Abstract
Colorectal cancer is a multistep process affecting several signaling pathways including EGFR (epidermal growth factor receptor), a therapeutic target for metastatic disease. Our aim was to characterize the mutational and expression profiles of the EGFR pathway in colorectal tumors and to integrate these results according to five previously defined groups. We screened seven genes for mutations (KRAS-BRAF-PIK3CA-PIK3R1-AKT1-MAP2K1-PTEN) and six proteins (EGFR-p110α-p85α-PTEN-phosphoAKT-phosphoMEK1) by immunohistochemistry, PTEN deletion, and MSI. At least one mutated gene was observed in 68% of tumors (KRAS 45%, PIK3CA 21%, BRAF 14%, and PTEN 7%). PTEN deletion was observed in 10.7% of tumors and 19.6% were MSI-High. In all, 54% of tumors showed a high EGFR expression, 48% p110α, 4.4% phosphoAKT, and 22% phosphoMEK1; and 43% showed low PTEN expression and 22% p85α. In total, five groups of tumors were defined based on MSI, BRAF, and KRAS mutations. Three groups gather mainly early-stage tumors, whereas a fourth group is mostly conformed by advanced tumors. We described here that 71.4% of tumors from one group have a mutated PI3K/PTEN pathway, in comparison to other groups having 32%, 27%, and 25%. In addition, the five groups are differentiated by molecular features such as EGFR, p85α, p110α, and PTEN, showing variable expression among tumor groups. In conclusion, alterations on the EGFR pathway were found in a high percentage of colorectal cancer patients. Using the integration of diverse molecular markers, we ratified previous classification in an ethnic group having relevant genetic differences and living in a different environmental background, adding complementary molecular targets related to therapy.
Introduction
Colorectal cancer (CRC) is one of the three leading causes of cancer mortality in developed countries. 1 In Chile, CRC is the fourth cause of cancer mortality, with a total number of 1867 registered deaths in 2011, according to the Department of Statistics and Health Information. A progressive increase in mortality rates (116%) has been observed over the past 26 years due to risk factors related to age, urban residence, lifestyle, nutrition, and ethnic background. 2
CRC is a multistep process that occurs as a consequence of life-accumulated genetic alterations. 3 The identification of molecular pathways involved in CRC has permitted to define at least three genetic pathways: chromosomal instability (CIN), microsatellite instability (MSI), and CpG island methylator phenotype (CIMP).4,5 CIN is characterized by alterations in the number and structure of chromosomes, affecting tumor suppressor genes and leading to a loss of heterozygosity (LOH). MSI is a consequence of a deficient DNA mismatch repair causing an augmented mutation rate. CIMP is caused by promoter hypermethylation leading into inactivation of specific tumor-suppressor genes. A particular interest has been placed in MSI as a screening method for the identification of Lynch syndrome cases. 6
However, mutations in oncogenes and tumor-suppressor genes participating in the epidermal growth factor receptor (EGFR) signaling pathway occur independently in these three genetic pathways defined in CRC. The EGFR signaling pathway through RAS/RAF/mitogen-activated protein kinase (MAPK) and phosphatidylinositol-4,5-bisphosphate 3-kinase (PI3K)/PTEN/AKT regulates proliferation, differentiation, apoptosis, and angiogenesis.7,8 This pathway is one of the most relevant since EGFR is a therapeutic target for metastatic CRC, and the identification of genetic alterations allows predicting therapeutic efficacy (sensitivity or resistance). Thus, it is well known that some patients will not respond to anti-EGFR therapy if they carry mutations in genes coding for proteins acting downstream EGFR. In this regard, 35%–40% of patients carry KRAS mutations, and among the remaining patients, a proportion will not respond to anti-EGFR therapy because other genes in the pathway such as BRAF, PTEN, or PIK3CA are mutated.9–13
Furthermore, due to different clinical outcomes and different responses to therapy, additional efforts have been made to improve classification of CRC. Jeremy Jass 14 proposed in 2007 the identification of five groups using classical markers such as MSI, CIMP, KRAS mutations, and BRAF mutations. These subgroups are associated with different clinicopathological features, one of them, group 5, gathering Lynch syndrome cases. More recently, Domingo et al. 15 added new markers such as mutations in NRAS, PIK3CA, TP53, and FBXW7/CDC4, obtaining seven CRC groups with different clinicopathological associations and survival. Others have proposed the identification of new markers in relation to gene expression, methylation patterns, and miRNAs; however, no new classifications have had a clinical impact up to now. 16 All these studies have added valuable information; however, tumor heterogeneity of CRC, in addition with the heterogeneity of the populations, support to continue further studies in the characterization of this disease.
