Abstract
This study was conducted to detect the expression of miR-19 and Pax6 (Paired box protein 6) in human osteosarcoma cells and the effects on biological characteristics of osteosarcoma cells. Quantitative real-time polymerase chain reaction was used to detect the expression of Pax6 and miR-19 in normal human osteoblasts (hFOB 1.19) and osteosarcoma cell lines (U2OS, Saos-2, and MG-63). Results showed that miR-19 was significantly upregulated in osteosarcoma cell lines compared with that in hFOB 1.19 cells, while the expression of Pax6 messenger RNA was significantly downregulated. Pax6 was defined as the target gene of miR-19 which was validated by luciferase reporter gene analysis. Results indicated that miR-19 had an interaction with Pax6 3′-untranslated region. At the same time, the protein expression of Pax6 was significantly decreased in the MG-63 cells transfected with miR-19 mimic and was notably enhanced in osteosarcoma MG-63 cells transfected with miR-19 inhibitor. These data suggested that Pax6 was a target of miR-19 in osteosarcoma MG-63 cells. The effects of miR-19 on the biological behavior of MG-63 cells were determined by 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide, flow cytometry, and Transwell assay. Results showed that the downregulation of miR-19 inhibited cell viability, reduced the percentage of cells in S phase and the number of cells passing through the Transwell chamber, and increased the number of apoptotic cells. Western blot analysis showed that the inhibition of miR-19 significantly increased the expression of epithelial proteins (E-cadherin and β-catenin) and decreased the expression of mesenchymal protein (Vimentin), extracellular signal–regulated kinase, and phosphorylated extracellular signal–regulated kinase in MG-63 cells. MiR-19 inhibitor and Pax6 small interfering RNA were simultaneously transfected into MG-63 cells. Results from 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide, flow cytometry, and Transwell assay demonstrated that the inhibition of Pax6 expression in MG-63 cells could reverse the cell biological effects induced by the inhibition of miR-19 expression. Based on these findings, it was suggested that miR-19, upregulated in osteosarcoma cells, negatively regulated the expression of Pax6, which can promote the malignant phenotypes of osteosarcoma cells via activation of the extracellular signal–regulated kinase signaling pathways. Therefore, miR-19/Pax6 may offer potential for use as a target for the treatment of osteosarcoma.
Osteosarcoma is the most common primary malignant bone tumor in teenagers that occurs in the long bone metaphyseal, and the incidence rate of osteosarcoma is around 4 ppm.1,2 Although neoadjuvant chemotherapy, surgery, and other comprehensive treatment have made the 5-year survival rate of patients with osteosarcoma increase from 50% to 65% in the past 30 years, there are still one-third of all patients suffering from pulmonary metastasis. With the rapid development of molecular biology technology, people have gradually realized that malignant tumor is actually a genetic disease and its occurrence and development involve a variety of genetic changes. The genetic changes could cause functional changes of oncogenes and anti-oncogenes in normal cells, cell signal transduction blocked, cell-cycle disorder, and cell apoptosis inhibited, finally resulting in abnormal cell proliferation, invasions, blood vessel hyperplasia, and other malignant phenotypes.3–5
Distant metastasis is one of the most important causes for the failure of treatment in most patients with osteosarcoma.6–8 The principal cause of distant metastasis of osteosarcoma is the abnormal expression of tumor metastasis–related genes and metastasis suppressor genes.9,10 Researchers have carried out a lot of scientific researches on the mechanism of the occurrence and development of osteosarcoma but failed to clarify the pathogenesis of osteosarcoma. So, it still requires further researches on the pathogenesis of osteosarcoma to clarify the pathogenesis and to provide scientific basis for the diagnosis, treatment, and prognosis of osteosarcoma, so as to effectively prevent the progression of osteosarcoma, improve the prognosis of patients, and lighten the burden of the patient’s family and society.9,11,12
Recent studies have made microRNA (miRNA) rapidly develop into an important potential molecular marker for cancer and other diseases.13–15 With the ongoing study between individual miRNA, miRNA gene cluster, and the occurrence and development of tumor, miR-17-92 gene cluster is a group of miRNAs which are studied in detail. The miR-17-92 gene cluster belongs to a cancer gene cluster and is located in 13q31.3.16–18 The miR-17-92 gene cluster is a highly conserved gene cluster which can encode miR-17-5p, miR-18a, miR-19a, miR-20, miR-19b-1, and miR-92-1.19,20 A large number of studies have indicated that miRNAs in miR-17-92 gene cluster showed high expression in a variety of tumors and serves as oncogene.21,22 MiR-17-92 gene cluster showed high expression in carcinoma tissues and cells of esophageal and lung cancer. 23 At the same time, more and more downstream target genes of the cluster have been identified, including inhibiting cell apoptosis by suppressing expression of tumor necrosis factor-α (TNF-α) and promoting the metastasis of tumors by regulating the expression of Fas-FasL.24–26 In addition, recent study has reported a correlation between the upregulation of miR-17-92 gene cluster expression and the occurrence and development of osteosarcoma. 27 But the effect and mechanism of miR-19 on the cell biological characteristics of osteosarcoma cells is not very clear.
