Abstract
GATA4, a transcription factor crucial for early liver development, has been implicated in the pathophysiology of hepatoblastoma, an embryonal tumor of childhood. However, the molecular and phenotypic consequences of GATA4 expression in hepatoblastoma are not fully understood. We surveyed GATA4 expression in 24 hepatoblastomas using RNA in situ hybridization and immunohistochemistry. RNA interference was used to inhibit GATA4 in human HUH6 hepatoblastoma cells, and changes in cell migration were measured with wound healing and transwell assays. RNA microarray hybridization was performed on control and GATA4 knockdown HUH6 cells, and differentially expressed genes were validated by quantitative polymerase chain reaction or immunostaining. Plasmid transfection was used to overexpress GATA4 in primary human hepatocytes and ensuring changes in gene expression were measured by quantitative polymerase chain reaction. We found that GATA4 expression was high in most hepatoblastomas but weak or negligible in normal hepatocytes. GATA4 gene silencing impaired HUH6 cell migration. We identified 106 differentially expressed genes (72 downregulated, 34 upregulated) in knockdown versus control HUH6 cells. GATA4 silencing altered the expression of genes associated with cytoskeleton organization, cell-to-cell adhesion, and extracellular matrix dynamics (e.g. ADD3, AHNAK, DOCK8, RHOU, MSF, IGFBP1, COL4A2). These changes in gene expression reflected a more epithelial (less malignant) phenotype. Consistent with this notion, there was reduced F-actin stress fiber formation in knockdown HUH6 cells. Forced expression of GATA4 in primary human hepatocytes triggered opposite changes in the expression of genes identified by GATA4 silencing in HUH6 cells. In conclusion, GATA4 is highly expressed in most hepatoblastomas and correlates with a mesenchymal, migratory phenotype of hepatoblastoma cells.
Introduction
Hepatoblastoma (HB) is a rare pediatric malignancy of the liver arising from hepatic progenitor cells and hepatoblasts and resembles fetal or embryonal liver tissue. In keeping with its probable fetal origin, HB is usually diagnosed in children under 4 years of age. 1 HB is associated with prematurity and certain specific genetic syndromes. 2 Activating mutations in Wnt pathway genes, especially CTNNB1, are frequent in these tumors. 3 However, the majority of the HBs are sporadic, and their molecular pathogenesis is poorly understood.
GATA transcription factors are a family of six evolutionarily highly conserved proteins that serve important physiological and pathological functions in various organs. 4 The genes encoding GATA4 and GATA6 are abundantly expressed in early fetal liver and crucial for the normal liver (NL) development.5,6 Later in gestation, their expression decreases in hepatocytes, and in postnatal liver, GATA4 expression is mainly restricted to endothelial cells and liver macrophages.7,8 GATA4 is expressed in the majority of HBs but not in adult hepatocellular carcinomas.7,9
The mechanisms by which GATA4 affects the malignant behavior of HB cells remains an area of active investigation. We have shown that GATA4 modulates expression of intrinsic apoptotic pathway components and protects HB cells from doxorubicin. 10 A recent report suggests that in addition to promoting cell survival, GATA4 regulates migration and invasion of HB cells. 11 Thus, an emerging paradigm is that GATA4 expression in HB cells favors a more mesenchymal, migratory phenotype. Here, we use silencing and overexpression of GATA4 in HUH6 HB cells and primary human hepatocytes, respectively, to unravel the molecular basis for this phenotypic switch. We show that modulation of GATA4 levels impacts the expression of genes involved in cytoskeleton organization, cell–cell adhesion, and extracellular matrix (ECM) dynamics.
Materials and methods
Human tissue samples
Human tissue samples from HB tumor patients and healthy liver transplantation donors were obtained from Helsinki University Hospital. The use of the samples in this study was approved by the Ethical Committee of Helsinki University Hospital and by the National Supervisory Authority for Welfare and Health. The study group included 24 patients with histologically diagnosed HB and 17 healthy organ donors (age 1–62 years). HB tumor distribution according to PRETEXT 12 staging in the study group was II (n = 7), III (n = 9), and IV (n = 8).
Cell culture
Established human HB cell lines HUH6 (mixed epithelial/mesenchymal HB) and HepG2 (epithelial HB) were cultured at 37°C in Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10% fetal calf serum, 2 mM
GATA4 silencing and overexpression in cell culture
GATA4 was downregulated in HUH6 and HepG2 cells via small interfering RNA (siRNA) transfection as previously described. 10 Cells were incubated for 24, 48, or 72 h after siRNA transfection and subsequently analyzed for migration and gene expression. In parallel experiments, a pMT2-GATA4 expression plasmid was used to overexpress GATA4 in human primary hepatocytes. 13 jetPEI™-Hepatocyte DNA reagent (Polyplus Transfection, Illkirch, France) was used for plasmid transfection. Cells were incubated 48 h after plasmid transfection and subsequently analyzed for gene expression.
