patients stratified by some risk factors such as age, weight, and BMI. The association between genotypic frequencies and the variables is summarized in Table S1 (Supplemental material). Unfortunately, the variables examined were not found to be significantly associated with CDH1 (-160C/A) and Exo1 (K589E) polymorphisms in prostate cancer patients.
Discussion
Prostate cancer is the most commonly occurring non-cutaneous malignancy in men in developing countries 1 and is the third-leading cause of cancer-related deaths among males in the current world. 3 Prostate cancer has been reported to result from complex interactions between environmental and genetic factors where genetic factors play the key roles in its pathogenesis.48–50 Individual differences in cancer occurrences have been attributed to some host factors including genetic polymorphisms.51,52SNPs in the genomic DNA are the most common genetic variants to be associated with progression of a number of cancers51–54 including prostate cancer.55,56
Promoter region of E-cadherin (CDH1) gene was found highly polymorphic in several populations. 13 CDH1 -160C/A SNP (rs16260), detected in the promoter region related to the transcriptional start site, was identified to alter the transcriptional competency of this gene.19,57 In prostate cancer cell lines, the transcriptional activity of the -160A allele was reduced by 68% compared with its C allele in vitro.17,57,58 The reduced transcriptional activity has been attributed to reduced expression. CDH1 -160C/A polymorphism within the promoter region has been reported to play important role in the susceptibility of various types of cancers including prostate, colorectal, urothelial, pancreatic, gastric cancers, and so on.13,16,57,59–61 However, in this study we have performed a case–control study to clarify the impact of E-cadherin -160C/A polymorphism on prostate cancer risk in Bangladeshi population.
Our study included Bangladeshi male prostate cancer patients and age-matched healthy controls. First, we investigated CDH1 (-160C/A) SNP for its association to the prostate cancer in our population. The frequency of heterozygote (C/A) genotype and heterozygote plus variant homozygote (C/A + A/A) was significantly (p < 0.05) increased in prostate cancer cases as compared with controls (48% vs 32% and 55% vs 37%, respectively). 18 Heterozygote (C/A) genotype showed 2.1-fold increased risk of prostate cancer (OR = 2.1, 95% CI = 1.1657–3.7830, p = 0.0135), whereas heterozygote plus variant homozygote (C/A + A/A) genotypes showed 2.08-fold increased risk (OR = 2.0811, 95% CI = 1.1820–3.6641, p = 0.0111). All the genotypes were compared with the wild-type normal homozygote (C/C). This finding of our study is in agreement with the previous findings of a meta-analysis where -160A allele carriers (CA + AA) of the E-cadherin gene showed to have a 21% elevated risk of prostate cancer. They also reported that the elevation in prostate cancer risk associated with -160A allele was evident in both European and Asian populations but not in Africans. 18 Wang et al. also showed that Asians and Europeans with -160A allele carriers exhibited more susceptibility to prostate cancers, whereas the North American carriers showed tolerance. There was no noticeable difference between Asians and Europeans. 17 In addition to prostate cancer, some groups studied the impact of -160C/A polymorphism in other types of cancers. In a meta-analysis, Wang et al. 61 showed -160A/C allele to increase the risk factor urothelial cancers, whereas other groups reported -160C/A allele not to be associated with breast cancer, 62 gastric cancer,63,64 and pancreatic cancer. 45 On the contrary, -160A/A variant homozygote allele was reported to decrease the gastric cancer risk in Japanese and Taiwanese population.65,66 The -160A/A homozygote allele was shown to play protective role in susceptibility to colorectal cancer in the Europeans 61 and to pancreatic cancer in Han Chinese population. 45 Interestingly, in our study population, mutant homozygote -160A/A allele of E-cadherin did not show any statistically significant association with prostate cancer susceptibility, although the OR value was 1.96 (95% CI = 0.5846–6.5711; p = 0.2756). Our finding related to variant homozygote (-160A/A) allele was in consistent with previous findings of some groups who reported no significant association with gastric cancer in Chinese and Italian population.63,64 On the other hand, Tipirisetti et al. 67 reported CDH1-160A/A variant to be associated with breast cancer among Indian women. This inconsistency in effect indicates that the role of -160A/A allele might be organ and ethnicity dependant. As our sample size was limited, further study using a larger population is necessary before drawing a conclusion.