Pax6 is a member of the Pax multigene family of the paired-homeodomain transcription factor and plays essential roles in embryonic differentiation. Recent studies demonstrated that Pax6 was related to the development of the eye, central nervous system, endocrine gland tissues, and so on. It has persistent expression in the brain.28,29 Pax6 first gained attention as a tumor suppressor gene in glioma cells, which directly bound the promoter region and regulated the expression of target gene, and then inhibited the invasion of glioma cells.30,31 In this study, we analyzed through online software (http://www.targetscan.org) that Pax6 was likely to be the regulatory target gene of miR-19.
Therefore, the expression of miR-19 and Pax6 in human osteosarcoma cells was first detected and luciferase reporter gene system was used to verify that Pax6 was the direct target of miR-19. Then, the effect of miR-19 on the biological characteristics of osteosarcoma cells was detected by 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT), flow cytometry, and cell invasion assay through inhibiting the expression of miR-19 in osteosarcoma cells. The rescue experiment confirmed that miR-19 played a biological role through Pax6. This study might provide experimental basis for the application of miR-19 and Pax6 in the treatment of osteosarcoma.
Materials and methods
Cell culture
Normal human osteoblasts hFOB 1.19 were cultured in mixed culture medium (Dulbecco’s modified Eagle’s medium (DMEM) and F-12 medium mixed by 1:1) containing 0.5 mM sodium pyruvate, 2.5 mM
Cell transfection
Normal cultured MG-63 cells were chosen and inoculated into six-well plate. The MG-63 cells were transfected with miR-19 inhibitor, miRNA inhibitor negative control (NC), Pax6 small interfering RNA (siRNA), and NC siRNA according to transfection specification of Lipofectamine 2000 (Invitrogen). Pax6 siRNA, miR-19 inhibitor, and their respective NC were provided by GenePharma (Shanghai, China). At the same time, normal control group (Normal) was set up. MiR-19 inhibitor, Pax6 siRNA, and their corresponding NC were diluted in MEM culture medium without serum. Then, Lipofectamine 2000 was diluted with MEM, mixed gently, and incubated at room temperature for 5 min. Then, we gently mixed diluted Lipofectamine 2000 with the above diluted mixture and incubated at room temperature for 20 min to form a complex. The complex was added to the culture plate containing MG-63 cells and placed into a culture box at 37°C, 5% CO2. After 48 h, the mixture was transferred to another MEM culture medium containing 10% FBS.
Quantitative real-time polymerase chain reaction
The quantitative real-time polymerase chain reaction (qRT-PCR) method was used to detect the expression of Pax6 messenger RNA (mRNA) and miR-19 in normal human osteoblasts (hFOB 1.19) and osteosarcoma cell lines (U2OS, MG-63, and Saos-2). The hFOB 1.19 and osteosarcoma cells cultured in vitro were collected. RNA was extracted using miRNA isolation kit. MiRNA Assay and TaqMan Universal PCR Master Mix were used to detect the expression of mature miR-19, and U6 was chosen as reference gene. The above reagents were purchased from Applied Biosystems (Foster city, CA, USA). Total RNA was extracted from hFOB 1.19 and osteosarcoma cells cultured in vitro by TRIzol reagents. Reverse transcription Kit (Qiagen, Hilden, Germany) was used to perform reverse transcription reaction. After the first strand was synthesized, we detected the expression of Pax6 mRNA, setting GAPDH as the reference gene. All responses were established with three complex holes. The experimental results were analyzed using the relative quantitative method of qRT-PCR.
Detection of luciferase activity
Through online software analysis (http://www.targetscan.org), Pax6 was predicted as one of the potential targets of miR-19. Then, luciferase reporter gene system was used to detect whether Pax6 was the target gene of miR-19. The sequence of Pax6 3′-UTR (untranslated region) which might be combined with miR-19 was cloned into pMIR-REPORT miRNA expression luciferase carrier (Applied Biosystems), so as to construct the luciferase reporter gene plasmid of the wild type (WT) of Pax6. We constructed a mutant (MUT) luciferase reporter gene plasmid by exerting point mutation in a nucleotide sequence of the Pax6 3′-UTR region that may be combined with miR-19. The Pax6 3′-UTR WT or Pax6 3′-UTR MUT type and miR-19 mimic or NC (miR-19 NC) were transfected simultaneously into HEK-293 cells by liposome transfection method. Cells were collected 48 h later. Dual-luciferase reporter assay system was used and Renilla luciferase (Renilla) was settled as reference to detect the activity of firefly luciferase (Firefly). The ratio of these two was regarded as the relative expression value (Firefly/Renilla).