Due to the high relevance of the EGFR pathway in CRC, our aim was to define the somatic mutational profile of seven genes (EGFR, KRAS, BRAF, MAP2K1, PIK3CA, PIK3R1, and PTEN) and the protein expression profile of six proteins (EGFR, PI3K catalytic subunit (p110α), PI3K regulatory subunit (p85α), PTEN, phosphoMEK1, and phosphoAKT) and integrate these results according to five subgroups defined by Jass. Our results permitted us to identify in our cohort the five groups of CRC previously described by Jass, adding other mutated genes and protein expression data, all related to the EGFR signal pathway, showing different patterns.
Materials and methods
Patients
Tissue samples (fresh-frozen and formalin-fixed, paraffin-embedded (FFPE)) derived from 56 patients with CRC, who underwent surgical resection at Clínica Las Condes between 2008 and 2013, were analyzed. No patients received radiation or chemotherapy prior to surgery. This study was approved by the institutional review board, and each patient provided written informed consents to donate the tissues after diagnostic procedures. Tumor staging was categorized according to the tumor/node/metastasis (TNM) classification (The Union for International Cancer Control (UICC)).
DNA extraction
First, initially 122 fresh-frozen CRC specimens were collected and analyzed by an experienced pathologist (H.K.) to guarantee that each tissue sample tested contained neoplastic cells in a 70% of the whole tissue. Only 56 tumors accomplished this criterion. DNA was isolated from fresh-frozen tissues for next-generation sequencing (NGS) sequencing and from FFPE sections for methylation, MSI and PTEN deletion analyses. The QIAamp DNA Mini Extraction Kit (Qiagen) was used according to the manufacturer’s protocol. Quality and quantity of total DNA had been determined in Nanodrop-1000 by agarose gel electrophoresis and Qubit 2.0.
MSI determination
The microsatellite status of each tumor was determined by the examination of seven microsatellite markers, as suggested by the National Cancer Institute: Bat-25, Bat-26, Bat-40, D2S123, D3S1029, D5S34619, and D17S250. Each marker was amplified by polymerase chain reaction (PCR) from tumoral and normal DNA from the same patient. PCR products were separated by capillary electrophoresis using the ABI310 sequencer and analyzed by Gene Mapper 4.0 software (Applied Biosystems). MSI status was defined according to the following criteria: two or more (>30%) markers with instability were defined as an MSI-High, one marker with instability was defined as an MSI-low and no markers with instability were defined as a stable microsatellite (MSS).
PTEN deletion analysis
PTEN deletion was determined with three short tandem repeat (STR) markers flanking PTEN (D10S1765, D10S215, and D10S541). Each marker was PCR amplified from tumor and normal DNA, and products were separated by capillary electrophoresis. The allele sizes and relative intensities from the normal and tumor tissues were compared. Markers showing, in normal DNA, two distinguishable alleles of similar intensities were termed as “informative” (heterozygous). Markers that depicted a single allele in the normal DNA were termed as “non-informative” (homozygous). To designate deletion at a specific STR marker, a reduction of the allele intensity in more than 50% was considered. Furthermore, two PTEN intragenic polymorphisms (IVS4+109ins/delTCTTA and IVS8+32T > G) were analyzed in each sample through Sanger sequencing in order to assure deletion or to detect shorter deletions.
MLH1 promoter methylation analysis
Methylation status of MLH1 promoter was examined by methylation-specific PCR analysis. DNA (200 ng) from tumor and normal tissues was modified using EZ DNA Methylation Gold Kit (Zymo Research) according to the manufacturer’s protocol. Modified DNA was then amplified using two sets of primers for methylated and unmethylated DNA from B/C and D regions as published previously. 17 PCR products were visualized by electrophoresis in a 3% agarose gel.
