Abstract
Hepatocellular carcinoma is one of the most prevalent neoplasms and the leading cause of cancer-related mortality worldwide. Mitochondrial ribosomal protein S23 is encoded by a nuclear gene and participates in mitochondrial protein translation. Mitochondrial ribosomal protein S23 overexpression has been found in many types of cancer. In this study, we explored mitochondrial ribosomal protein S23 expression in primary hepatocellular carcinoma tissues compared with matched adjacent non-tumoral liver tissues using mitochondrial ribosomal protein S23 messenger RNA and protein levels collected from public databases and clinical samples. Immunohistochemistry was performed to analyze the relationship between mitochondrial ribosomal protein S23 and various clinicopathological features. The results indicated that mitochondrial ribosomal protein S23 was significantly overexpressed in hepatocellular carcinoma. High mitochondrial ribosomal protein S23 expression was correlated with the tumor size and tumor–metastasis–node stage. Moreover, patients with high mitochondrial ribosomal protein S23 expression levels presented poorer survival rates. Mitochondrial ribosomal protein S23 was an independent prognostic factor for survival, especially at the early stage of hepatocellular carcinoma. In addition, the downregulation of mitochondrial ribosomal protein S23 decreased the proliferation of hepatocellular carcinoma in vitro and in vivo. In conclusion, we verified for the first time that mitochondrial ribosomal protein S23 expression was upregulated in hepatocellular carcinoma. High mitochondrial ribosomal protein S23 levels can predict poor clinical outcomes in hepatocellular carcinoma, and this protein plays a key role in tumor proliferation. Therefore, mitochondrial ribosomal protein S23 may be a potential therapeutic target for hepatocellular carcinoma.
Introduction
Hepatocellular carcinoma (HCC) is one of the most complicated cancers, and its incidence has increased worldwide in recent years. 1 HCC is derived from hepatocytes and represents the most common histological liver cancer subtype, 2 accounting for almost 90% of liver cancers. 3 The latest data show that HCC is the most common malignant cancer and the leading cause of cancer-associated deaths worldwide.4,5 Disturbingly, the rapid increase in the number of HCC patients worldwide is due to the increased incidence of viral infections 6 and metabolic diseases. 7 In most cases, HCC develops in individuals with chronic hepatitis B or hepatitis C virus infections. 8 Patients with HCC often have other diseases, which makes HCC treatment more complex and variable between individuals.
At present, liver resection remains the standard and most beneficial treatment for most HCC patients, with the exception of patients with terminal HCC. 9 A high incidence of tumor recurrence and metastasis is the main obstacle for the survival of HCC patients. 10 Although pathological classification is currently the most important tool to estimate the HCC prognosis, specific markers are needed to assist in the diagnosis and treatment of HCC, especially in the early stages. 3 The most commonly used clinical indicator is alpha-fetoprotein (AFP). However, because the limitations and insidious onset of HCC often make early detection and diagnosis difficult, 11 most HCC patients are diagnosed in the intermediate or advanced disease stages. Successful curative and palliative treatments are difficult to achieve without early detection of HCC.3,12 In addition, the underlying mechanisms that promote the pathogenesis of this deadly disease need further investigation. In particular, the use of gene expression data to predict the progression and prognosis of HCC is promising, especially in the early stages of tumor development. 10 Thus, developing more effective and novel genetic biomarkers for the early detection and treatment of HCC is crucial because these markers can aid in targeted gene therapy for HCC. 13
Mitochondria are bioenergetic, biosynthetic, signaling organelles that are involved in the adaptation to environmental changes. 14 Mitochondria are the main energy and power centers in cells and generate almost 90% of cellular energy. Recently, mitochondria have been found to not only be involved in energy metabolism but also participate in many other important functions, including apoptosis, autophagy, and cell death. 15 Mitochondria have their own DNA (mtDNA) and a unique translation system, which contains approximately 70 protein components and produces 13 proteins essential for oxidative phosphorylation. 16
Mitochondria are also important mediators of tumorigenesis, and many studies have focused on the relationship between mitochondria and tumorigenesis. 14 A group of unique proteins called mitochondrial ribosomal proteins (MRPs) are completely encoded by nuclear genes but mainly assist with the mitochondrial protein translation process within the mitochondria. 16 In addition, MRPs may have other functions in tumorigenesis.14,16,17
Here, we report the mitochondrial ribosomal protein S23 (MRPS23) gene, which encodes a 28S subunit protein. Early research found that MRPS23 was overexpressed in breast cancer 18 and uterine cervical cancer19,20 and suggested that MRPS23 overexpression might be a driver of proliferation in the luminal subtype of human breast cancer. 18 In addition, overexpressed MRPS23 is a molecular marker for advanced cervical carcinomas and is strongly associated with rapid proliferation, oxidative phosphorylation, invasiveness, and tumor size. 19 However, few studies have investigated the roles of MRPS23 in HCC.