MTT assay
MG-63 cells were transfected with miR-19 inhibitor and inhibitor NC. MTT analysis method was adopted to detect the effect of miR-19 on cell viability. MG-63 cells transfected with miR-19 inhibitor or inhibitor NC were seeded on 96-well plates. Then, 100 μL MTT solution (0.5 mg/mL) was added into each hole and incubated at 37°C, 5% CO2 for 4 h. Following, each hole was added with 100 μL 20% sodium dodecyl sulfate (SDS) for 24 h at 37°C. Optical density (OD) was measured at 570 nm by microplate reader (American BioTek Instruments Inc., Winooski, VT, USA). At the same time, normal control group (treated with nothing; Normal) was set; 10 replicates were examined in each experimental group, and each experiment was repeated 3 times.
Flow cytometry assay
MG-63 cells were transfected with miR-19 inhibitor and inhibitor NC. We washed the cells with phosphate-buffered saline (PBS) for 2 times, digested the cells with trypsin (Invitrogen), and then again washed the cells with PBS for 2 times. The cells were collected by centrifugation and propidium iodide (PI) dye was added for 30 min at 4°C kept in dark place. After sieving and filtration, the cells were detected and analyzed by flow cytometry (BD Biosciences, San Jose, CA, USA). FCM CellQuest software was used to count the cells in order to detect cell proliferation. Macquit data analysis software (BD Biosciences) was used to analyze the data. Next, the effect of miR-19 on apoptosis was detected. Cells were washed with PBS for 2 times after 48 h of the transfection. Annexin V-FITC (Roche, New York, NY, USA) and PI dye liquid (Roche) were added and incubated for 15 min at room temperature kept in dark place. After sieving and filtration, the cells were detected and analyzed by flow cytometry. FCM CellQuest software was used to count the cells; Macquit data analysis software was used to analyze the data.
Transwell invasion assay
MG-63 cells were transfected with miR-19 inhibitor and inhibitor NC and then were inoculated onto the upper chamber of the Transwell invasion chamber (Corning Inc., Corning, NY, USA). The upper chamber was pre-layered with a layer of Matrigel (BD Biosciences). The cells were cultured for 24 h at normal condition, then washed with PBS for 1–2 times and stained by crystal violet (Beyotime Institute of Biotechnology, Shanghai, China). Eight fields of view were randomly chosen, and the number of cells crossing polycarbonate membrane was counted under the Leica microscope. Cells that crossed the Transwell polycarbonate membrane were invasive cells.
Western blot
Total protein from transfected MG-63 cells was isolated. After protein quantity and electrophoretic transfer, the rabbit anti-human E-cadherin and Pax6 polyclonal antibodies (1:1000 dilution; Abcam, Cambridge, MA, USA), rabbit anti-human β-catenin monoclonal antibody (1:1000 dilution; Abcam), mouse anti-human Vimentin, extracellular signal–regulated kinase (ERK), phosphorylated ERK (p-ERK) polyclonal antibodies (1:1000 dilution; Abcam), and mouse anti-human β-catenin and β-actin monoclonal antibodies (1:2000 dilution; Sigma-Aldrich, St. Louis, MO, USA) were added and incubated with polyvinylidene fluoride (PVDF) membrane overnight at 4°C. The PVDF membrane was washed for 5 min with Tris-buffered saline with Tween 20 (TBST) for 3 times. Then, horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG or HRP-conjugated goat anti-mouse IgG antibodies (Sigma-Aldrich) were added and incubated at 37°C for 1 h. The PVDF membrane was washed with TBST for 3 times × 5 min. Enhanced chemiluminescence (ECL) autograph was conducted with chemiluminescence reagent. The relative content of each protein was expressed by target protein/β-actin gray scale ratio, and the relative expression changes were analyzed by PDQuest software (Bio-Rad Laboratories, Inc., Hercules, CA, USA).
Statistical analysis
SPSS17.0 statistical analysis software was adopted for statistical treatment of the experimental data. Data of more than two groups were compared using variance analysis at the 0.05 level.
Results
The expression of miR-19 and Pax6 mRNA in human osteosarcoma cells
The qRT-PCR results showed that the expression of miR-19 in osteosarcoma cell lines (U2OS, MG-63, and Saos-2) was significantly higher than that in normal human osteoblasts (hFOB 1.19; p < 0.01; Figure 1(a)), which indicated that miR-19 was upregulated in human osteosarcoma cell line. While the expression trend of Pax6 mRNA in human osteosarcoma cell lines was just the opposite of the expression of miR-19. The expression of Pax6 mRNA in osteosarcoma cells was significantly lower than that in hFOB 1.19 (p < 0.01; Figure 1(b)), which indicated that Pax6 mRNA was downregulated in human osteosarcoma cell line. These data demonstrated that the expression of miR-19 and Pax6 mRNA displayed a negative correlation in human osteosarcoma cells.
