Abstract
In this study, we aimed to explore new downstream effectors of TWIST1/2 in inducing epithelial-to-mesenchymal transition in gastric cancer. Bioinformatic data mining was performed using data in The Cancer Genome Atlas Stomach Adenocarcinoma. Survival curves were generated using Kaplan–Meier plotter. Gastric cancer cell lines (AGS and SGC-7901) were used as in vitro cell model to investigate the regulative effect of TWIST1/2 on epithelial membrane protein 3 expression and the progression of epithelial-to-mesenchymal transition. Results showed that TWIST1 and TWIST2 are usually co-upregulated in patients with primary gastric cancer. High TWIST1 expression is associated with worse overall survival (hazard ratio = 1.26; 95% confidence interval = 1.06–1.49; p = 0.007) and also worse first progression-free survival (hazard ratio = 1.47; 95% confidence interval = 1.18–1.82; p < 0.0001). Similarly, high TWIST2 expression is associated with unfavorable overall survival (hazard ratio = 1.71; 95% confidence interval = 1.32–2.22; p < 0.0001) and progression-free survival (hazard ratio = 1.99; 95% confidence interval = 1.45–2.72; p < 0.0001). Epithelial membrane protein 3 is negatively correlated to CDH1 expression (Pearson’s r = −0.46) but is positively correlated to VIM expression (Pearson’s r = 0.83). Knockdown of epithelial membrane protein 3 significantly increased E-cadherin but significantly decreased Vimentin expression in AGS cells. Gastric cancer patients with metastasis have significantly higher epithelial membrane protein 3 expression than the cases without metastasis. In addition, high epithelial membrane protein 3 expression is associated with worse overall survival (hazard ratio = 2.59; 95% confidence interval = 2.06–3.26; p < 0.0001) and also worse progression-free survival (hazard ratio = 2.21; 95% confidence interval = 1.78–2.74; p < 0.0001). In conclusion, epithelial membrane protein 3 is a downstream effector of TWIST1/2 in inducing epithelial-to-mesenchymal transition in gastric cancer. Epithelial membrane protein 3 upregulation might be associated with gastric cancer metastasis and is a potential indicator of unfavorable overall survival and progression-free survival in gastric cancer patients.
Introduction
Gastric cancer (GC) is a highly invasive and aggressive malignancy and is one of the leading causes of cancer-related deaths across the world.1,2 Metastasis is an important indicator of poor prognosis of GC.3–5 However, the mechanisms underlying metastasis are quite complex and are far from been fully understood. Among the recognized mechanisms, epithelial-to-mesenchymal transition (EMT) is considered as one of the fundamental changes of tumor cells to gain stronger migration and invasion potential.6–8 TWIST1 and TWIST2 are two genes encoding twist1 and twist2, two basic helix-loop-helix (bHLH) transcription factors. 9 These two transcription factors have critical roles in the progression of EMT in multiple types of cancer, such as pancreatic cancer, 10 cervical cancer 11 and GC. 12 As two similar transcriptional factors, twist1 and twist2 may form homodimer or heterodimer and regulate the expression of a series of genes. 13 The known downstream effectors of twist1 in GC include FoxM1, 14 FGFR2, 15 and Siah2. 16 However, less is known about the downstream effectors of twist2 in GC.
The epithelial membrane protein 3 (EMP3) is amyelin-related gene that belongs to the peripheral myelin protein 22-kDa (PMP22) gene family of small hydrophobic membrane glycoproteins. 17 It has paradoxical roles in different types of cancer. Some studies found that EMP3 might act as a tumor suppressor gene in low-grade glioma, 18 esophageal squamous cell carcinoma, 19 and non–small cell lung cancer. 20 In comparison, it also showed oncogenic effect in glioblastoma 21 and hepatocellular carcinoma. 22 However, its functional role in GC is not yet explored.
In this study, we found that TWIST1 and TWIST2 are co-upregulated with EMP3 in GC and can induce its upregulation in GC cells. In addition, high EMP3 expression is associated with EMT of GC cells and is also associated with unfavorable overall survival (OS) and first progression-free survival (PFS) in GC patients.
