Abstract
Lung cancer is one of the most frequently occurring cancers throughout the world as well as in Bangladesh. This study aimed to correlate the prognostic and/or predictive value of functional polymorphisms in SULT1A1 (rs9282861) and XRCC1 (rs25487) genes and lung cancer risk in Bangladeshi population. A case-control study was conducted which comprises 202 lung cancer patients and 242 healthy volunteers taking into account the age, sex, and smoking status. After isolation of genomic DNA, genotyping was done by polymerase chain reaction–restriction fragment length polymorphism method and the lung cancer risk was evaluated as odds ratio that was adjusted for age, sex, and smoking status. A significant association was found between SULT1A1 rs9282861 and XRCC1 rs25487 polymorphisms and lung cancer risk. In case of rs9282861 polymorphism, Arg/His (adjusted odds ratio = 5.06, 95% confidence interval = 3.05–8.41, p < 0.05) and His/His (adjusted odds ratio = 3.88, 95% confidence interval = 2.20–6.82, p < 0.05) genotypes were strongly associated with increased risk of lung cancer in comparison to the Arg/Arg genotype. In case of rs25487 polymorphism, Arg/Gln heterozygote (adjusted odds ratio = 4.57, 95% confidence interval = 2.79–7.46, p < 0.05) and Gln/Gln mutant homozygote (adjusted odds ratio = 4.99, 95% confidence interval = 2.66–9.36, p < 0.05) were also found to be significantly associated with increased risk of lung cancer. This study demonstrates that the presence of His allele and Gln allele in case of SULT1A1 rs9282861 and XRCC1 rs25487, respectively, involve in lung cancer prognosis in Bangladeshi population.
Keywords
Introduction
Lung cancer has been the most common cancer in the world for several decades and is the most common cause of cancer-related death worldwide. 1 It is the most commonly diagnosed cancer and leading cause of death in Bangladeshi males. Patients with advanced and metastatic disease have 5-year survival rate less than 5%. 2 According to published report (2008–2010) from January 2008 to December 2010 time period, 46,110 new cases attended the Outpatient Department of National Institute of Cancer Research and Hospital (NICRH) in Bangladesh. Among them, 27,281 were diagnosed as cancer cases, and out of them, 28.39% male cancer patients were fighting against lung cancer. 2 International Agency for Research on Cancer (IARC) published that 7.5% death had been projected to be caused by cancer in 2005 in Bangladesh and it will rise up to 13% in 2030. A recent World Health Organization (WHO) study anticipated that there are 196,000 lung cancer cases in Bangladesh and 85% of those are aged 30 years or above. 3
Lung cancer is the result of complex pathway, multilateral steps and multifactorial process. The actual etiology of lung cancer is yet to understand. Smoking cigarette is a prominent risk factor for lung cancer. 4 Despite the exposure to cigarette smoking, many other factors such as genetic factors also may play a significant role in the development of lung cancer. 4 Previous studies revealed that polymorphisms of DNA-repair genes and metabolic genes of carcinogens are a key factor for lung cancer susceptibility, as because DNA-repair activities play a critical role on gene function modification and protection of the genome, and genetic polymorphisms of metabolic enzymes of carcinogens are strongly predicted to modify an individual’s susceptibility to cancer.5–8
Sulfotransferases (SULTs) are multifunctional and important enzymes that regulate catalytic function of sulfonate conjugation, which is the main pathway in phase II metabolism of numerous endogenous and exogenous compounds. 9 It acts via the transfer of the sulfuryl moiety (–SO3) from the nucleotide donor, 3-phosphoadenosine 5′-phosphosulfate (PAPS), to the hydroxyls and primary amines of acceptors. Sulfotransferase 1A1 (SULT1A1) is a member of SULT superfamily, considered to be the predominant type of SULTs because of its extensive tissue distribution and abundance. 10 SULT1A1 gene is mapped to chromosome 16p11.2-12.1. A single nucleotide transition (G > A) at codon 213 in exon 7 of SULT1A1 gene causes an Arg to His amino acid substitution (rs9282861). Previous studies showed that SULT1A1 rs9282861 genetic polymorphism leads to a decrease in enzymatic activity of SULT1A1 and the sulfonation efficiency thus associating with susceptibility to several cancers including lung cancer.11,12