Migration assays
Two complementary methods were used to assess the influence of GATA4 on cell migration. (1) For the wound healing assay, HUH6 cells were transfected with GATA4 siRNA (G4) or control non-target siRNA (NT) and cultured for 24 h on a 6-well plate coated with collagen I (Sigma Aldrich, MO, USA). A scratch was created in the cell monolayer with 120 µL pipette tip and images were captured at the outset (0 h) and after 24 h. Total of eight images per well were analyzed by calculating remainder of migrating cell area from the two time-points. (2) For the transwell assay, HUH6 cells were transfected with siRNA (G4 or NT) on a 10-cm plate, cultured for 48 h, and detached. A suspension of 6 × 104 transfected cells was applied to 8-µm-pore inserts, with serum-free media in the upper chamber and normal growth media in the lower chamber. The bottom of the insert was coated with collagen I (1 mg/mL). Cells were allowed to migrate for 40 h and then the cells were fixed with 4% paraformaldehyde in phosphate-buffered saline (PBS) and stained with crystal violet. The number of migrated cells was counted by light microscopy from five fields per well, with magnification of 10×. Both migration assays were performed in triplicate.
RNA microarray hybridization
RNA was extracted from two sample sets of HUH6 cells transfected with NT or G4 siRNA. Each sample set included three NT siRNA and three G4 siRNA samples. RNA quality was assessed with Bioanalyzer (Agilent, CA, USA). All 12 (6 + 6) samples were subjected to RNA microarray hybridization, performed by the Functional Genomics Unit at the University of Helsinki using an Illumina HumanHT-12 v4 oligonucleotide expression BeadChip (Illumina, CA, USA).
Quantitative real-time polymerase chain reaction
Total RNA was extracted using the NucleoSpin RNA/Protein kit (Macherey-Nagel, Düren, DE). Reverse transcription was performed using the Reverse Transcriptase Core Kit, and quantitative polymerase chain reaction (qPCR) was carried out using the MESA GREEN qPCR MasterMix Plus for SYBR Assay (both from Eurogentec, Seraing, Belgium). The geometric mean of two housekeeping genes, PPIG and GAPDH, was used as reference for messenger RNA (mRNA) expression. Primer pairs used are listed in Supplementary Table 1.
RNA in situ hybridization
Fluorescence RNA in situ hybridization was performed as previously described. 14 The probes used were as follows: Hs-GATA4 (#579821), positive control probe Hs-PPIB (#313901), and negative control probe DapB (#310043). TSA Plus Cyanine 3 fluorophore (NEL744001KT, Perkin Elmer) was used for Hs-GATA4 probe at 1:1500 dilution.
Western blotting
Western blotting was performed as previously described. 14 The primary antibodies used were as follows: goat antihuman GATA4 (AF2606, Novus Biologicals, Abingdon, UK) at a 1:500 dilution, mouse antihuman E-cadherin (610181) at a 1:1000 dilution (BD Transduction laboratories, CA, USA), mouse antihuman N-cadherin (610920) at a 1:1000 dilution (BD Transduction laboratories), and goat antihuman GAPDH (Ab-9485) at a 1:1000 dilution (Abcam, Cambridge, UK).
Immunohistochemistry
Formalin-fixed, paraffin-embedded liver sections were deparaffinized, hydrated, and treated with citratuffer (Target retrieval solution; Dako, Glostrup, Denmark) at 97°C for 30 min. Endogenous peroxidase activity was blocked with 3% hydrogen peroxide, and nonspecific binding was blocked with 1.5% normal serum. A polymerized reporter enzyme staining system (ImmPRESS reagent kit; Vector Laboratories, CA, USA or Novolink Polymer Detection System; Leica Biosystems, Newcastle, UK) was used to visualize the bound antibody. Primary antibody was incubated overnight at 4°C. The primary antibody was mouse antihuman GATA4 IgG (MAB2606, R&D systems, MN, USA) at a 1:5000 dilution. In control experiments, nonimmune serum replaced the primary antibody. Immunoreactivity was scored by two independent researchers as follows: negative = <10% positive nuclei, low/intermediate = <50% positive nuclei, and high = >50% positive nuclei.
Immunofluorescence staining
HUH6 cells were cultured in Matrigel-coated Nunc LabTek 2-well chamber slide systems (Thermo Scientific, Waltham, MA, USA) for 48 h after transfection. Cells were fixed and permeabilized in 100% methanol and washed in PBS. Unspecific binding was blocked with UltraVision Protein Block (Thermo Scientific). Cells were incubated with appropriate primary antibodies for 1 h at room temperature. Primary antibodies used were as follows: mouse antihuman GATA4 (MAB2606) at dilution 1:150, mouse antihuman E-cadherin (610181) at a 1:50 dilution (BD Transduction laboratories), and mouse antihuman N-cadherin (610920) at a 1:200 dilution (BD Transduction laboratories). Secondary antibody incubation was performed with AlexaFluor 488 anti-mouse antibody (Life Technologies, CA, USA) at a 1:200 dilution at room temperature covered from light. F-actin immunostaining with phalloidin-fluorescein isothiocyanate (FITC)-conjugated antibody (P5282) at a 1:250 dilution (Sigma Aldrich, St. Louis, MO, USA) was performed according to manufacturer’s instructions.
Statistical analyses
For microarray data, background correction was done on raw array data using BeadStudio software (Illumina) before quantile normalization and log 2 transformation were performed using the BeadArray Bioconductor package. 15 Differentially expressed genes were identified using linear models for microarray data (LIMMA) 16 with Benjamini–Hochberg correction. The expression level changes with a false discovery rate (FDR) below 5% were considered as significantly differentially expressed. Microarray data were subjected to average linkage clustering with uncentered correlation using Cluster, 17 and heatmap was generated with R software. 18 GOstats Bioconductor package was used to perform gene set enrichment analysis of the differentially expressed. 19 Hypergeometric tests with the Benjamini–Hochberg FDR were performed to adjust the p-value. qPCR and migration assay experiments were performed in triplicates and statistically analyzed using Student’s T-test; p < 0.05 was considered significant in all experiments.