The Exo1 gene comprises one untranslated exon followed by 13 coding exons that encode an 846-amino acid protein.21–23 The Exo1 gene product, a member of the RAD2 nuclease family, functions in a number cellular pathways including DNA repair, replication, recombination, and telomere integrity. 60 Exo1 is the only exonuclease involved in the human MMR system that corrects the mismatches between bases and small insertion or deletion loops.68,69 A functional MMR system preserves genomic stability, modulates DNA recombination, and mediates cell cycle arrest. 70 If DNA damage is not properly repaired and the cells are not induced to undergo apoptotic elimination, defects in the DNA mount up and spread through the cell progeny, finally resulting in carcinogenesis. 71 SNPs of DNA repair genes have been shown to be associated with several types of cancer occurrence in different ethnicities.29–32,72 Functional defects in this DNA repair system have been shown to play important role in cancer progression. Because of its key role in the MMR system, Exo1 gene has become a significant focus to be studied widely for its association with susceptibility of various cancers.46,73,74 Several SNPs of Exo1 gene have been reported as genetic risk factors of cancer. However, findings of the studies regarding effect of Exo1 polymorphisms on cancer susceptibility remain inconsistent. In this study, we investigated the association of Exo1 K589E (rs1047840) SNP with prostate cancer susceptibility in Bangladeshi population.
In our study, we found Exo1 K589E (G/A) SNP to be significantly associated with prostate cancer susceptibility in Bangladeshi population. Heterozygote (G/A) and variant homozygote (A/A) genotypes showed 2.3-fold (OR = 2.3021, 95% CI = 1.2956–4.0905, p = 0.0031) and 4.85-fold (OR = 4.8462, 95% CI = 1.0198–23.0284 p = 0.0291) significant increases in prostate cancer susceptibility, respectively. Similarly, combined heterozygote plus variant homozygote (G/A + A/A) was shown to raise the risk of prostate cancer by 3.04 times (OR = 3.0362, 95% CI = 1.7054–5.4053, p = 0.0001) when compared with wild-type normal homozygote (G/G). The outcome of the Exo1 K589E polymorphism, located on exon 13, is alteration in amino acid at codon 589 from negatively charged glutamate (Glu, E) residue (GAG) to a positively charged lysine (Lys, K) residue (AAG). As the position of K589E SNP is at an exonic splicing enhancer region, the amino acid alteration at this position might manipulate the protein functions. 33
Our findings in Bangladeshi population is in consistent with the work investigating association of Exo1 K589E polymorphism with gastric cancer risk in central Taiwanese population,33,36 with lung cancer in Chinese population, 33 with breast cancer in Taiwanese population, 37 and with cervical cancer in Chinese population. 75 In another study in Turkish population, Bayram et al. 76 have shown that Lys/Lys genotype of the Exo1 K589E polymorphism is significantly associated with increased risk of HCC development. However, Zienolddiny et al. 77 did not find any significant association between Exo1 K589E SNP and non–small cell lung cancer susceptibility in Caucasian Norwegian population. So, the observed variation in the genotypic distribution of Exo1 K589E polymorphism might be due to the dissimilarity in the carcinogenesis in different tissues. In a meta-analysis, Duan et al. have shown that Exo1 K589E SNP was significantly associated with increased cancer risk in Asian population but not in the Caucasian population. This suggested the effect to be ethnicity dependent. 43
We also estimated the allele frequency of both CDH1 (-160C/A) and Exo1 (K589E) SNPs for both prostate cancer cases and controls. Our results demonstrated that the variant A allele in CDH1 (-160C/A) and Exo1 (K589E) genes was significantly associated with increased risk of prostate cancer. Then we also have carried a petite assessment of allelic distribution for both CDH1 (-160C/A) and Exo1 (K589E) genes in Bangladeshi prostate cancer cases and different regions around the globe. In our study, the normal C and variant A allele frequencies in CDH1 gene were found to be 69.0% and 31.0% in cases. On the other hand, frequency of variant A allele was found to be 16.6% in Han Chinese population for pancreatic cancer, 45 15.9% and 20.0% in Chinese population for gastric and esophageal cancer, 44 17.0% in Mexico for gastric cancer, 42 and 34.2% in India for breast cancer. 67 For Exo1 (K589E) gene, variant A allele frequency in our study group was found to be 35.0%, whereas in other studies carried out in China and Taiwan, the frequencies were 24.3%, 21.0%, 22.8%, and 25.0% for gastric, 36 breast, 37 lung, 38 and cervical cancer, 75 respectively. Among Iranian population, A allele frequency was also found 31.3% for colorectal cancer. 47