X-Ray Repair Cross Complementing 1 (XRCC1) gene is one of the >20 genes that is involved in both the base excision repair (BER) and single-stranded break repair (SSBR) processes 13 and removes oxidative DNA damage caused by exposures to ionizing radiation or alkylating agents. 14 Being located on chromosome 19q13.2-13.3, XRCC1 gene encodes a scaffolding protein that forms a complex with polymerase beta (pol β), DNA ligase III (lig III), and poly(ADP-ribose) polymerase (PARP).15,16 Within these repair processes, XRCC1 functions in the ligation step, linking healthy DNA strands back together after the removal of damaged bases has occurred. 14 XRCC1 also binds polynucleotide kinase (PNK) through the PNK forkhead–associated domain and stimulates repair process at damaged DNA termini. 17 This gene is, hence, deemed to be a vital candidate gene influencing the susceptibility to lung cancer.18–21 One of the most extensively studied single-nucleotide polymorphisms (SNPs) of XRCC1 gene is XRCC1 rs25487 (Arg399Gln) where a transition of G to A nucleotide at codon 399 in exon 10 happens. Many studies have shown that Arg399Gln is associated with increased risk of lung cancer,22,23 while some non-significant or negative association is also reported by other studies.24,25 Furthermore, data have suggested that the effect of the XRCC1 SNPs on lung cancer risk may be dependent on ethnicity. 26
Cigarette smoking is thought to be responsible for 90% of lung cancer cases in men and 78% in women. 27 Processed tobacco contains more than 3000 compounds, including 30 carcinogens. 28 SULT1A1 is known to be involved in the metabolism of procarcinogens such as heterocyclic amines and polycyclic aromatic hydrocarbons, both of which are present in tobacco smoke. In some studies, smokers with SULT1A1 His213 allele have been shown to possess higher risk of developing lung cancer.29,30 In addition, carcinogens present in tobacco products generate free radicals. These free radicals increase the oxidative burden and can cause potentially dangerous DNA lesions. If those oxidative DNA lesions are not removed quickly enough, they can become self-perpetuating mutations that contribute to the development of disease states, particularly cancer. 31 Polymorphisms in genes coding for DNA-repair proteins may play a role in susceptibility to environmental carcinogens. As XRCC1 plays an important role in DNA repair, studies relating smoking status and XRCC1 rs25487 polymorphism have been conducted indicating the involvement of heterozygous Arg/Gln and homozygous Gln/Gln variants with lung cancer risk in smokers. 19
Since SULT1A1 rs9282861 and XRCC1 rs25487 polymorphisms have been proven to be a risk factor of lung cancer in different ethnicities, for example, Caucasian, Indian, Chinese, Korean, and Turkish,32–38 we had therefore undertaken a study to find out a possible correlation of these SNPs with lung cancer patients in Bangladeshi population for the very first time.
Methods
Subject selection
A case-control study was conducted on 202 lung cancer patients and 242 healthy volunteers matched by age, sex, lifestyle, and smoking status. Lung cancer patients were recruited from NICRH, Mohakhali, Dhaka Medical College Hospital, Ahsania Mission Cancer Hospital and Bangabandhu Sheikh Mujib Medical University (PG Hospital), Dhaka, Bangladesh. These patients were diagnosed histologically with lung cancer as per the International Association for the Study of Lung Cancer/American Thoracic Society/European Respiratory Society International Multidisciplinary Classification of Lung Adenocarcinoma 39 between May 2014 and September 2015. After physical examination, controls were selected by matching age, sex, and smoking status with the lung cancer patients. Detailed history on smoking status, demographic characteristics, occupational exposure, and lifestyle factors of all cases and controls were collected by trained nurses in the presence of expert physicians. The study was conducted in accordance with the Helsinki Declaration and its further amendments.40,41 Written ethical permission from the ethical review committee of the respective hospitals was taken to approve the protocol. Prior to study, all the cases and controls had to give written consent in a prescribed form. The clinicopathological characteristics were recorded in a prescribed questionnaire. All the patients and controls were included and excluded following the criteria of our previous article. 42 The genetic study was conducted in the laboratory of pharmacogenetics and pharmacokinetics at the Department of Clinical Pharmacy and Pharmacology, Faculty of Pharmacy, University of Dhaka, Bangladesh.