Antibodies
Antibodies against EGFR (D38B1; dilution 1:50), p110α (C73F8; dilution 1:400), PTEN (D4.3; dilution 1:125) and phosphoAKT (Ser 473; D9E; dilution 1:50) were purchased from Cell Signalling Technology. Antibodies against phosphoMEK1 (Ser298) (EPR3338; dilution 1:100) and p85α (M253; dilution 1:100) were purchased from Abcam. Antibodies against MSH6 (44; dilution 1:50) and PMS2 (MRQ-28) were purchased from Cell Marque. Antibodies against MSH2 (25D12; dilution 1:50) and anti-CD3 (LN10; dilution 1:100) were purchased from Novocastra. Antibody against MLH1 (ES05) was purchased from Dako.
Immunohistochemistry
Tissue microarrays (TMAs) containing 46 CRC cases and negative controls were constructed. Non-sufficient quantity or quality tissue was available for 10 tumors. Immunohistochemical staining was carried out on TMAs using antibodies mentioned above. Essentially, the sections were deparaffinized with standard xylene and hydrated through graded alcohols into water. Immunoreactivity retrieval in PT link (Dako) was performed for 20 min at 95°C in an Envision Kit (Dako) high-pH solution. Staining was performed on the automated Autostainer 48 (Dako) using the EnVision (Dako) kit according to manufacturer’s instructions. For nuclear contrast, Mayer’s hematoxylin staining was used. The interpretation of antibody staining was performed considering a scale of four categories: strong (3), moderate (2), low (1), or negative (0).
Library preparation and NGS
EGFR (exons 18–21), KRAS (exons 1 and 2), BRAF (exon 15), PIK3CA (exons 9 and 20), and PIK3R1 (exons 1–11) and the complete coding region of the PTEN and MAP2K1 were sequenced. Libraries of DNA fragments were obtained and captured according to SureSelectXT protocol (Agilent Technologies). Libraries’ quality was determined in a Bioanalyzer 2100, High-Sensitivity Assay; and quantification was determined by real-time PCR in a LightCycler 480 (Roche). NGS was performed at “Sistemas Genómicos SL.”
Statistical analysis
Categorical variables were expressed in numbers and/or percentages. Fisher’s exact test was applied to determine the relationship between two categories including clinicopathological features, protein expression levels and gene mutations. Chi-square test was used to compare the proportion of gene mutations among groups with three different clinicopathological characteristics. All reported p values were two-sided, and statistical significance was set at the 95% level (p < 0.05). The statistical analyses were made using GraphPad Prism software version 5.01.
Results
Clinicopathological features
Table 1 includes clinical information of the 56 CRC patients studied. The study included 33 (59%) male and 23 (41%) female patients, with a median age at diagnosis of 64 years (range: 45–97 years). Seven patients were diagnosed with metastasis at the moment of diagnosis (stage IV; CE021, CE105, CE108, CE124, CE125, and CE132) or during the follow-up (stage III; CE112). Two patients did not have lymph node status since they underwent local resections without lymphadenectomy. Most of the patients had an early-stage disease (60.4%). Four patients died over the course of the study, three of them diagnosed in stage IV (CE021, CE124, and CE125), and one in stage III (CE098). Figure 1 describes the workflow analyses in the 56 patients.
Clinical description of cohort.