Results
GATA4 is overexpressed in most HBs
To confirm and extend our earlier report that GATA4 is present in most HBs, 7 we surveyed the expression of this transcription factor in a larger set of NL (n = 17) and HB (n = 24) samples using two different methods: RNA in situ hybridization and immunostaining. Hepatocytes in NL expressed low or negligible amounts of GATA4 mRNA (Figure 1(a) and (b)) and protein (Table 1, Figure 1(c) and (d)). Consistent with our previous findings, 7 sinusoidal endothelial cells and Kupffer cells expressed GATA4 and served as internal controls in the analyses. Whereas GATA4 expression was insignificant in NL hepatocytes, GATA4 was highly expressed in the majority (58%) of HB tumors at both the mRNA and protein levels (Table 1, Figure 1(e)–(h)). Only 17% of the HB specimens were devoid of GATA4 immunoreactivity. High GATA4 expression localized to portions of tumor with an embryonal histology. There was no correlation between GATA4 immunoreactivity level and the PRETEXT group or initial serum alpha-fetoprotein level.
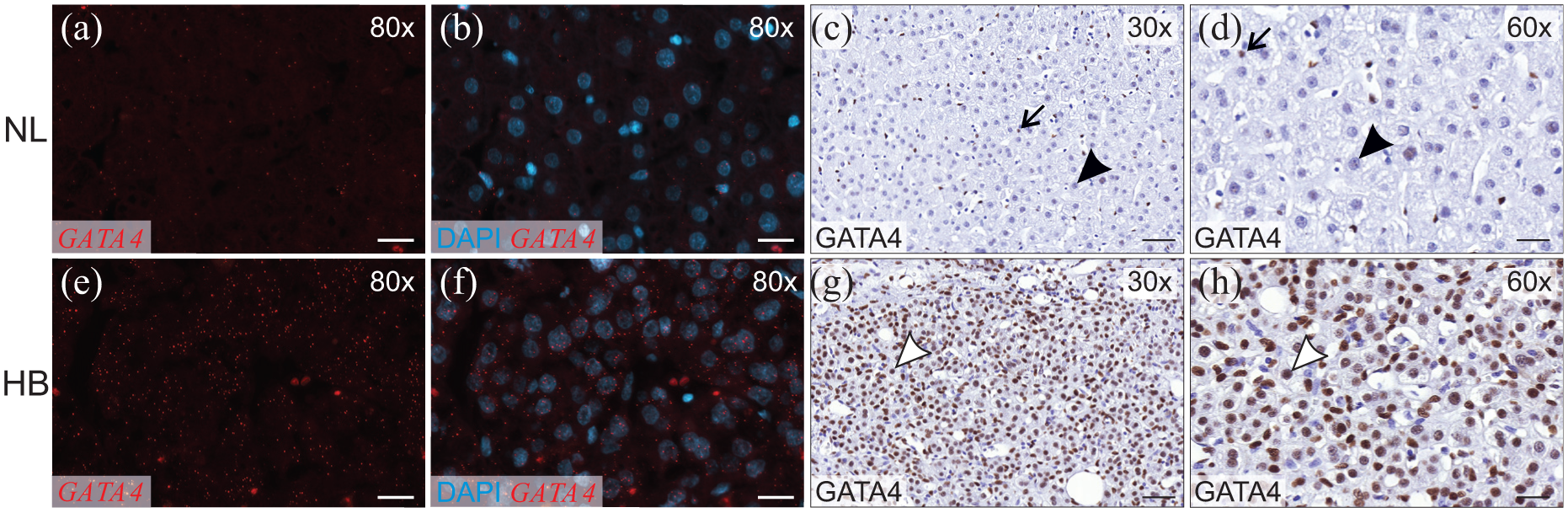
GATA4 expression in normal liver (NL) and hepatoblastoma (HB). GATA4 was detected with RNA in situ hybridization and immunohistochemistry in human liver samples. RNA in situ hybridization: red indicates positive signal of GATA4, and blue indicates DAPI staining of the nuclei. Immunohistochemistry: Brown indicates positive signal of GATA4 in nuclei. (a–d) Normal hepatocytes (black arrowhead) show weak or negative signal of GATA4 mRNA (a and b) and protein (c and d). Kupffer cells (arrow) are positive for GATA4 and serve as internal controls. (e–h) HB cells (white arrowhead) are highly positive for GATA4 at mRNA (e and f) and protein level (g and h). Scale bar: 100 µm.
GATA4 immunoreactivity in normal liver (NL) and hepatoblastoma (HB) samples.

Silencing of GATA4 in the HUH6 HB cell line
siRNA transfection was used to suppress GATA4 expression in HUH6 cells. Silencing efficiency was verified by qPCR. In all experiments, GATA4 mRNA levels were reduced by a minimum of 80% in GATA4 siRNA-treated cells when compared to controls (Supplementary Figure 1(a)). GATA4 reduction was also documented at the protein level with Western blotting (Supplementary Figure 1(b)). Immunofluorescence staining confirmed markedly decreased GATA4 immunoreactivity in the nuclei of GATA4 siRNA-targeted cells (Supplementary Figure 1(c)–(f)).