Genotyping
We collected 3 mL blood samples from all the patients and controls in sterile tubes containing ethylenediaminetetraacetic acid (EDTA)-Na2 and stored at −80°C until DNA extraction. DNA extraction and amplification of genomic DNA using the predesigned forward and reverse primers were performed as described previously.43,44 Polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) method was used to genotype the SULT1A1 rs9282861 and XRCC1 rs25487 polymorphisms. PCR products were confirmed by running on agarose gel (1%) and the confirmed PCR products were digested overnight with the respective restriction endonucleases and were visualized on 2% agarose gel after ethidium bromide staining. The results were validated by performing sequencing of 2% the heterozygous and homozygous samples.
Statistical analysis
Chi-square (χ2) test was applied to compare the sociodemographic and clinicopathological data between the cases and controls. We evaluated the risk as odds ratios (ORs) with 95% confidence intervals (95% CI) by multivariate logistic regression, and ORs were adjusted for different variables like age, sex, and smoking status. All statistical analyses were done applying the SPSS software, version 22.0, and the adjusted OR has been expressed as AOR. In all of the analyses, p < 0.05 was considered as statistically significant.
Results
Characteristic features of study population
A total of 202 lung cancer patients were included in this case-control study. The study included 178 males (88.12%) and 24 females (11.88%) diagnosed with lung cancers, with a mean age of 54.87 years. The control group comprised 173 males (71.49%) and 69 females (28.51%) with a mean age of 43.15 years. The percentage of the patients below 60 years of age was 28.22% and that of controls was 26.45%. Patients smoking profile seemed to have positive contribution in causing lung cancer as