Materials and methods
Cell culture and transfection
The human GC cell lines SGC-7901 and AGS were obtained from American Type Culture Collection (Manassas, VA, USA). The cancer cells were cultured in RPMI-1640 medium supplemented with 10% fetal bovine serum (FBS), 100 μg/mL penicillin, and 100 U/mL streptomycin. EMP3 small interfering RNA (siRNA; sc-97634) was obtained from Santa Cruz Biotechnology (Santa Cruz, CA, USA). AGS cells were transfected with 50 nM of EMP3 siRNA or the scramble negative control using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). After 24 h, the cells transfected with siRNA were harvested for quantitative real-time polymerase chain reaction (qRT-PCR) to determine the inhibiting efficiency. TWIST1 and TWIST2 Lentifect™ purified lentiviral particles and the empty controls were obtained from GeneCopoeia (Rockville, MD, USA). Cells were infected with the lentiviral particles in the presence of polybrene.
Data mining in The Cancer Genome Atlas Stomach Adenocarcinoma
The messenger RNA (mRNA) expression of TWIST1, TWIST2, EMP3, CDH1, and VIM in The Cancer Genome Atlas Stomach Adenocarcinoma (TCGA-STAD) was analyzed using UCSC Xena Browser. Regression analysis was performed between TWIST1 and EMP3, between TWIST2 and EMP3, between EMP3 and CDH1, and between EMP3 and VIM.
Data mining in Kaplan–Meier plotter
The prognostic value of TWIST1, TWIST2, and EMP3 mRNA expression was evaluated using Kaplan–Meier plotter, an online database containing gene expression data and survival information of 1065 GC patients. 23 OS and PFS were calculated by splitting the patients into high and low expression groups, according to the best performing threshold. The hazard ratio (HR) with 95% confidence intervals (CIs) and log-rank p value were calculated. The number at risk is indicated below the survival curves.
qRT-PCR analysis
RNA was extracted from cells using the RNA Mini Kit (QIAGEN, Valencia, CA, USA) according to the manufacturer’s instructions. Then, the RNA samples were used as template for first-strand complementary DNA (cDNA) synthesis using the AffinityScript cDNA kit (Stratagene, La Jolla, CA, USA). qRT-PCR was then performed using the Power SYBR Green PCR Master Mix (Applied Biosystems, Foster City, CA, USA) with gene-specific primers and a standard thermal cycling procedure on an ABI 7900HT instrument (Applied Biosystems). The primers used were as follows—TWIST1: 5′-GGACAAGCTGAGCAAGATTCA-3′ (forward) and 5′-CGGAGAAGGCGTAGCTGAG-3′ (reverse), TWIST2: 5′-ACGAGCGTCTCAGCTACGCC-3′ (forward) and 5′-AGGTGGGTCCTGGCTTGCGG-3′ (reverse), and EMP3: 5′-ACGAGCGTCTCAGCTACGCC-3′ (forward) and 5′-GCATAGAAGAGACCTCCT-3′ (reverse). RNA quantity was normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH) and was calculated according to the 2−ΔΔCt method.
Western blotting
Whole cell extracts were prepared using radioimmunoprecipitation assay (RIPA) lysis buffer (Santa Cruz Biotechnology). A volume of 30 ng of the protein was blotted onto nitrocellulose filters after sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE). Then, the membranes were incubated with anti-twist1, anti-twist2, anti-EMP3, or anti-β-actin (Abcam, Cambridge, UK). After that the membranes were further incubated with horseradish peroxidase–conjugated secondary antibody. After washing, the protein bands were visualized using the enhanced chemiluminescence (ECL) system (Amersham Biosciences, San Francisco, CA, USA).
Immunofluorescent staining
AGS cells with or without knockdown of EMP3 were grown on coverslips. After overnight incubation, the cells were fixed, permeabilized with 0.1% Triton X-100, and blocked with 1% bovine serum albumin (BSA). The coverslips were incubated with anti-E-cadherin (ab15148; Abcam) or anti-Vimentin (ab92547; Abcam) at 4°C for overnight. After washing, the coverslips were further incubated with anti-rabbit Alexa Fluor® 488 conjugated secondary antibody (#4412; Cell Signaling Technology, Danvers, MA, USA) or anti-rabbit Alexa Fluor® 555 conjugated secondary antibody (#4413; Cell Signaling Technology) for 1 h at room temperature in the dark. Nuclei were stained using Prolong® Gold Antifade Reagent with 4′,6-diamidino-2-phenylindole (DAPI; #8961; Cell Signaling Technology). Immunofluorescent images were obtained using an Olympus IX73 inverse microscope (Tokyo, Japan).