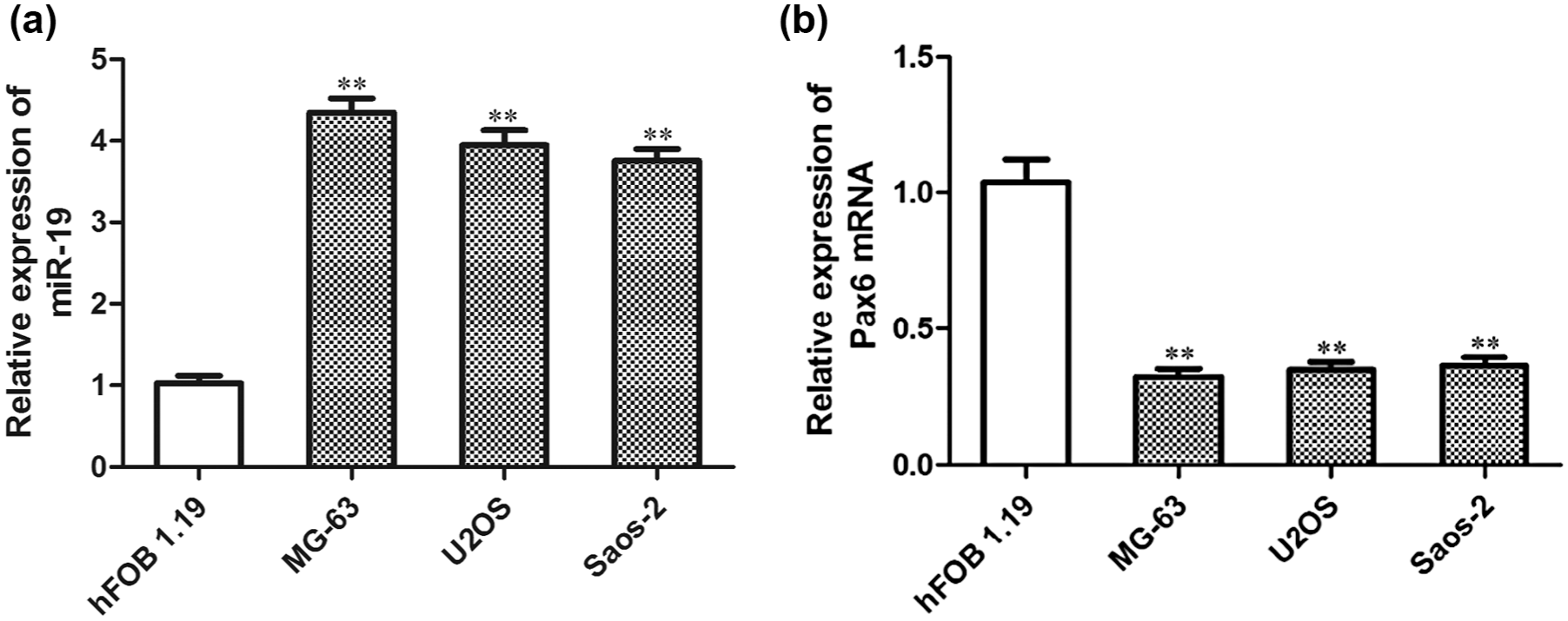
Expression of miR-19 and Pax6 mRNA in normal human osteoblasts (hFOB 1.19) and osteosarcoma cell lines (U2OS, Saos-2, and MG-63). (a) Relative expression of miR-19 in normal human osteoblasts (hFOB 1.19) and osteosarcoma cell lines (U2OS, Saos-2, and MG-63). (b) Relative expression of Pax6 mRNA in normal human osteoblasts and osteosarcoma cell lines. All data in bar graphs are presented as mean ± standard error (SE) from at least three independent experiments (**p < 0.01 compared to hFOB 1.19).
Pax6 was the target gene of miR-19
Based on the results of qRT-PCR, we found that the expression trend of Pax6 mRNA in human osteosarcoma cell lines was just the contrast of miR-19. We analyzed and predicted that Pax6 might be the target of miR-19 through online software (http://www.targetscan.org). Therefore, the luciferase reporter gene system was used to further verify that Pax6 was the target of miR-19. Luciferase assay showed that the luciferase activity was significantly suppressed in the miR-19 mimic transfection group compared with the mimic NC (p < 0.01). Results are shown in Figure 2(a), which indicated that miR-19 could directly interact with the target site of Pax6 3′-UTR.