GATA4 silencing impairs migration of HUH6 cells
The impact of GATA4 silencing on HUH6 cell migration was determined using two complementary methods. In a wound healing assay, the area of migrating cells was 2.6 times smaller in GATA4-silenced cells compared to NT control cells (Figure 2(a)–(e)). In transwell assay, the number of migrating cells was 1.8 times lower in GATA4-silenced cells compared to NT control cells (Figure 2(f)–(h)). These results reinforce the hypothesis that GATA4 expression enhances the migratory capacity of HB cells. We previously demonstrated that HUH6 cell proliferation is unaffected by GATA4 silencing, 7 implying that the wound healing and transwell assay results reflect altered cell migration rather than impaired cell growth.

GATA4 deficiency reduces migration of HUH6 cells. (a–e) Wound healing assay of HUH6 cells shows 2.6-fold (e) larger area of migrating cells after 24 h in control (a, c) versus GATA4 siRNA treated (b and d) cells. Area of migrating cells is presented relative to NT siRNA as the mean ± SD of three independent experiments performed in triplicate. **p < 0.01. (f–h) Transwell assay verifies this result with 1.8-fold (h) greater number of migrated cells in NT (F) versus G4 (g) siRNA-treated HUH6 cells. Number of migrating cells is presented relative to NT siRNA as the mean ± SD of three independent experiments performed in triplicate. *p < 0.05. NT: nontarget siRNA; G4: GATA4 siRNA. Magnification: 10×.
Impact of GATA4 silencing on the HUH6 transcriptome
We used RNA microarray hybridization to assess global changes in gene expression associated with GATA4 silencing in HUH6 cells (Figure 3). We identified 106 genes that were differentially expressed in GATA4 silenced versus control cells. A total of 79 probes (72 genes) were downregulated, and 34 probes (34 genes) were upregulated. The data have been deposited in the NCBI Gene Expression Omnibus 20 and are accessible through GEO Series accession number GSE110788. Results were ranked according to their log 2 fold change (FC) values. To identify biological processes associated with the inhibition of GATA4 in HUH6 cells, we performed Gene Ontology (GO) analysis 21 with all differentially expressed genes. A substantial proportion of the altered biological processes were related to epithelial-mesenchymal balance (i.e. cell polarity, migration, and adhesion to adjacent cells or substrate; Table 2).

GATA4 silencing alters the gene expression profile in HUH6 cells toward epithelial phenotype. Heatmap of the 50 most differentially expressed genes in RNA microarray hybridization performed on GATA4 siRNA (G4) and control nontarget (NT) siRNA-treated cells.
Gene set enrichment analysis of GATA4 silenced HUH6 microarray data.
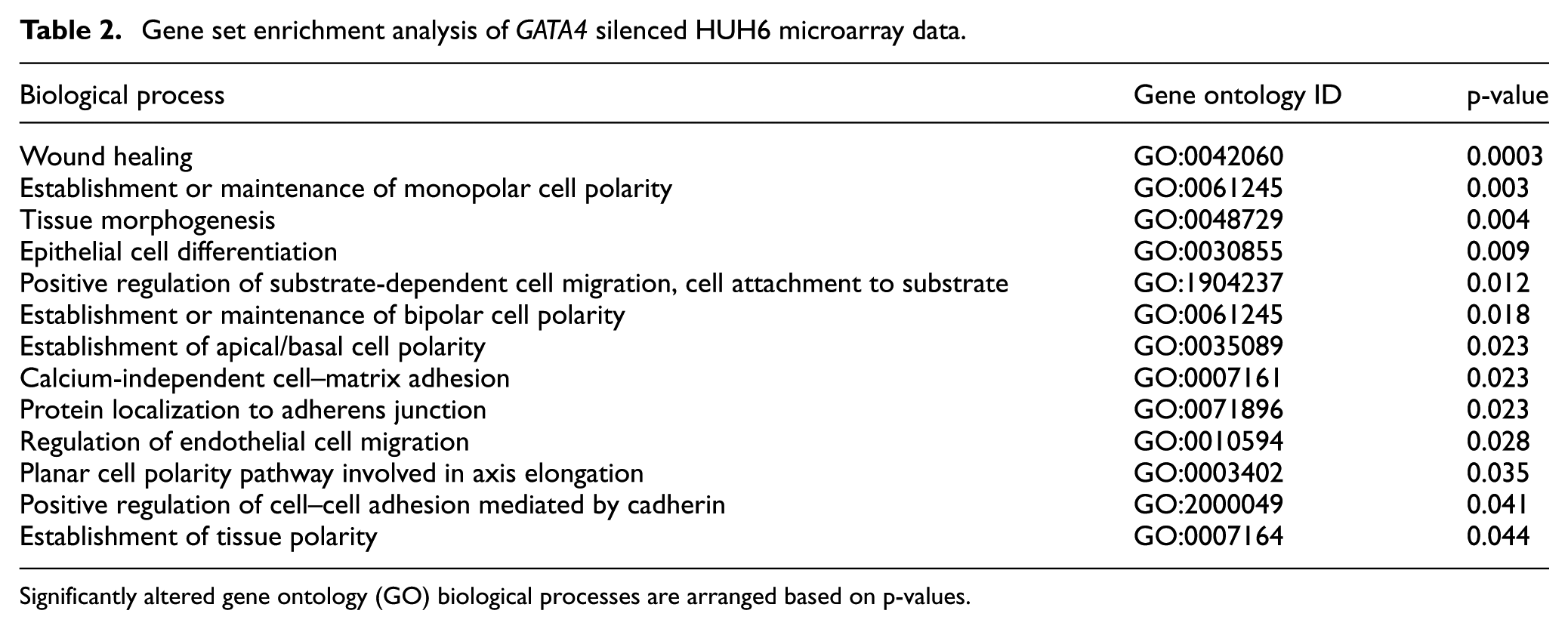

Significantly altered gene ontology (GO) biological processes are arranged based on p-values.