current, former and ever smokers showed statistically significant risk of lung cancer development when compared to never smokers. No significant deviation (p > 0.05) was found between patients and controls considering different demographic parameters like education, age, family history of lung cancer in first- and second-degree relatives (Table 1).
Demographic characteristics of the patients and controls.
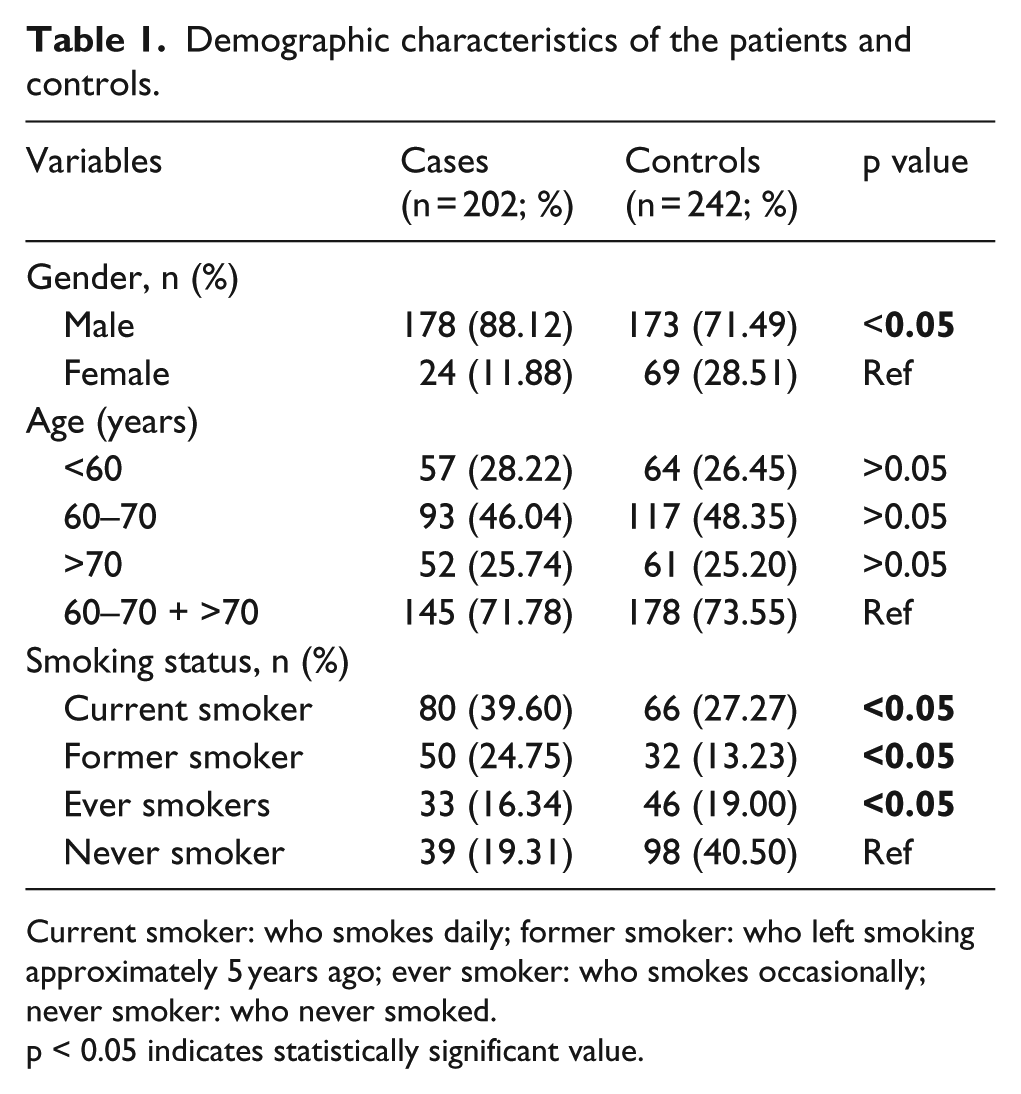
Current smoker: who smokes daily; former smoker: who left smoking approximately 5 years ago; ever smoker: who smokes occasionally; never smoker: who never smoked.
p < 0.05 indicates statistically significant value.
Impact of SULT1A1 rs9282861 polymorphism
The rs9282861 polymorphism of SULT1A1 is located outside exon 7 in a sequence of guanine (G) mutated to adenine (A), forming a HaeII (NEB, USA) restriction site that yielded a 281-bp fragment length PCR product. The electrophoresis result for the product after HaeII digestion is shown in Figure 1. The mutant homozygous produced two fragments (180 and 101 bp), and the heterozygous produced three fragments (281, 180, and 101 bp), whereas wild-type could not be digested.
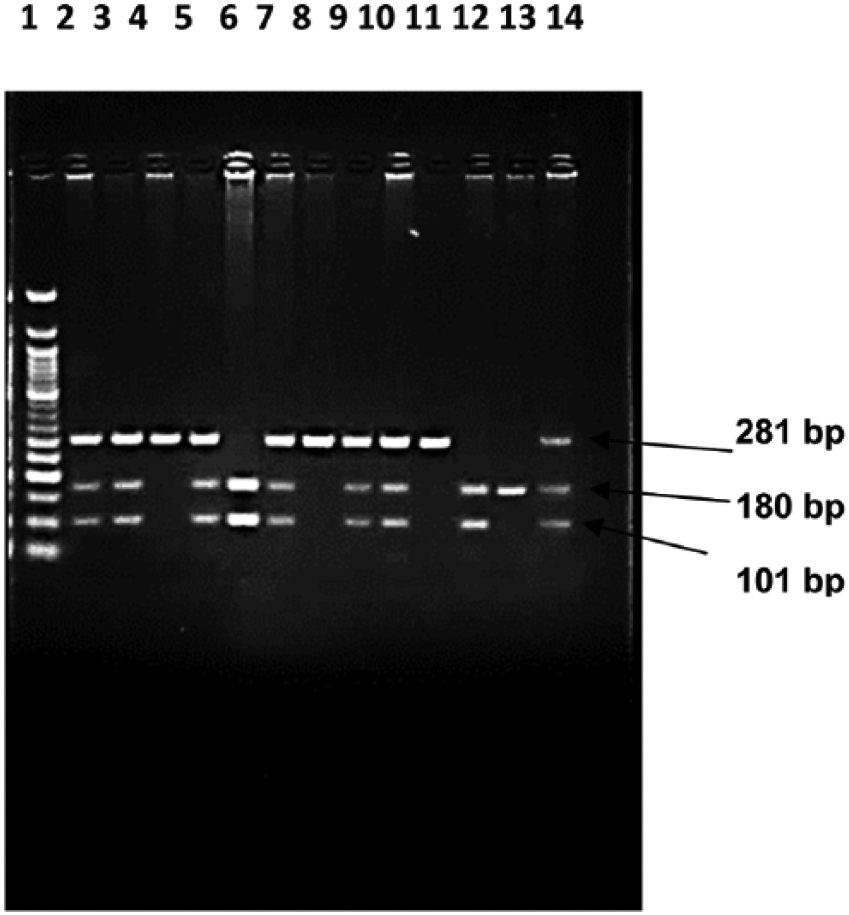
Restriction endonuclease (
The distribution of the SULT1A1 rs9282861 genotype frequencies in patients and control is presented in Table 2. The A allele was prevalent in our study population. Out of 202 patients, 49% carried the SULT1A1 rs9282861 variant A allele, whereas in controls these carriers were 35.95%. The patients with Arg/His heterozygous genotype were found to have 5.06-fold increased chance (AOR = 5.06, 95% CI = 3.05–8.41, p < 0.05) of developing lung cancer, whereas the carriers of His/His mutant homozygous genotype were at 3.88-fold greater risk of developing lung cancer (AOR = 3.88, 95% CI = 2.2–6.82, p < 0.05) compared with Arg/Arg normal homozygous genotype.