Statistical analysis
Data were reported as mean ± standard deviation (SD). Group difference was assessed by two-tailed Student’s t test or analysis of variance (ANOVA) with Student–Newman–Keuls test as a post hoc test. p < 0.05 was considered to be statistically significant.
Results
High TWIST1/2 expression is associated with worse OS and PFS in GC patients
Previous studies found that aberrant expression of TWIST1 and/or TWIST2 is associated with malignant phenotypes of GC, such as enhanced invasion, poor differentiation, and fast disease progression.15,24 By retrieving data in TCGA-STAD, we characterized the expression of TWIST1 and TWIST2 in 415 patients with primary stomach adenocarcinoma (Figure 1(a)). TWIST1 and TWIST2 are usually co-upregulated in the patients (Figure 1(a)). To assess the association between the expression of TWIST1 or TWIST2 and survival outcomes, OS curves and PFS curves were generated using Kaplan–Meier plotter by setting the best performing threshold as cutoff. Log-rank test showed that high TWIST1 expression is associated with worse OS (HR = 1.26; 95% CI = 1.06–1.49; p = 0.007; Figure 1(b)) and also worse PFS (HR = 1.47; 95% CI = 1.18–1.82; p < 0.0001; Figure 1(c)). Similarly, high TWIST2 expression is associated with unfavorable OS (HR = 1.71; 95% CI = 1.32–2.22; p < 0.0001; Figure 1(d)) and PFS (HR = 1.99; 95% CI = 1.45–2.72; p < 0.0001; Figure 1(e)).

High TWIST1/2 expression is associated with worse OS and PFS in GC patients. (a) The heatmap of TWIST1 and TWIST2 expression in TCGA-STAD. Patients with primary tumors were screened for analysis. Kaplan–Meier curves of (b and d) overall survival (OS) and (c and e) first progression-free survival (PFS) in patients with high and low (b and c) TWIST1 or (d and e) TWIST2 expression. The curves were generated using Kaplan–Meier plotter.
TWIST1/2 induces EMP3 expression in GC
To further examine the downstream signaling pathways of TWIST1/2 in GC, we further investigated the genes co-expressed with TWIST1/2 in TCGA-STAD. Interestingly, we found that EMP3 is moderately co-upregulated with TWIST1 (Pearson’s r = 0.56), while EMP3 is strongly co-upregulated with TWIST2 (Pearson’s r = 0.70; Figure 2(a)–(c)). To further examine the regulative effect of TWIST1/2 on EMP3 expression, AGS and SGC-7901 cells were transfected with TWIST1 or TWIST2 expression vector, respectively (Figure 2(d) and (e)). In both the cell lines, TWIST1 or TWIST2 upregulation significantly elevated EMP3 expression (Figure 2(f) and (g)).

TWIST1/2 induces EMP3 expression in GC. (a) The heatmap of TWIST1, TWIST2, and EMP3 expression in TCGA-STAD. Patients with primary tumors were screened for analysis. Regression analysis of the expression between (b) TWIST1 and EMP3 and (c) between TWIST2 and EMP3 in the sample group in (a). qRT-PCR analysis of (d) TWIST1 or (e) TWIST2 expression in AGS and SGC-7901 cells after 24 h of infection of lentiviral TWIST1 or TWIST2 expression particles. Western blotting of Twist1, Twist2, and EMP3 expression in SGC-7901 and AGS cells after 48 h of infection of lentiviral (f) TWIST1 or (g) TWIST2 expression particles (**p < 0.01).
EMP3 inhibition partly reverses EMT in AGS cells
EMP3 is an important mesenchymal gene. As a downstream effector of TWIST1/2, we decided to further investigate its functional role in EMT of GC cells. In TCGA-STAD, we examined the correlation between EMP3 and CDH1 (E-cadherin) and between EMP3 and VIM (Vimentin). Heatmap and following regression analysis showed that EMP3 is negatively correlated to CDH1 expression (Pearson’s r = −0.46; Figure 3(a) and (b)) but is positively correlated to VIM expression (Pearson’s r = 0.83; Figure 3(a) and (c)). To further verify the regulative effect of EMP3 on EMT in GC cells, AGS cells were transfected with EMP3 siRNA (Figure 3(d)). Knockdown of EMP3 significantly increased E-cadherin but significantly decreased Vimentin expression in AGS cells (Figure 3(e)). The following immunofluorescent staining confirmed these changes (Figure 3(f)).