GATA4 affects the expression of genes involved in cytoskeleton remodeling and cell migration
To validate the microarray results, we performed qPCR analysis on selected genes that were differentially expressed in GATA4 silenced versus control HUH6 cells. In addition, we analyzed expression of other genes important for epithelial-mesenchymal transition (EMT) or with previous connection to GATA4. We verified altered expression of several genes regulating actin dynamics and cell migration. Downregulated genes included ADD3 (Adducin3), AHNAK (Ahnak nucleoprotein), DOCK8 (Dedicator of Cytokinesis 8), RHOU (Ras homolog family member U), and SYTL2 (Synaptotagmin-like 2; Figure 4(a)–(e)), all known to promote cell motility and EMT in other cell types.22–28 Although not detected in the microarray, qPCR analysis showed that SRC, a proto-oncogene tyrosine-protein kinase crucial for promoting carcinogenesis and EMT,28,29 was significantly downregulated in HUH6 cells treated with GATA4 siRNA (Figure 4(f)). On the other hand, the EMT inhibitor and tumor suppressor RHOB (Ras homolog family member B) was upregulated in response to GATA4 silencing in both microarray and qPCR (Figure 4(g)).

Alterations in genes involved in cytoskeleton reorganization and cell migration after GATA4 silencing in HUH6 cells. (a–g) RNA microarray hybridization and/or qPCR revealed altered expression of several genes affecting actin dynamics and cytoskeleton reorganization. mRNA expression is presented relative to NT siRNA transfection as the mean ± SD of three independent experiments performed in triplicate. *p < 0.05, **p < 0.01. NT: nontarget siRNA; G4: GATA4 siRNA. (h and i) Immunofluorescence staining of F-actin demonstrates reduced stress fiber formation and relocalization of F-actin filaments to cortical regions of the cell after GATA4 silencing (i) compared to control cells (h). Scale bar: 50 µm. (j and k): Close-up images of (h) and (i). Arrowhead points to a stress fiber and arrow indicates the cortical, epithelial-like localization of F-actin. Scale bar: 100 µm.
GATA4 regulates the organization of filamentous actin cytoskeleton
To visualize the morphological alterations in the filamentous actin (F-actin) cytoskeleton, GATA4-silenced and control HUH6 cells were stained with phalloidin-FITC-conjugated antibody. Control HUH6 cells exhibited actin stress fiber formation, indicating a motile, mesenchymal-like phenotype (Figure 4(h) and (j)). This phenotype was abolished in GATA4-silenced cells (Figure 4(i) and (k)). GATA4 downregulation also led to re-localization and reorganization of actin fibers to the cortical region of the cells, a distribution typical of immobile epithelial cells (Figure 4(i) and (k)).
GATA4 promotes cadherin switch in HUH6 cells
As cadherin-encoding genes play important roles in cancer cell motility, 30 we measured their expression in GATA4-depleted HUH6 cells. qPCR analyses demonstrated a significant increase in expression of E-cadherin coding gene CDH1 and concomitant downregulation of N-cadherin coding CDH2 (Figure 5(a) and (b)) after GATA4 silencing, indicating that GATA4-depleted cells are more adhesive and adopt an epithelial-like phenotype. These changes in cadherin expression were validated at the protein level using Western blotting (Figure 5(c)) and immunofluorescence microscopy (Figure 5(d)–(g)).

GATA4 induces cadherin switch in HUH6 cells. (a and b) mRNA expression of CDH1 encoding epithelial type E-cadherin is increased, whereas CDH2 encoding mesenchymal type N-cadherin is reduced (b) after GATA4 siRNA. mRNA expression is presented relative to NT siRNA transfection as the mean ± SD of three independent experiments performed in triplicate. *p < 0.05, **p < 0.01. (c) Western blotting shows increased expression of E-cadherin and reduced expression of N-cadherin after GATA4 siRNA treatment. GAPDH served as loading control. (d–g) Immunofluorescence staining demonstrates a reversed cadherin switch induced by GATA4 silencing in HUH6 cells. The cell membranous (arrowhead) E-cadherin expression is increased (d and e) and N-cadherin expression is reduced (f and g). Scale bar: 50 µm. NT: nontarget siRNA; G4: GATA4 siRNA.
Based on these in vitro findings, we next assessed the expression of E-cadherin and N-cadherin in GATA4-positive HB tumor specimens. In three HBs with high GATA4 immunoreactivity, we found a reversed expression of E- and N-cadherin compared to NL (Supplementary Figure 2).