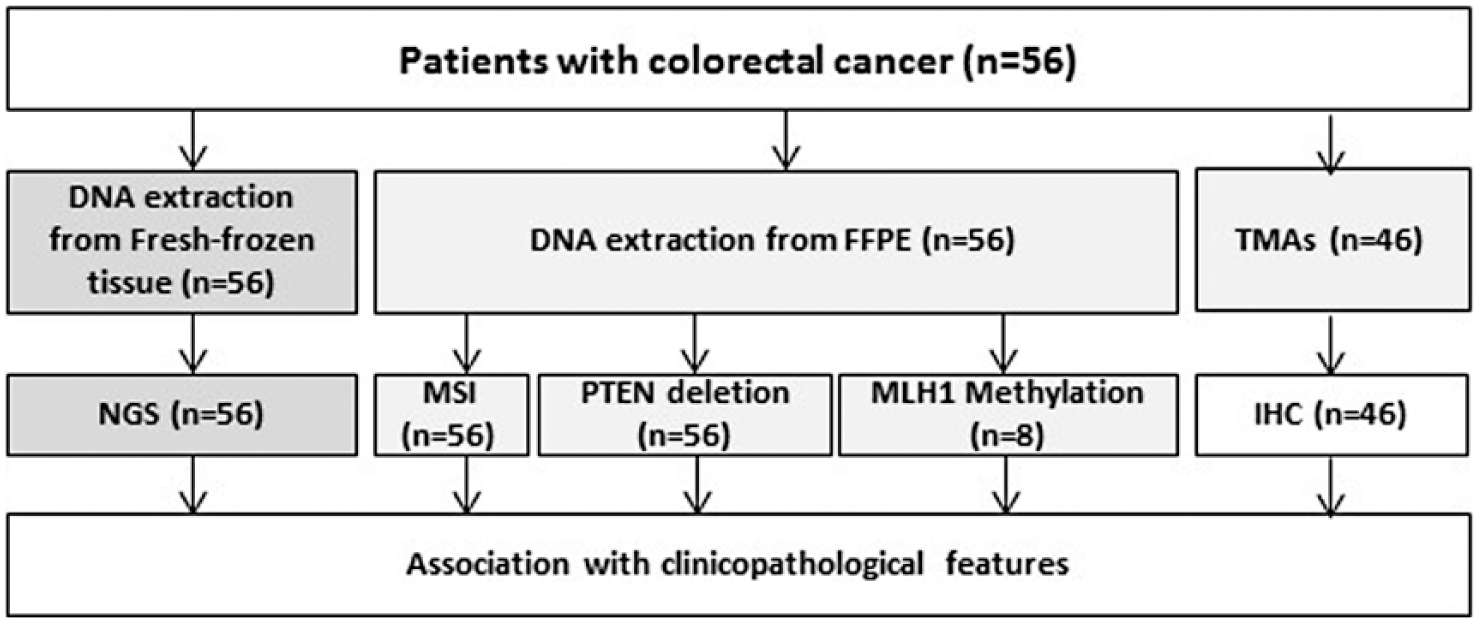
Flowchart of sample analysis.
Mutational analysis and clinicopathological associations
From the 56 tumors sequenced, 68% (38/56) bear at least one genetic alteration in any of the analyzed genes; 45% of patients harbor a KRAS mutation, 21% a PIK3CA mutation, 14% a BRAF mutation, and 7% a PTEN mutation, while no mutation was identified in EGFR, MAP2K1, and PIK3R1. Table 2 shows specific nucleotide alterations, protein changes, COSMIC Mutation ID, tumor ID, and mutation frequency by tumor (mutated/total reads) for each mutated gene.
Frequency of mutations identified in 38 colorectal cancer samples.

KRAS mutations were focused in codon 12 (77%), followed by codons 13 (19%) and 59 (4%). Of the KRAS wild-type tumors, eight harbored a BRAF mutation in exon 15, being V600E the most frequent, while G606E was rarely detected. PIK3CA mutations were identified in 12 tumors, localized in exon 9 (58%) and exon 20 (42%), producing a gain of function. Finally, four different mutations in PTEN were found in five tumors; two of them correspond to frameshift mutations and the other two are intronic variants of uncertain significance.
The mutation frequencies detected in each tumor ranged mainly between 0.13 and 0.75. Only one pathogenic mutation in PTEN was found at a very low level 0.014 (Table 2). The mutational profile showed in Figure 2 includes the results of all somatic mutations found, the MSI status, and PTEN deletions. In this work, six tumors showed PTEN deletion detected through STR markers adjacent to PTEN, in addition to two intronic PTEN polymorphisms. In total, eight tumors have at least one allele affected: two tumors showed a mutation in PTEN (CE069 and CE105), four tumors presented a PTEN deletion (CE021, CE124, CE008, and CE135), and two tumors presented coincident mutation and deletion (CE066 and CE109).

Summary of genetic alterations identified in 56 colorectal cancer samples. Alterations include MSI-High status, somatic mutations, and PTEN deletions.