Genotype frequencies of SULT1A1 rs9282861 and XRCC1 rs25487 gene polymorphisms in cases and controls.

AOR: adjusted odds ratio; CI: confidence interval.
Impact of XRCC1 rs25487 polymorphism
In case of XRCC1 rs25487 polymorphism, restriction endonuclease EcoNI (NEB, USA) cleaved the mutated Gln allele giving two fragments of 148 and 94 bp but did not cleaved the wild-type Arg allele leaving the 242 bp fragment length PCR product undigested. Digestion by EcoNI illustrated the mutation of XRCC1 rs25487 which is shown in Figure 2.

Restriction endonuclease (
Table 2 also contains the distribution of rs25487 genotypes in cases and controls. Percent distribution of Arg/Gln heterozygous genotype was found higher in patient group compared to that of in control group (45.05% vs 28.51%) and showed a high risk factor of 4.57 (AOR = 4.57, 95% CI = 2.79–7.46, p < 0.05) of developing lung cancer compared to Arg/Arg normal homozygous genotype. The Gln/Gln mutant homozygous genotype was statistically found to be responsible for developing lung cancer and comprises a 4.99-fold (AOR = 4.99, 95% CI = 2.66–9.39, p < 0.05) elevated risk of lung cancer susceptibility compared with Arg/Arg homozygous genotype. Similarly, A allele frequency was found as high as 48.76% out of 202 lung cancer patients and 29.13% in a group of 242 controls.
Impact of smoking status in SULT1A1 rs9282861 and XRCC1 rs25487 polymorphisms in lung cancer risk
In case of SULT1A1 rs9282861 polymorphism, we found that current smokers carrying GG (Arg/Arg) and GA (Arg/His) genotypes have an increased risk of developing lung cancer with ORs of 3.03 and 3.34, respectively, compared to never smokers with GG (Arg/Arg) normal homozygous, and both of these genotypes are statistically significant (p < 0.05). Former smokers and ever smokers with heterozygous GA (Arg/His) genotypes have a 4.90-fold (OR = 4.90, 95% CI = 1.66–14.48, p < 0.05) and 3.62-fold (OR = 3.62, 95% CI = 1.29–10.15, p < 0.05) increased risk of lung cancer susceptibility, respectively. No other smoker type showed statistically significant (p < 0.05) risk association for lung cancer development (Table 3).
Correlation of SULT1A1 rs9282861 polymorphisms with smoking status.