GATA4 silencing in HUH6 cells alters the expression of genes involved in ECM remodeling
Microarray and/or qPCR analysis showed that GATA4 silencing in HUH6 cells also affects the expression of genes involved in the ECM remodeling and cell-to-ECM adhesion. In addition to the differentially expressed genes detected by microarray, we assessed the expression of several other ECM-related genes by qPCR. Two of the downregulated genes, MSF (migration-stimulating factor; a truncated isoform of fibronectin 1) and IGFBP1 (insulin-like growth factor binding protein 1), are both known promoters of EMT and migration (Figure 6(a) and (b)).31,32 The ECM remodeling proteins TIMP2 (tissue inhibitor of metalloproteinases 2) and MMP1 (matrix metalloproteinase 1) were also downregulated (Figure 6(c) and (d)). Epithelial basement membrane component COL4A2 (collagen, type IV, alpha 2) was upregulated after GATA4 silencing (Figure 6(e)).
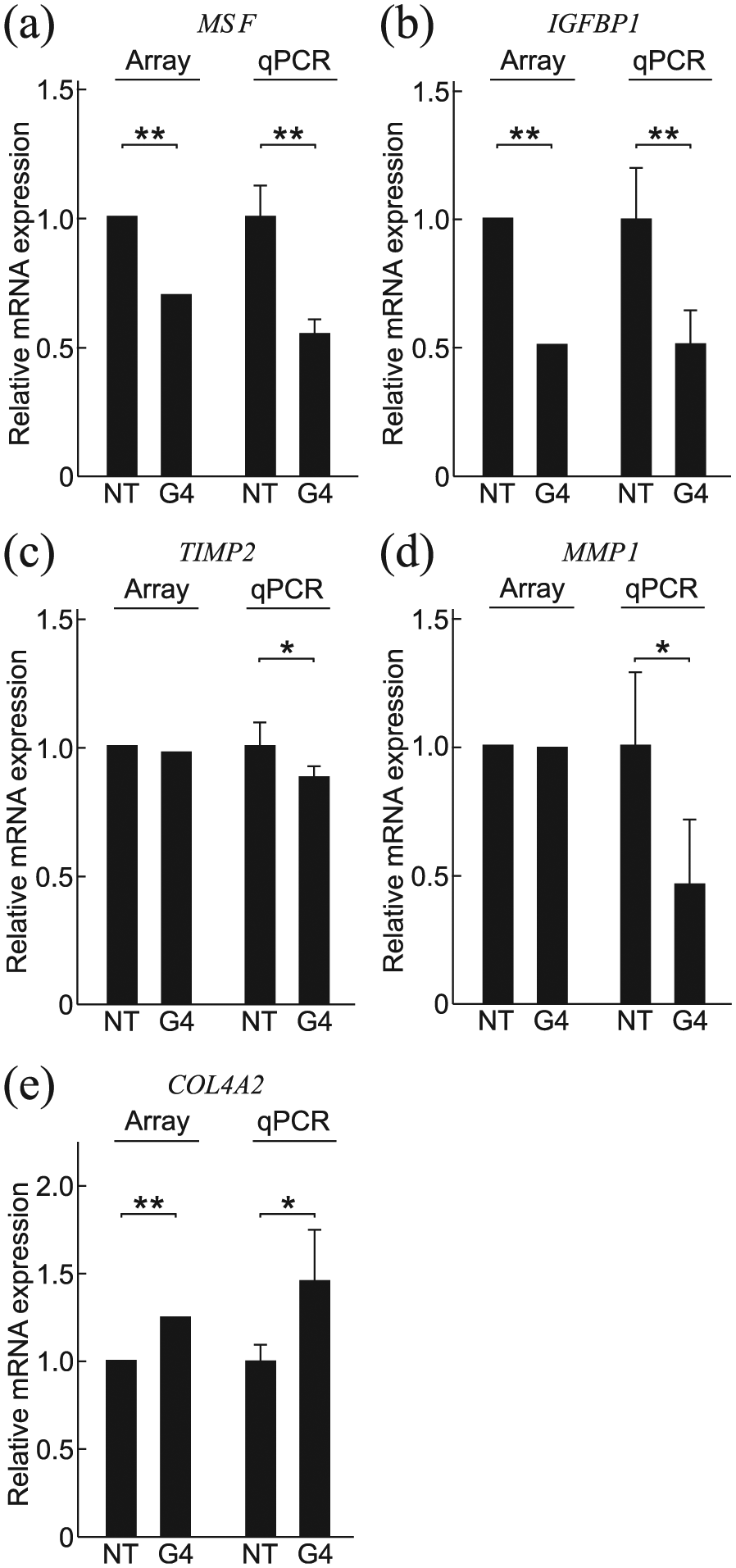
Differential expression in genes involved in ECM remodeling after GATA4 silencing in HUH6 cells. (a–e) RNA microarray hybridization and/or qPCR of control (NT) versus GATA4 silenced (G4) HUH6 cells show altered expression of several genes involved in ECM remodeling or cell-ECM contacts. mRNA expression is presented relative to NT siRNA transfection as the mean ± SD of three independent experiments performed in triplicate. *p < 0.05, **p < 0.01. NT: nontarget siRNA; G4: GATA4 siRNA.