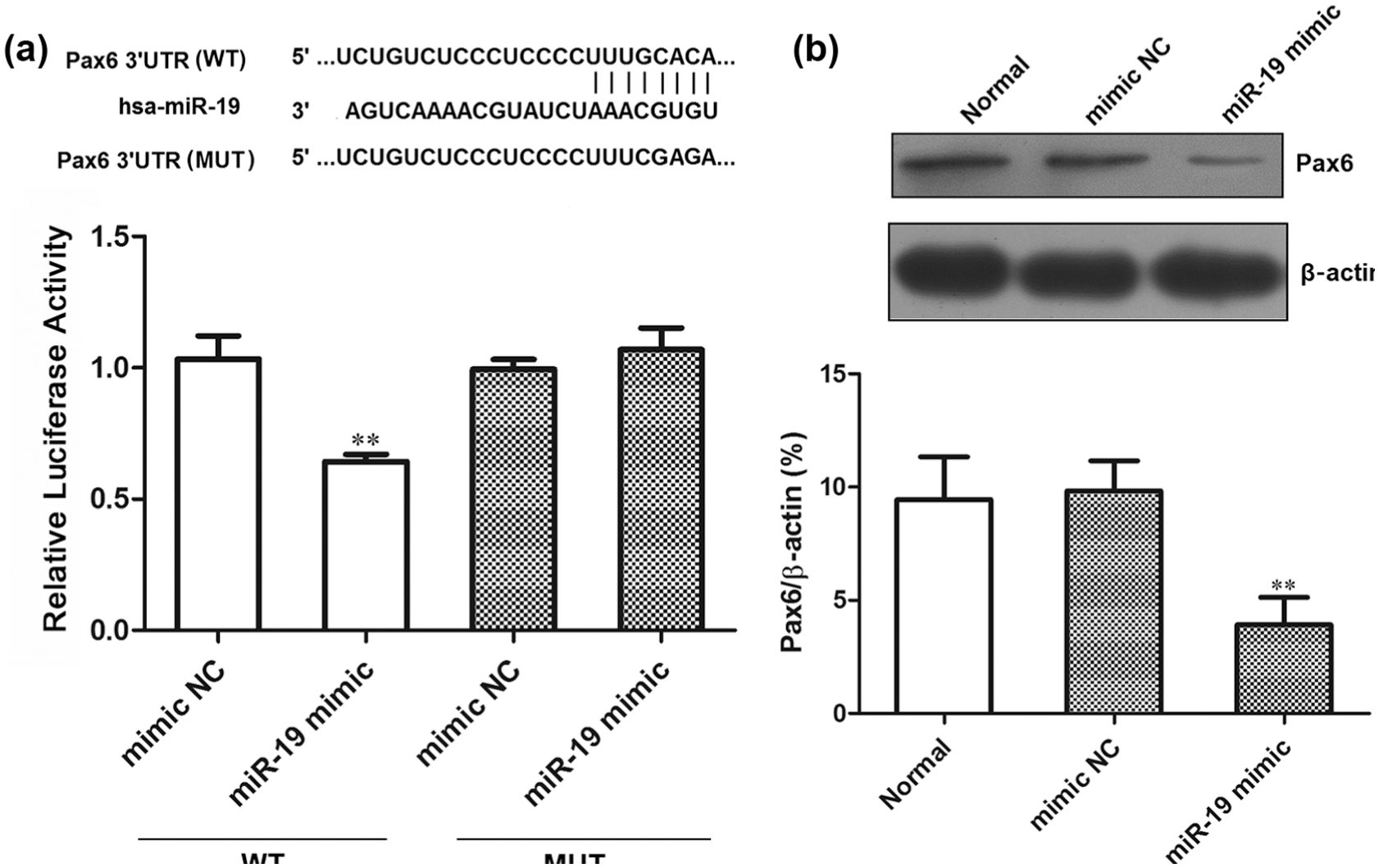
Pax6 was the target gene of miR-19. (a) Schematic representation of Pax6 3′-UTR showing putative miR-19 target site, and relative luciferase activity of HEK-293 cells that were co-transfected with miR-19 mimic or negative control (mimic NC). (b) The effect of miR-19 on the expression of Pax6 protein. All data were obtained from at least three independent experiments (**p < 0.01 compared to mimic NC group and Normal group).
The effect of miR-19 on Pax6 expression was detected by western blot assay. The results showed that the Pax6 protein expression in miR-19 mimic transfection group was significantly lower than that in the normal control group (Normal; p < 0.01) and the NC group (p < 0.01; Figure 2(b)), which indicated that overexpression of miR-19 inhibited the expression of Pax6 protein. These data demonstrated that Pax6 was the target gene of miR-19.
MiR-19 inhibition decreased MG-63 cell viability
The expression of miR-19 increased in osteosarcoma cell line MG-63, so we hypothesized whether the inhibition of miR-19 expression could inhibit its tumorigenicity. First, we transfected miR-19 inhibitor into MG-63 cells and detected the expression change of miR-19 by qRT-PCR. The results showed that the expression of miR-19 in miR-19 inhibitor transfection group was significantly lower than that in the normal control group (p < 0.01) and NC group (p < 0.01; Figure 3(a)). MTT analysis showed that after the transfection of miR-19 inhibitor, the cell viability was significantly lower than that of the normal control group and the NC group (p < 0.01; Figure 3(b)), which indicated that the inhibition of miR-19 expression could decrease the viability of MG-63 cells.

The effect of miR-19 on the cell viability of MG-63 cells: (a) The relative expression of miR-19 in MG-63 cells was determined by qRT-PCR (**p < 0.01 versus Normal group or inhibitor NC group). (b) The effect of miR-19 on the cell viability of MG-63 cells was determined by MTT assay. All data were obtained from at least three independent experiments (**p < 0.01 compared to Normal group or inhibitor NC group).