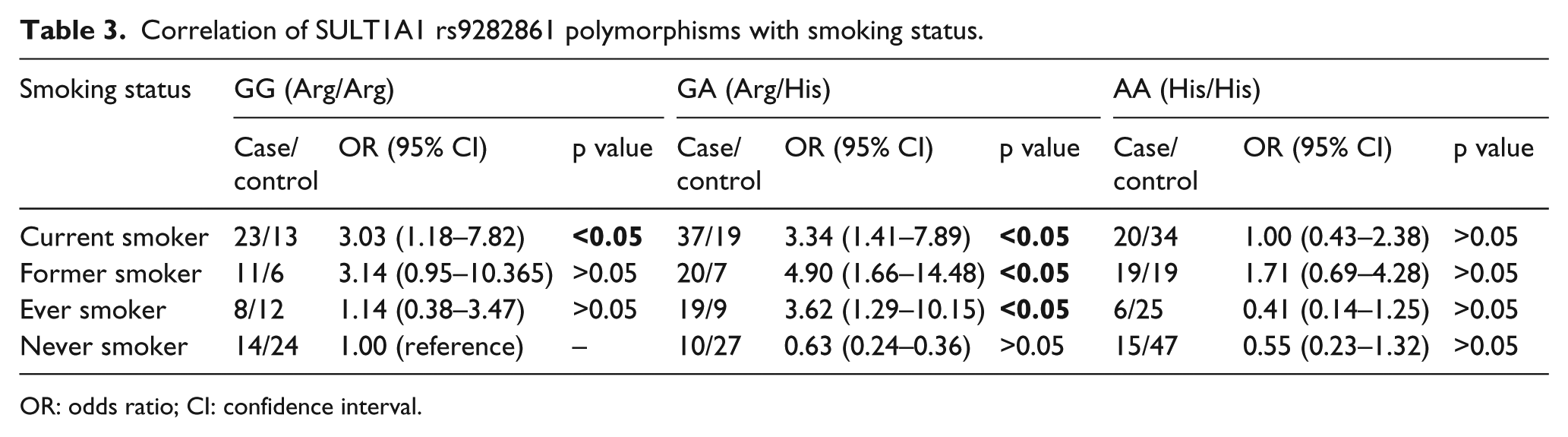
OR: odds ratio; CI: confidence interval.
Correlation between XRCC1 rs25487 polymorphism and smoking status for lung cancer susceptibility is presented in Table 4. In case of XRCC1 rs25487 polymorphism, carriers of GG (Arg/Arg) genotypes in current smokers showed higher risk of lung cancer development when compared with never smokers carrying wild-type GG (Arg/Arg) genotype. It showed 5.6-fold (OR = 5.6, 95% CI = 1.64–19.15, p < 0.05) more chance of developing lung cancer in current smokers. Other smoker groups demonstrated no significant risk of lung cancer susceptibility.
Correlation of XRCC1 rs25487 polymorphisms with smoking status.

OR: odds ratio; CI: confidence interval.
Discussion
Genetic alteration in the genomic DNA due to SNP has an association with tumor progression.45–48 Various incidences of polymorphisms in genomic DNA, their susceptibility to genetic alterations, and the risk of tumor progression substantially differ among different ethnic groups.45–48 As human SULT1A1 is an important enzyme in the metabolism of endogenous and exogenous carcinogens and the G638A polymorphism in SULT1A1 results in reduced enzyme activity, we investigated whether the SULT1A1 rs9282861 genotype has an effect on the individual susceptibility to lung cancer development. We analyzed 202 lung cancer patients and 242 controls in Bangladeshi population.
Our experiment on SULT1A1 rs9282861 polymorphism reveals that the percentage of heterozygous GA (Arg/His) genotype was 25.62% in controls, whereas 42.58% in cases, and the percentage of mutant AA (His/His) genotype was 23.14% in control, whereas 27.72% in cases (Table 2). Our study results go in line with other studies conducted on different populations. Arslan et al. 33 detected a high prevalence of Arg/His heterozygosity in the Turkish population with 36.5% of controls and 49.1% of patients. A somewhat similar situation has been observed in a Southeast Asian population, where the SULT1A1 histidine homozygous genotype has been found associated with lung cancer. 49 Similar findings have also been reported in studies on some other cancer types. Wu et al. 50 conducted a study consisting of 187 patients with esophageal cancer and 308 controls in a Chinese population and found a 3.53-fold increased risk (95% CI = 2.12–5.87) of the cancer for the SULT1A1 638GA genotype compared with the GG genotype. Bamber et al. 51 in a comprehensive study have also reported that a significantly reduced risk of cancer (OR = 0.47; 95% CI = 0.27–0.83, p < 0.05) was associated with the SULT1A1 638GG homozygotes in subjects under the age of 80 years in a group of Caucasian colorectal cancer patients. Finding rationale, several studies have demonstrated that the homozygosity of arginine allows certain SULT1A1 proteins to form stable complex with other metabolic compounds.52,53 It indicates that the SULT1A1 Arg allele is preferentially retained in Arg/His germline heterozygotes. It increases the possibility of having unknown mutation in the SULT1A1 gene (either in the exon or in the intronic region) and/or additional genetic changes in other gene(s) which may make Arg/His heterozygous patients to be at high risk for an invasive cancer early progression. However, inconsistent results also exist to show null association between SULT1A1 genotype and risk of lung cancer 6 and urothelial epithelial cancers in a Japanese population 54 and colorectal cancer 55 and prostate cancer 56 in Caucasian populations.