Reduced GATA4 expression leads to altered gene expression in HepG2 cells
To confirm and extend the analysis, we silenced GATA4 in another HB-derived human cell line HepG2. After silencing GATA4, we assessed the expression of the same set of genes identified in the analysis of HUH6 cells. We found significant changes in the expression of 8 of 12 genes (ADD3 (1.50-fold, p = 0.005), AHNAK (0.72-fold, p = 0.005), CDH1 (2.20-fold, p = 0.002), CDH2 (1.39-fold, p = 0.007), RHOB (1.55-fold, p = 0.0001), MMP1 (2.37-fold, p = 0.0008), SRC (1.26-fold, p = 0.025), and IGFBP1 (0.70-fold, p = 0.003)) (data not shown). In four of these cases, the change in expression was in the same direction as in HUH6 cells.
Forced expression of GATA4 in cultured human primary hepatocytes shifts the gene expression pattern toward that of a mesenchymal phenotype
Next, we examined the impact of GATA4 overexpression on epithelial-mesenchymal balance in normal hepatocytes with a low endogenous level of this transcription factor (the baseline expression of GATA4 mRNA was 4.3-fold lower in primary human hepatocytes compared to HUH6 cells; p = 0.0002). Transduction of primary human hepatocytes with a GATA4 expression plasmid led to a 229% (p = 0.007) increase in the level of GATA4 mRNA in these cells and concomitant changes in genes affecting cytoskeleton reorganization and cell motility (AHNAK, RHOU, SYTL2, SRC, and RHOB; Figure 7(a)), the cadherin switch (CDH1 and CDH2; Figure 7(b)), and ECM remodeling (MSF, TIMP2, and MMP1; Figure 7(c)). The expression of ADD3, DOCK8, IGFBP1, and COL4A2 remained unaltered (data not shown). The changes in gene expression (elicited by overexpression of GATA4) in primary human hepatocytes were opposite to those triggered by GATA4 silencing in HUH6 cells, reinforcing the premise that GATA4 promotes a mesenchymal phenotype.

GATA4 overexpression alters gene expression in human primary hepatocytes. (a–c) Plasmid-mediated overexpression of GATA4 causes significant changes in the expression of genes regulating cytoskeleton reorganization (a), cadherin switch (b), and ECM remodulation (c). mRNA expression is presented relative to control plasmid transfection as the mean ± SD of five independent experiments performed in triplicate. *p < 0.05, **p < 0.01. Ctrl: control plasmid; G4↑: GATA4 expression plasmid.
Discussion
In this study, we have documented changes in migration, morphology, and gene expression of human HUH6 HB cells following depletion of GATA4, a transcription factor frequently overexpressed in human HBs. We found that a major proportion of the genes altered by GATA4 silencing in HUH6 cells are involved in the process of cell motility. An inverse pattern of gene expression was seen in normal hepatocytes following forced overexpression of this transcription factor. Based on these results, we suggest that GATA4 promotes a migratory, mesenchymal phenotype in HB cells.
GATA transcription factors have been linked to the regulation of cell polarity and movement, intercellular junctions, and ECM remodeling in several cell types. For example, GATA4 and GATA6 are known to promote EMT in human liver sinusoidal endothelial cells and in canine kidney (MDCK) cell line by reducing E-cadherin expression.33,34 GATA4 has been implicated in ECM remodeling in murine granulosa 35 and Sertoli cells. 36 GATA6 is required for the functional polarization of peritoneal macrophages. 37
Remodeling of the actin cytoskeleton is essential for cell migration, and genes related to actin dynamics were differentially expressed in GATA4 knockdown HUH6 cells. ADD3, AHNAK, and SYTL2 are essential for tumor cell migration and invasion in several cancer cell types.22–24,38 RHOU and RHOB are members of the Rho family of GTPases, which regulate epithelial apico-basal polarity and actin dynamics. RHOU is also an inducer of EMT and is overexpressed in several liver cancer cell lines, including HUH6.25,39 RHOB in turn is a tumor suppressor gene, loss of which induces migration and invasion.26,27,40 Proto-oncogene tyrosine-protein kinase SRC has multiple roles in promoting a malignant and mesenchymal phenotype in many types of cancer, as its expression induces weakening of cadherin-dependent cell–cell adhesion, loss of actin organization, and cancer cell migration (reviewed in Frame 29 ). SRC has been linked to HB cell migration and pathogenesis. 41 Moreover, SRC has been shown to regulate the expression of DOCK8, an EMT-promoting gene, in hepatocellular carcinoma cells. 28 We observed downregulation of both SRC and DOCK8 after GATA4 silencing in HUH6 cells. Taken together, the alterations in HUH6 gene expression after GATA4 silencing suggest that GATA4 plays a central role in cytoskeletal reorganization, contractility, and motility of HB cells.
Reduction of cell adhesion and loosening of intercellular junctions are necessary for cell migration. A hallmark molecular event of EMT is the cadherin switch, that is, downregulation of E-cadherin and concomitant upregulation of mesenchymal cell marker N-cadherin. N-cadherin forms weaker adhesions with adjacent cells than E-cadherin and facilitates migration by forming contacts to ECM. 42 The cadherin switch leads to a collective type of migration, in which the cells move forward in one front instead of migrating as single fibroblast-like cells. 43 Collective migration is also evident in normal HUH6 cells. We demonstrated a reduced migration and a reversed cadherin switch leading to a more epithelial phenotype in GATA4-silenced HUH6 cells. These findings are in line with previous reports demonstrating GATA4 as regulator of both E- and N-cadherin expression and localization in other cell types.34,35 Importantly, in a subset of HBs, we found that the GATA4-positive areas with embryonal histology exhibited a mesenchymal-type cadherin expression profile. This finding suggests that GATA4 may also regulate the cadherin balance in vivo.