MiR-19 inhibition suppressed MG-63 cell proliferation and induced MG-63 cell apoptosis
MG-63 cells were transfected with miR-19 inhibitor and corresponding NC. After 40 h, the cell cycle was detected by flow cytometry and the proportion of S-phase cells was counted. The results showed that the proportion of S cells in the miR-19 inhibitor transfection group was significantly lower than that in normal control group and the NC group (p < 0.01). Results are shown in Figure 4. The effect of miR-19 on cell apoptosis was also analyzed by flow cytometry. The results showed that apoptotic cells in the miR-19 inhibitor transfection group were significantly higher than that in the normal control group and the NC group (p < 0.01; Figure 5). These data indicated that the inhibition of miR-19 expression could inhibit the proliferation and promote the apoptosis of MG-63 cells.

The effect of miR-19 on the proliferation of MG-63 cells. All data were obtained from at least three independent experiments (*p < 0.05 compared to Normal group or inhibitor NC group).

The effect of miR-19 on the apoptosis of MG-63 cells. All data were obtained from at least three independent experiments (**p < 0.01 compared to Normal group or inhibitor NC group).
MiR-19 inhibition decreased the invasion ability of MG-63 cells
The effect of miR-19 on MG-63 cell invasion ability was analyzed by Transwell invasion assay. Crystal violet staining results showed that the downregulation of miR-19 significantly reduced the number of cells passing through the Transwell invasion chamber (p < 0.01). According to Figure 6, the invasion of MG-63 cells was inhibited due to the downregulation of miR-19.
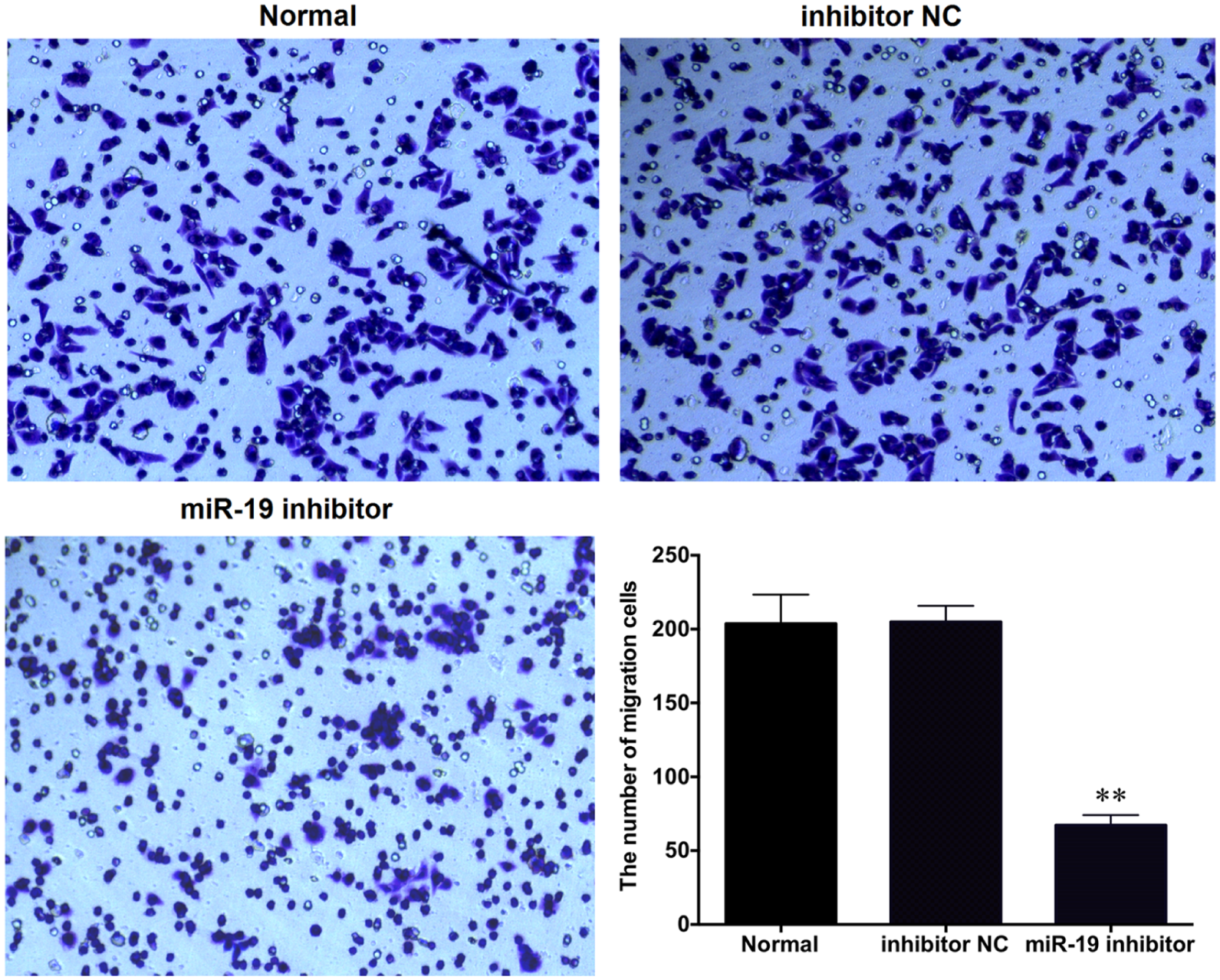
The effect of miR-19 on MG-63 cell invasion. All data were obtained from at least three independent experiments (**p < 0.01 compared to Normal group or inhibitor NC group).