In this study, we assessed another most common SNP of XRCC1 in Bangladeshi population for the first time, since the role of rs25487 polymorphism in relation to lung cancer risk had not yet been reported. It was found that heterozygous GA (Arg/Gln) frequency was 28.51% in controls, whereas 45.05% in cases, and mutant AA (Gln/Gln) frequency was 14.88% in control, whereas 26.24% in cases (Table 2). It is a polymorphism which was reported in the population of North-India with 48.9% of controls and 50.0% of patients for Arg/Gln heterozygosity. 36 Study conducted by Matullo et al. 57 reported a 50.0% polymorphism in patients and 44.1% in controls where due to the presence of heterozygous Arg/Gln genotype, the risk of lung cancer occurrence was found by 1.53-fold higher than the wild-type Arg/Arg genotype. Thus, AA (Gln/Gln) frequency shows statistically significant association with lung cancer development (p < 0.05). The GA (Arg/Gln) genotype also possesses the risk of developing lung cancer (p < 0.05). The risk factor for GA (Arg/Gln) and AA (Gln/Gln) variants are 4.57 and 4.99, respectively (Table 2). However, studies like Popanda et al., 58 Hou et al., 24 and Gao et al. 25 failed to correlate XRCC1 rs25487 polymorphism with lung cancer susceptibility.
Furthermore, we also investigated the association of smoking and SULT1A1 rs9282861 and XRCC1 rs25487 polymorphisms and lung cancer risk. Our case-control study reported that current smokers with Arg/Arg and Arg/His genotypes of SULT1A1 are in elevated risk of lung cancer occurrence compared with never smokers with Arg/Arg wild-type genotypes. Similar risk association has also been found in case of former and ever smokers having heterozygous Arg/His genotype (Table 3). For XRCC1 rs25487 polymorphism, only current smokers, carrier of Arg/Arg normal homozygous, showed a statistically significant risk association for lung cancer development in comparison with never smokers having Arg/Arg normal homozygous (Table 4). However, these findings are needed to be substantiated in a larger population.
Conclusion
In conclusion, our results indicate combined Arg/His and His/His genotypes and ‘A’ allele of SULT1A1 rs9282861 are associated with increased lung cancer risk. Our result also highlights a prominent association between the XRCC1 rs25487 polymorphism and lung cancer risk in Bangladeshi population. Both heterozygous Arg/Gln and mutant homozygous Gln/Gln and ‘A’ allele contributes strongly to lung cancer prognosis. Our study will be helpful in correlating gene–gene and gene–environment interactions among different processes of carcinogenesis and smoking. Establishment of this approach could facilitate the association between genetic anomaly in these genes and occurrence of lung cancer.
Footnotes
Acknowledgements
The authors are thankful to the many patients, volunteers, nurses, physicians, and scientists of National Institute of Cancer Research and Hospital (NICRH), Dhaka Medical College Hospital, Ahsania Mission Cancer Hospital, and Bangabandhu Sheikh Mujib Medical University (BSMMU). The authors are also thankful to the Department of Clinical Pharmacy and Pharmacology, Faculty of Pharmacy, University of Dhaka, Bangladesh to give the opportunity to carry out the whole research work. Tasnova Tasnim and Mir Md Abdullah Al-Mamun were involved in sample collection, genetic analysis, and manuscript preparation. Mohammad Safiqul Islam, Maizbha Uddin Ahmed, Noor Ahmed Nahid, Md Reazul Islam, Mohd Nazmul Hasan Apu, Most Umme Bushra, Sikder Nahidul Islam Rabbi, Zabun Nahar, and Jakir Ahmed Chowdhury designed the research work and drafted the manuscript and coordinated the study. A.H. conceived and supervised the study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