In summary, the mutational screening revealed (1) co-occurrence of MSI-High with BRAF or KRAS mutations in seven and three tumors, respectively; (2) co-occurrence of PIK3CA mutations and BRAF or KRAS mutations in four tumors each; and (3) co-occurrence of PTEN mutations with any of the mutated genes in three tumors, in addition to two other tumors with a PTEN deletion. Thus, 12 tumors had mutated both intracellular pathways (KRAS/BRAF and PIK3CA/PTEN).
We tested whether the somatic mutational status in CRC correlated with some clinicopathological features, which is shown in Table 3. Associations were found between KRAS and BRAF mutation with clinicopathological characteristics. KRAS mutations were associated with gender (p = 0.029), site of primary tumors (p = 0.0134), lymph node status (p = 0.0426), and tumor stage (p = 0.0214). The presence of BRAF mutation was associated to gender (p = 0.006), site of primary tumor (p = 0.0062), mucinous component (p = 0.0445), and MSI (p < 0.0001). The most significant association was found between KRAS mutations and positive lymph nodes, as well as with an advanced stage of CRC. We did not find significant associations between somatic mutations and age at diagnosis, invasion depth, and metastatic disease.
Association between KRAS and BRAF mutations and clinicopathological characteristics in colorectal cancer.

WT: wild type; MUT: mutated; MSS: microsatellite stable; MSI: microsatellite instability.
All tests were performed with the Fisher’s exact test, in exception for the tumor location that was performed with the Chi-square test.
Unknown data were excluded from the analysis.
Significant values are shown in bold.
As it has been previously described in the literature, mutations in KRAS and BRAF were mutually exclusive in tumors of our study (Table 4). Conversely, PIK3CA and BRAF mutations coincide in 33.3% of tumors (p = 0.05).
Association between KRAS, PIK3CA, and BRAF mutations in colorectal cancer.
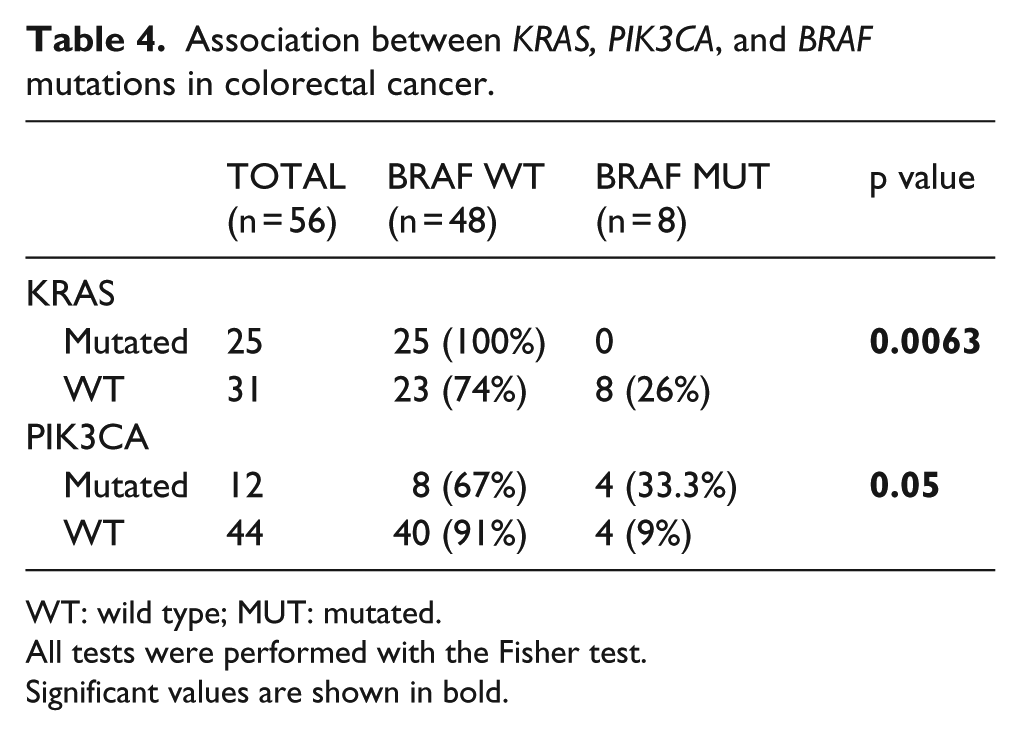
WT: wild type; MUT: mutated.