Concurrently with achieving an induced migratory capacity and reduced adhesion to adjacent cells, ECM remodeling enables the movement of cancer cells away from their original location. Several ECM-related genes were differentially expressed after GATA4 silencing in HUH6 cells. Among the downregulated genes were MSF, IGFBP1, MMP1, and TIMP2. MSF is one of the oncofetal isoforms of FN1, a key marker of mesenchymal cells, linked to cancer metastasis and the EMT process.31,44 IGFBP1 has several IGF-receptor-independent roles in cancer progression, including growth, migration, and modulation of cell-ECM adhesions,45,46 and it is known to activate the SRC/FAK complex, also important for epithelial-mesenchymal balance in HB tumors.41,45 MMPs are ECM-degrading proteins secreted by the cells undergoing EMT, and their activity is regulated by TIMPs. MMP1 is frequently overexpressed in malignant tumors and regulated by GATA4 in canine kidney cells.34,47 Silencing Gata4 induces Timp2 downregulation in murine Sertoli cells 36 further supporting the role of GATA4 in tissue remodeling. Epithelial basement membrane component, COL4A2, was upregulated in HUH6 cells after GATA4 silencing. This upregulation is connected to EMT process in other cancer cell types,48,49 and Gata4 knockdown has been shown to upregulate Col4A2 expression also in murine liver sinusoidal endothelial cells. 33 Collectively, the observed changes in ECM and basement membrane–related genes support the role of GATA4 in promoting cancer cell migration and invasion.
To confirm our findings in HUH6 cell, we studied HepG2, another malignant liver cell line, to investigate the expressional changes induced by GATA4 silencing between different HB types. The HepG2 cell line was derived from a liver tumor that arose in an adolescent and has frequently been referred to as a hepatocellular carcinoma cell line, as this liver tumor type is much more common in older children. However, HepG2 was recently re-characterized and confirmed to represent an epithelial-type HB. 50 GATA4 silencing in HepG2 cells altered the expression of only a subset of the genes dysregulated by GATA4 silencing in HUH6 cells. This discrepancy may be due to the more epithelial-like character of HepG2 cells.
Expression of GATA4 is lower in normal postnatal hepatocytes than in HB cells (Soini et al. 7 and this study). To explore whether GATA4 can alter the pattern of gene expression in normal adult hepatocytes, we overexpressed GATA4 in human primary hepatocytes. Our findings suggest that in addition to its effects in tumor cells, GATA4 can induce the expression of several genes related to malignant phenotype.
Based on these findings, we hypothesize that GATA4 is an important regulator of migration in HB cells. It is not known, however, whether GATA4 is a direct regulator of the genes identified in this study. To address this issue, chromatin immunoprecipitation experiments and subsequent transactivation assays in HB cells need to be performed.
Taken together, our results demonstrate that high GATA4 expression favors a motile, mesenchymal-like phenotype of cultured HB cells. By inference, GATA4 expression may affect tumor metastatic capacity in vivo. Our findings shed new light on the molecular pathogenesis of HB and provide an avenue to develop novel treatments such as miRNA- and siRNA-based approaches, recently suggested in HBs and other liver diseases.51,52
Supplemental Material
Supplemental_table_1_(2) – Supplemental material for Transcription factor GATA4 associates with mesenchymal-like gene expression in human hepatoblastoma cells
Supplemental material, Supplemental_table_1_(2) for Transcription factor GATA4 associates with mesenchymal-like gene expression in human hepatoblastoma cells by Tea Soini, Katja Eloranta, Marjut Pihlajoki, Antti Kyrönlahti, Oyediran Akinrinade, Noora Andersson, Jouko Lohi, Mikko P Pakarinen, David B Wilson and Markku Heikinheimo in Tumor Biology
Supplemental Material
Supplementary_figure_1 – Supplemental material for Transcription factor GATA4 associates with mesenchymal-like gene expression in human hepatoblastoma cells
Supplemental material, Supplementary_figure_1 for Transcription factor GATA4 associates with mesenchymal-like gene expression in human hepatoblastoma cells by Tea Soini, Katja Eloranta, Marjut Pihlajoki, Antti Kyrönlahti, Oyediran Akinrinade, Noora Andersson, Jouko Lohi, Mikko P Pakarinen, David B Wilson and Markku Heikinheimo in Tumor Biology
Supplemental Material
Supplementary_figure_2 – Supplemental material for Transcription factor GATA4 associates with mesenchymal-like gene expression in human hepatoblastoma cells
Supplemental material, Supplementary_figure_2 for Transcription factor GATA4 associates with mesenchymal-like gene expression in human hepatoblastoma cells by Tea Soini, Katja Eloranta, Marjut Pihlajoki, Antti Kyrönlahti, Oyediran Akinrinade, Noora Andersson, Jouko Lohi, Mikko P Pakarinen, David B Wilson and Markku Heikinheimo in Tumor Biology
Footnotes
Acknowledgements
We thank Dr Gregory Longmore for comments and suggestions. Tea Soini, Katja Eloranta, and Marjut Pihlajoki contributed equally to this study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Academy of Finland; Finnish Pediatric Research Foundation; Helsinki University Central Hospital Research Grants, Finland; Sigrid Jusélius Foundation, Finland; and Pediatric Cancer Foundation Väre, Finland.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