EMP3 inhibition partly reverses EMT in AGS cells. (a) The heatmap of EMP3, CDH1, and VIM expression in TCGA-STAD. Patients with primary tumors were screened for analysis. Regression analysis of the expression between (b) EMP3 and CDH1 and (c) between EMP3 and VIM in the sample group in (a). (d) qRT-PCR analysis of EMP3 expression in AGS cells after 24 h of transfection of EMP3 siRNA. (e) Western blotting of EMP3, E-cadherin, and Vimentin expression in AGS cells after 48 h of transfection of EMP3 siRNA. (f) Immunofluorescent staining of E-cadherin and Vimentin in AGS cells after 48 h of transfection of EMP3 siRNA (**p < 0.01).
High EMP3 expression might be associated with metastasis and unfavorable survival
EMT is one of the important mechanisms of tumor cell invasion and metastasis. By comparing EMP3 expression between M0 (without metastasis) cases and M1 (with metastasis) cases, we observed a significant upregulation of EMP3 in M1 group (Figure 4(a)). Then, we further assessed the association between EMP3 and OS/PFS using Kaplan–Meier plotter. Log-rank test showed that high EMP3 expression is associated with worse OS (HR = 2.59; 95% CI = 2.06–3.26; p < 0.0001; Figure 4(b)) and also worse PFS (HR = 2.21; 95% CI = 1.78–2.74; p < 0.0001; Figure 4(c)).
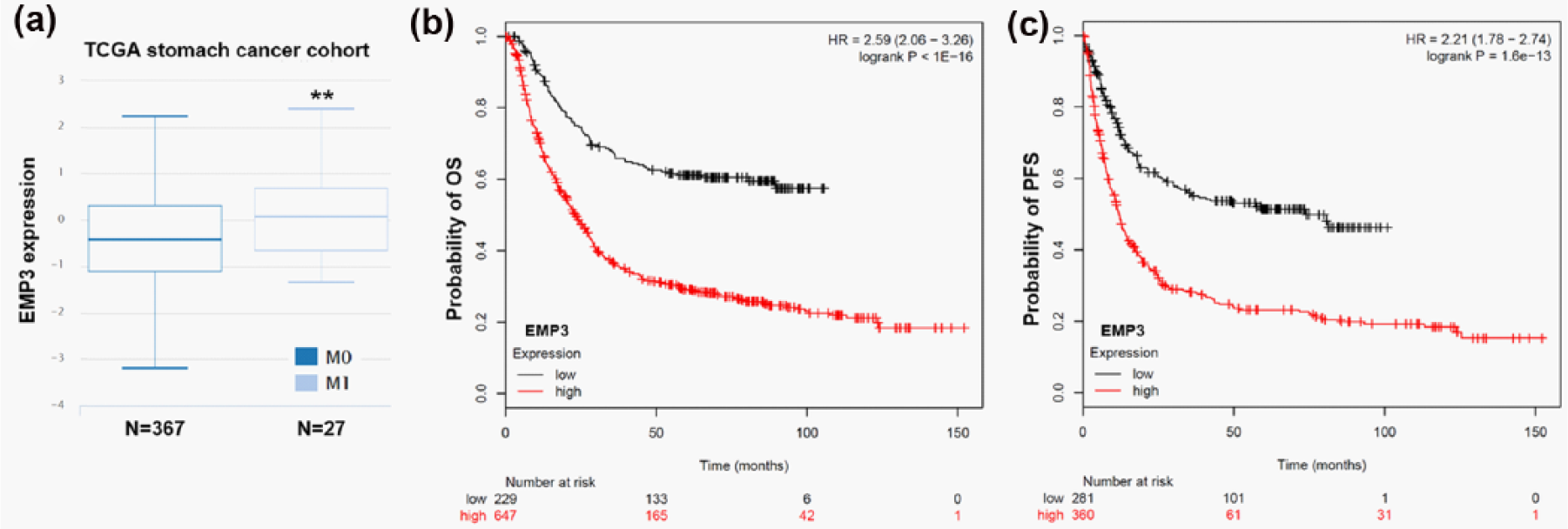
High EMP3 expression might be associated with metastasis and unfavorable survival. (a) Boxplot of the EMP3 expression in M0 and M1 gastric cancer cases. Kaplan–Meier curves of (b) OS and (c) PFS in patients with high/low EMP3 expression. The curves were generated using Kaplan–Meier plotter (**p < 0.01).