All tests were performed with the Fisher test.
Significant values are shown in bold.
Immunohistochemistry of EGFR signaling–related protein and correlations
The expression levels in six EGFR signaling–related proteins were determined by immunohistochemistry on tissue microarrays containing 46 CRC tumors. Expression levels were measured with a scale including negative, low, moderate, and strong immunostaining (Figure 3(a)). As shown in Figure 3(b), strong and moderate levels of EGFR were detected in 25/46 tumors (54%), p110α in 22/46 (48%), phosphoAKT (Ser473) in 2/45 (4.4%), and phosphoMEK1 (Ser298) in 10/45 (22%). However, low or negative expression of p85α and PTEN tumor suppressors were found in 10/45 tumors (22%) and 20/46 tumors (43%), respectively.

Immunohistochemical staining of EGFR signaling pathway–related proteins in colorectal cancer samples. (a) Representative images of negative, low, moderate, and strong immunohistochemistry of EGFR (A–D), p110α (E–G), phosphoAKT (H–K), phosphoMEK1 (L–O), p85α (P–S), and PTEN (T–W) proteins. Note that for p110α expression analysis, none of the tumors presented negative expression. Images correspond to amplifications of selected regions of microphotographs 20×. (b) Comparison of colorectal cancer samples with protein expression 0/1+ (negative and low) and 2+/3+ (moderate and high) for each analysis.
We tested whether expression levels of EGFR-related proteins in CRC correlated with clinicopathological and molecular features. Immunostains were grouped in low (negative and low) and high (moderate and high) expressions. As shown in Table 5, associations were observed between EGFR expression levels and BRAF mutation (p = 0.05); p110α expression and location of primary tumors (p = 0.015), BRAF mutation (p = 0.048), and PTEN mutation/deletion (p = 0.003); p85α expression and gender (p = 0.01), BRAF mutation (p = 0.049), and PIK3CA mutation (p = 0.007); and PTEN expression and mutated/deleted PTEN (p = 0.036). No association was found for phospoMEK1 and phosphoAKT.
Association between expression levels of EGFR, p110α, p85α, and PTEN with clinicopathological and molecular characteristics in colorectal cancer (low = scores 0 and 1; high = scores 2 and 3).

CC: colon cancer; EGFR: epidermal growth factor receptor; MSI: microsatellite instability; MSI-High: high-frequency MSI; MSS: microsatellite stable; LOH: loss of heterozygosity.
All tests were performed with the Fisher test.
Significant values are shown in bold.
Identification of CRC subgroups
We analyzed all tumors in order to classify them according to Jass based on MSI, BRAF, and KRAS mutations. 14 As shown in Table 6, group 1, group 2, group 3, group 4, and group 5 included 7, 1, 22, 22, and 4 tumors, respectively. Then, we added the clinicopathological features to each group, in order to complete the features described previously by Jass. A high frequency of women, right colon cancer, mucinous tumors, high levels of CD3+ lymphocytes and early stage characterized group 1. On the contrary, group 3 included mainly men, left colon and rectum cancer, low frequency of mucinous tumors, low frequency of CD3+ lymphocytes, and high percentage of advanced tumors (III and IV). Group 4 was characterized mainly by left colon and rectum cancer and early-stage CRC. Group 5 was represented by right colon cancer in men.
Molecular, clinical, and IHC features of colorectal cancer groups 1–5.

IHC: immunohistochemistry; CRC: colorectal cancer.
Our study contribution is the addition of PI3K/PTEN pathway mutational analysis and the expression analysis of proteins involved in the EGFR signal transduction pathway. Interestingly, from group 1, four tumors have a mutation in PIK3CA and one in PTEN, leading to a 71.4% (5/7) of tumors belonging to group 1 having a PI3K-/PTEN-mutated pathway. In the other groups, a low frequency of mutations was observed in this pathway: group 3 (32%), group 4 (27%), and group 5 (25%). In relation to protein expression detected by IHC, group 1 shows a 100% of tumors with high EGFR and 50% of tumors with low p85α. Groups 3 and 5 are characterized mainly by a high expression of p110α, while group 4 shows a high frequency of tumors with low PTEN.