MiR-19 inhibition decreased the expression of epithelial–mesenchymal transition–related proteins and suppressed ERK pathway
Western blot analysis showed that the expression of epithelial proteins (E-cadherin and β-catenin) in miR-19 inhibitor transfection group was significantly higher than that in the normal control group and the NC group (p < 0.01), while the expression of mesenchymal protein (Vimentin), ERK, and p-ERK in miR-19 inhibitor transfection group was significantly lower than that in the normal control group and the NC group (p < 0.01). According to Figure 7, the inhibition of miR-19 significantly increased the expression of epithelial proteins (E-cadherin and β-catenin) and decreased the expression of mesenchymal protein (Vimentin). MiR-19 inhibitor suppressed the expression of ERK while it has little effect in ERK activation.

MiR-19 inhibition significantly increased the expression of epithelial proteins (E-cadherin and β-catenin) and decreased the expression of mesenchymal protein (Vimentin) and extracellular signal–regulated kinase (ERK) pathway. (a) The effect of miR-19 on the expression of epithelial–mesenchymal transition–related proteins (E-cadherin, β-catenin, and Vimentin). (b) The effect of miR-19 on the expression of ERK pathway–related proteins (ERK and p-ERK). All data were obtained from at least three independent experiments (**p < 0.01 compared to Normal group or inhibitor NC group).
Validation of the biological role of miR-19 through regulating Pax6
Based on the above results, we might make a conclusion that miR-19 upregulation played a role of oncogene in MG-63 cells through inhibiting the expression of Pax6. Therefore, we assumed whether inhibiting the expression of Pax6 could reverse the biological effects induced by miR-19 downregulation. The miR-19 inhibitor and Pax6 siRNA were simultaneously transfected into MG-63 cells. MTT, flow cytometry, and Transwell analysis method were used to further verify that miR-19 played a cell biological role through regulating the expression of Pax6.
First, the expression changes of Pax6 mRNA were detected in each group by qRT-PCR. The results showed that the transfection of Pax6 siRNA into MG-63 cells could significantly inhibit the expression of Pax6 mRNA (p < 0.01), while the expression of Pax6 mRNA in the transfection group (miR-19 inhibitor + Pax6 siRNA) was significantly higher than that in the Pax6 siRNA transfection group (p < 0.01) and had no significant differences compared with the normal control group (Normal) and siRNA NC group (siRNA scramble; Figure 8(a)).

Validation of the biological role of miR-19 through regulating Pax6 in osteosarcoma cells. (a) The expression of Pax6 in MG-63 cells. (b) The viability of MG-63 cells. (c) The apoptosis of MG-63 cells. (d) The invasion of MG-63 cells. All data were obtained from at least three independent experiments (**p < 0.01 compared to Normal group and siRNA scramble group; ##p < 0.01 indicated Pax6 siRNA compared to miR-19 inhibitor + Pax6 siRNA).
The results from MTT and Transwell analysis showed that cell viability and the number of cells migrating through Transwell chamber in miR-19 inhibitor and Pax6 siRNA transfection group was significantly lower than that in the Pax6 siRNA transfection group (p < 0.01) and had no significant differences compared with the normal control group (Normal) and the siRNA NC group (siRNA scramble). The results are shown in Figure 8(b) and (d). The results from flow cytometry analysis (Figure 8(c)) suggested that the number of apoptosis cells in miR-19 inhibitor and Pax6 siRNA transfection group was significantly higher than that in the Pax6 siRNA transfection group (p < 0.01) and had no significant differences compared with the normal control group (Normal) and the siRNA NC group (siRNA scramble). These data demonstrated that Pax6 inhibition could reverse the biological effect induced by miR-19 inhibition in MG-63 cells.