Discussion
Mechanistically, Twist1 and Twist2 can form homodimer or heterodimer, which together with other bHLH family members regulate target gene expression via binding to the E-box DNA sequence 5′-NCANNTGN-3′. 13 These two transcriptional factors can induce GC growth and development through multiple pathways. For example, Twist1 can directly bind to the promoter of FOXM1 and thereby enhance GC cell proliferation. 14 It can also facilitate invasion and EMT of GC cells via promotion of FGFR2 expression. 15 In Helicobacter pylori–infected GC cells, twsit1 can increase cell motility and invasiveness through Siah2 induction. 16 Aberrant expression of TWIST2 was also observed in GC and contributes to EMT of GC cells. 25 In this study, by data mining in TCGA-STAD, we observed a significant co-expression between TWIST1 and TWIST2. Since TWIST1 and TWIST2 can induce aggressive phenotype of GC cells, we further investigated the prognostic value of TWIST1 and TWIST2 in GC. By data mining in Kaplan–Meier plotter, we found that high TWIST1/2 expression is associated with worse OS and PFS in GC patients. This finding is in line with one previous study that found increased TWIST1 expression in GC-associated fibroblasts, which is associated with poor patient survival. 12
To further investigate the downstream effectors of TWIST1 and TWIST2 in GC, we identified their co-upregulated genes in TCGA-STAD. Interestingly, we found that EMP3 is moderately co-upregulated with TWIST1 and is strongly co-upregulated with TWIST2. In fact, EMP3 is an important mesenchymal gene 26 and may regulate EMT of some cancer cells. 27 By promoter scanning, we failed to identify any high score E-Box site in EMP3 promoter. However, in both AGS and SGC-7901 cells, we found that TWIST1 or TWIST2 overexpression significantly elevated EMP3 expression. Therefore, we infer that TWIST1/2 can induce EMP3 upregulation, but the exact mechanisms should be explored in future studies.
Although EMP3 was initially identified as a tumor suppressor in some types of solid tumors, some recent studies also revealed its oncogenic effect. For example, in glioblastoma cells, EMP3 can directly interact with transforming growth factor beta receptor 2 (TGFBR2) upon TGF-β stimulation, which subsequently activates TGF-β/Smad2/3 signaling activation and enhances cell proliferation in vitro and in vivo. 21 In upper urinary tract urothelial carcinoma, EMP3 can enhance cancer cell proliferation and migration via activating the ErbB2/phosphoinositide 3-kinase (PI3K)/AKT pathway. 28 Knockdown of EMP3 can suppress proliferation and invasion of hepatocellular carcinoma cells through inactivation of the PI3K/Akt pathway. 22 In oral squamous cancer cells, EMP3 could reduce the expression of p66Shc and enhance cell migration through the weakened E-cadherin and ZO-1 signals. 27 Considering the possible effect of EMP3 on modulating EMT, we compared the expression between EMP3 and the EMT markers in TCGA-STAD. Results showed that EMP3 is negatively correlated to CDH1 expression but is positively correlated to VIM expression. In vitro assay confirmed that knockdown of EMP3 significantly increased E-cadherin but significantly decreased Vimentin expression in AGS cells. Based on these findings, we infer that TWIST1/2 may induce EMT of GC cells via upregulating EMP3.
Till now, no studies investigated the prognostic value of EMP3 in GC. In this study, we further assessed the association between EMT expression and GC metastasis and survival outcomes. Our data showed that the GC patients with metastasis have significantly higher EMP3 expression than the cases without metastasis. In addition, high EMP3 expression is associated with worse OS and also worse PFS in the patients.
Conclusion
EMP3 is a downstream effector of TWIST1/2 in inducing EMT in GC. EMP3 upregulation might be associated with GC metastasis and is a potential indicator of unfavorable OS and PFS in GC patients.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