Identification of Lynch syndrome cases
As defined previously, some molecular features, such as MSI status, DNA repair protein’s expression levels, MLH1 promoter methylation status, and BRAF mutations, allow to classify CRC tumors as sporadic or hereditary. In this work, 9/43 (20.9%) tumors show loss of expression of DNA repair proteins (MLH1, MSH2, PMS2, and MSH6; Table 7). These results suggest that six patients correspond to sporadic CRC cases because of MLH1 methylation, and three patients could be hereditary cases (CE021, CE109, and CE112). These patients’ clinical features are shown in Table 7; none of them has a family history of cancer. Germline mutations screening revealed the existence of an MLH1 mutation c.2041G > A (p.Ala681Thr) in CE112 patient, while MLH1 mutation was not identified in subject CE109.
Molecular and clinical characteristics of MSI-High tumors and/or loss expression of DNA repair proteins.

IHC: positive (+) and negative (−); UM: unmethylated; M: methylated; WT: wild type; ND: not determined.
Discussion
In this study, we determined mutation frequencies of KRAS, BRAF, PIK3CA, PIK3R1, PTEN, EGFR, and MAP2K1 and protein expression levels of EGFR, p110α, p85α, PTEN, phosphoMEK1, and phosphoAKT in Chilean CRC patients. We examined associations between molecular markers and clinicopathological characteristics. In addition, we ratified the Jass classification and added molecular targets of EGFR signaling pathway in a population with significant genetic differences and different environmental conditions from other populations previously studied.18,19
The EGFR signaling pathway is one of the most relevant in CRC development, prognosis, and treatment, acting through two intracellular pathways: KRAS/BRAF/MAPK and PI3K/AKT/mammalian target of rapamycin (mTOR), in which key proteins promote or participate in pathways promoting cell proliferation. In this study, we found that at least 68% of tumors presented a mutation in one or two of the analyzed genes. The highest mutation rate was found in KRAS (45%), followed by PIK3CA (21%), BRAF (14%), and PTEN (7%). No mutation was identified in EGFR, PIK3R1, and MAP2K1, in concordance with the low-mutation frequencies described in other reports and in COSMIC database.20,21 Our findings are similar to what was observed in previous studies using NGS.22,23 Previous reports in Chilean population are consistent with our results: 42% KRAS mutation and 12% BRAF mutation.24,25 This study confirms the usefulness of assessing mutational detection in EGFR pathway–related genes by NGS as part of the characterization of CRC.
Associations were mainly found for KRAS and BRAF mutations, with different clinicopathological features such as gender, site of primary tumors, mucinous component, MSI, lymph node status, or tumor stage, which is consistent with what has been widely described in the literature.26,27 Difficulty in establishing an association to survival may be explained by the early detection of the disease in our cohort, showing a 95% survival and including a small number of patients in stage IV. Previously, PTEN and PIK3CA mutations have been associated with gender, site of primary tumor, MSI status and stage, among others. 28 In this study, we found a higher frequency of PIK3CA mutations in women, mucinous carcinoma, MSI-High, and negative lymph nodes; however, no association was found between PIK3CA mutations in women and these clinicopathological features. In the case of PTEN mutations, the absence of association could be explained by the low percentage of mutated samples (7%) in this gene. Alternatively, the absence of association in our cohort may be explained by the genetic differences among ethnic groups.