We evaluated the combined effect of two polymorphisms under study that may modify the effects of individual genetic variation. Single CDH1 -160A/A polymorphism was not found to be associated with prostate cancer risk in this study population. However, combination of this allele with Exo1 G/A (Lys/Glu) allele was shown to increase the risk of prostate cancer by 6.3-fold (OR, 6.33; 95% CI, 1.106–36.276; p = 0.036), which clearly indicated the role of combination of cancer-related polymorphisms. Combined effect of the genotypes of CDH1 (-160C/A) and Exo1 (G/A) showed a trend of increased risk to a great extent for CDH1 (-160A/A) and Exo1 G/G combination (Table 5). The highest increase in risk was observed with CDH1 (-160C/A)–Exo1 any A combination (OR = 9.410; 95% CI, 3.371–26.265; p < 0.0001) with 9.4-fold increase.
We have also tried to find an association of some selected risk factors such as age, weight, and BMI with the prostate cancer susceptibility among the Bangladeshi men. But, in this case, we did not have any significant association of these risk factors with the PCa prevalence in Bangladesh.
Conclusion
In conclusion, for the first time in Bangladesh, our data provided some evidence that CDH1 -160C/A and Exo1 K589E SNPs are significantly associated with increased susceptibility to prostate cancer in Bangladeshi population. However, our findings regarding the direct effect of the polymorphic variants on the prostate cancer risk need to be validated by further functional analysis. Moreover, well-designed population-based studies with larger sample sizes will be of greater value to determine the true effects of these polymorphisms for prostate cancer.
Supplemental Material
Figure_S1 – Supplemental material for Genetic polymorphisms in CDH1 and Exo1 genes elevate the prostate cancer risk in Bangladeshi population
Supplemental material, Figure_S1 for Genetic polymorphisms in CDH1 and Exo1 genes elevate the prostate cancer risk in Bangladeshi population by Hasnain Imtiaz, Sharmin Afroz, Md Amir Hossain, Sm Faysal Bellah, Md Mostafizur Rahman, Md Shahin Kadir, Razia Sultana, Md Abdul Mazid and Md Mustafizur Rahman in Tumor Biology
Supplemental Material
Table_S1 – Supplemental material for Genetic polymorphisms in CDH1 and Exo1 genes elevate the prostate cancer risk in Bangladeshi population
Supplemental material, Table_S1 for Genetic polymorphisms in CDH1 and Exo1 genes elevate the prostate cancer risk in Bangladeshi population by Hasnain Imtiaz, Sharmin Afroz, Md Amir Hossain, Sm Faysal Bellah, Md Mostafizur Rahman, Md Shahin Kadir, Razia Sultana, Md Abdul Mazid and Md Mustafizur Rahman in Tumor Biology
Footnotes
Acknowledgements
The authors are thankful to the Pharmacy Discipline of Khulna University, Bangladesh, for providing laboratory facilities and instrumental support. We are also thankful to the staff, nurses, physicians, and the authority of Ahsania Mission Cancer and General Hospital, Dhaka Medical College Hospital, Dhaka, and Bangabandhu Sheikh Mujib Medical University (BSMMU), Dhaka, Bangladesh, for kind permission and support to collect blood samples from prostate cancer patients. We would like to express our gratitude to Ministry of Science and Technology, Govt. of People’s Republic of Bangladesh for providing partial financial support for this study. The authors have no other relevant affiliation or financial involvement with any organization. H.I. and S.A. contributed equally.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
The ethical committees of the hospitals approved the study protocol, and the study was conducted according to the Helsinki declaration and its subsequent revisions.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