Discussion
This study first detected the expression of miR-19 in normal human osteoblasts (hFOB 1.19) and osteosarcoma cell lines (U2OS, MG-63, and Saos-2) by qRT-PCR. The results showed an upregulation of the expression of miR-19 in osteosarcoma cell lines, which indicated that miR-19 might play an important role in the tumorigenesis of osteosarcoma. Pax6 protein serves as a transcription factor. There have been reported that Pax6 protein mainly expresses in the nucleus and plays the role of transcription factors. When cell proliferation is inhibited, Pax6 will shift from nucleus to cytoplasm, thus losing the regulation function on transcription.32,33 The abnormal expression of Pax6 gene is often existent in many tumor cells. There are reports on the downregulation of Pax6 by promoter methylation, which was associated with poor prognosis in non-small-cell lung cancer, 34 and suppression of Pax6 promoted cell proliferation and inhibited apoptosis in human retinoblastoma cells. 35 While a study of Zhao et al. 36 showed that downregulation of Pax6 by short hairpin RNA (shRNA) inhibited proliferation and cell-cycle progression of human non-small-cell lung cancer cell lines, and Pax6 was expressed in pancreatic cancer and actively participated in cancer progression through activation of the MET tyrosine kinase receptor gene. 37 In addition, a report demonstrated that Pax6 could downregulate miR-124 expression to promote cell migration. 38 Therefore, we are eager to explore the expression of Pax6 in osteosarcoma. The qRT-PCR assay showed that Pax6 mRNA expression decreased in osteosarcoma cell lines (U2OS, MG-63, and Saos-2), and the expression trend was the opposite of miR-19.
Through online software TargetScan, we predicted that Pax6 might be a target gene of miR-19. In order to further verify that Pax6 is a target gene of miR-19 regulation, luciferase reporter gene system was used to conduct the experiment. The results showed that miR-19 had an interaction with Pax6 3′-UTR. And western blot was used to detect the regulation effect of miR-19 on the expression of Pax6. The results showed that the overexpression of miR-19 inhibited the expression of Pax6 protein.
There are reports on the expression of miR-19, which is upregulated in a variety of tumor cells.39–41 In this study, miR-19 expression lined up in osteosarcoma cells, which may play a role of proto-oncogene in osteosarcoma cells. In order to investigate the role of miR-19 in human osteosarcoma cells, miR-19 expression was inhibited in osteosarcoma MG-63 cells. Then, MTT, flow cytometry, and Transwell were used to explore the effects of miR-19 on the biological characteristics of human osteosarcoma cell MG-63. The results showed that miR-19 inhibition could the decrease the cell viability of MG-63 cells and the proliferation and invasion activity of MG-63 cells and promote cell apoptosis. These results also confirmed the role of miR-19 as the proto-oncogene miRNAs.
The main cause for the distant metastasis of osteosarcoma is the abnormal expression of tumor metastasis–related genes and metastasis suppressor genes.9,10 E-cadherin becomes a hotpot of research because it plays an important role in the invasion and metastasis of tumor. E-cadherin, a transmembrane glycoprotein with a molecular weight of 120 kDa, which belongs to the cadherin superfamily, is an important molecular cell and matrix. The expression level and activity trend of E-cadherin will affect the detachment and reattachment of tumor cells, so it plays a key role in the evolution of tumor invasion. 42 β-catenin is expressed in many tissues, and it has an independent effect on cell adhesion mediated by calcium-binding protein complex. 43
The deficiency or downregulation of β-catenin expression will lead to the functional loss of complex E-cadherin/β-catenin, the result of which is a loss of contact inhibition of tumor cells and unlimited proliferation but without differentiation; connection loss between cells enables tumor cells to easily fall off from the local adhesion sites. 44 Vimentin belongs to cell intermediate filament protein family and is an important component of the cytoskeleton, which plays an important role in maintaining cell and organelle morphology, participating in mitosis and cell differentiation, wound healing, signal transduction, and so on. It has been found that Vimentin displayed upregulation in a variety of epithelial tumor cells and had a close affiliation with the migration, invasion, and metastasis ability of tumors. 45
In this study, the effect of miR-19 on the expression of epithelial–mesenchymal transition (EMT)-related proteins (E-cadherin, β-catenin, and Vimentin) in MG-63 cells was detected by western blot. The results suggested that inhibiting the expression of miR-19 would promote the expression of E-cadherin and β-catenin, but the expression of Vimentin could be inhibited. However, it indicated that miR-19 could promote the expression of Vimentin and inhibit the expression of E-cadherin and β-catenin, so as to promote the invasion of osteosarcoma cells. This study further confirmed that miR-19 may play a biological role through the ERK pathway.
In order to further verify whether miR-19 plays the cell biological role in MG-63 cells through the regulation of Pax6, Pax6 siRNA and miR-19 inhibitor were simultaneously transfected into MG-63 cells. MTT, flow cytometry, and Transwell analysis results suggested that the inhibition of Pax6 could reverse the biological effect induced by miR-19 inhibition in MG-63.
In summary, the expression of miR-19 in human osteosarcoma cell lines is upregulated and the expression of Pax6 is downregulated. MiR-19 can negatively regulate the expression of Pax6 protein, which can promote the malignant phenotypes of osteosarcoma cells via activation of the ERK signaling pathways. Therefore, miR-19/Pax6 may offer potential for use as a target for the treatment of osteosarcoma.
Footnotes
Acknowledgements
Qingbing Meng and Ming Dai have contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by National Natural Science Foundation of China (Grant No. 81402226).