Noteworthy is that in some cases, the evaluation by IHC gives additional information in relation to protein expression, when mutations or other molecular features are not observed. In this study, overexpression of EGFR and p110α was observed in 54% and 48% of tumors, respectively. The absence of point mutations in these genes suggests other mechanisms of overexpression, previously described in the CRC, such as gene amplification. 20 However, PTEN expression analysis has shown a decreased or negative protein level in 46% of tumors, but no mutation or deletion was detected. Similar results have been described in other types of cancer, suggesting that other mechanisms of inactivation leading to a loss of protein expression must be involved, such as the level of gene expression regulation. 29 PTEN promoter methylation was initially described in some cancer tumors;30,31 however, it was later demonstrated that methylation occurs in the PTEN pseudogene.31,32 Actually, miRNAs targeting PTEN have been described to regulate its expression in cancer, as well as alterations in the PTEN protein turnover by deregulation of post-translational modification. 33 Although mutations in the gene for PIK3R1 were not identified in our cohort, IHC analysis showed a decrease or absence of the protein in 24% of tumors. Recently, numerous miRNAs have been described as regulators of this gene in other types of cancer, which are probably participating in the regulation of protein expression and in CRC development.34–36
Until today, several efforts have been made to find markers allowing a classification of CRC in order to improve patient’s clinical outcome and therapies.14–16,37 Even if those studies permitted a better clinicopathological classification of tumors, no improvement in translational medicine has been made. One of the best known classifications for CRC was defined by Jass, which includes the recognition of five molecular groups mainly defined by MSI, BRAF, and KRAS analyses. 14 These molecular markers have clinical utility themselves or have been associated with prognosis or therapy response. In this way, the presence of KRAS mutations is used as a predictor of resistance to anti-EGFR therapy in metastatic colorectal cancer; MSI-High is a marker for better prognosis and MSI-High in the absence of mutated BRAF predicts Lynch syndrome cases.6,10,38 Tumors with mutated BRAF in the absence of MSI-High have been associated with a poorer clinical outcome.39,40
In this work, we were able to identify the five groups defined by Jass out of 56 tumors from Chilean CRC patients. Similar to what has been described by Jass, we found that demographic, clinical, and pathological features were related with each molecular group. Groups 1, 4, and 5 show a high frequency of tumors in CRC early stages, while group 3 had a worse prognosis due to a high frequency of tumors in advanced stages including three patients who died. Group 2 is the less represented in our cohort of patients, including only one patient who was diagnosed with CRC at stage IV, was BRAF mutated, and died at 59 years.
Through the incorporation of PI3K/AKT/mTOR pathway in Jass classification, we found that PIK3CA and PTEN mutations are distributed in different groups of CRC, being more frequent in group 1. Indeed, PIK3CA and PTEN mutations can occur in conjunction with KRAS and BRAF mutations as was observed in 12 of the 56 tumors. Mutation identification in these genes could define a potential therapy response, redirecting patients who are refractory to anti-EGFR to a more effective therapy like mTOR inhibition. Interestingly, MSS tumors without KRAS and BRAF mutations present in group 4 carry mutations in PIK3CA and PTEN that could help to define patients with different clinicopathological features. In this regard, the identification of PIK3CA mutations serves as a predictive marker for the use of aspirin after diagnosis of CRC. In fact, regular use of aspirin has been widely associated with longer survival rates in patients with mutated PIK3CA.41,42
Conclusion
In summary, this study may improve the current classification of CRC by adding mutational and expression profiles of the EGFR signaling pathway. The possibility to integrate results from different tumor analyses and patient’s clinical features will allow improving CRC treatment.
Footnotes
Acknowledgements
We also thank all participants for their participation and contribution to this study. K.A. and P.O. contributed to design concept, data acquisition, data analysis and interpretation, and drafted the manuscript. H.K. and M.K. were responsible for the histological classification of fresh-frozen CRC samples. P.O. and C.V. carried out DNA isolation for mutation screening by next-generation sequencing. C.V. performed MLH1 methylation studies. A.M.W analyzed the MSI status of CRC samples. L.C. performed data acquisition, analysis, and interpretation of immunohistochemistry assay. J.C.T. performed the statistical analysis. F.L.-K. and U.K. contributed to concept design and patient recruitment and reviewed the manuscript critically. M.D.L.F. helped to draft the manuscript. P.C. contributed to data analysis, results’ interpretation, and manuscript review. All authors read and approved the final manuscript.
Declarations of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics approval
This study was approved by the ethics committee from Clínica Las Condes, Santiago, Chile. All patients gave their written informed consent.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the grants (1111020 and 1140012) from the National Fund for Scientific and Technological Development (FONDECYT), Santiago, Chile.