In this study, we used reverse transcription quantitative polymerase chain reaction (RT-qPCR), western blotting, and immunohistochemistry (IHC) to examine MRPS23 messenger RNA (mRNA) and protein expression in HCC tissues and adjacent non-tumoral liver tissues (ANLTs). In addition, we analyzed data from The Cancer Genome Atlas (TCGA) online databases to explore whether an MRPS23 expression signature was predictive of the survival prognosis. We used IHC to explore the relationship between MRPS23 expression and some clinicopathological features. MRPS23 protein expression was also analyzed to investigate its association with clinicopathological features and the prognosis. In vitro and in vivo studies were performed to investigate the function of MRPS23 in HCC cell lines. The findings of this study identify MRPS23 as a novel biomarker for predictions of the HCC prognosis and expound on its function in HCC cell lines. The in vitro and in vivo experiments show that MRPS23 may be a candidate oncogene that affects the proliferative capacity of HCC cells. However, the detailed molecular mechanisms by which MRPS23 contributes to HCC are unknown and require further research.
Materials and methods
Clinical specimens
Matched HCC specimens and ANLTs were randomly obtained from patients during hepatic resection at the Department of Hepatobiliary Surgery, Xijing Hospital of the Fourth Military University, from 2010 to 2014. All specimens were collected using the same standardization process and were confirmed by pathological examination. We randomly selected 8 paired specimens for protein extraction and 50 paired specimens for total RNA extraction. The extractions were performed according to the standard protocols.
Data collection and analysis from the TCGA database
All HCC samples from TCGA were collected from the TCGA website (https://tcga-data.nci.nih.gov/) and used following strict human subject protection guidelines. A total of 50 pairs of samples (HCC vs ANLTs) were chosen for the MRPS23 expression profile analysis. As for survival analysis, a total of 342 available data were filtered and used for the statistical analysis. TCGA RNA sequencing and corresponding clinical data were downloaded following approval of this project by the consortium. Then, these data were filtered and standardized before they enter the follow-up statistical analysis. In order to avoid errors caused by inappropriate grouping, a quality control is needed after standardization, the data were dichotomized into high-level and low-level groups, followed by the Kaplan–Meier survival analysis.
HCC tissue microarray and IHC analysis
Commercially available tissue microarray (TMA) was purchased (HLivHCC180Sur-05; Shanghai Biochip Co., Ltd, Shanghai, China). This TMA contained 85 matched pairs of primary HCC samples with adjacent liver tissues and 10 additional HCC samples. All the samples were applied to evaluate the prognostic value of MRPS23 based on its detailed survival data. The sections were dewaxed with xylene, gradually hydrated, and then boiled in 10 mM citrate buffer (pH 6.0) for 5 min for antigen retrieval. Endogenous peroxidases were inactivated with 3% H2O2. Then, the sections were blocked with goat serum, incubated with primary antibody diluted at 1:200 (MRPS23; Sigma-Aldrich, St. Louis, MO, USA) for overnight at 4°C, and incubated with biotinylated secondary antibody at room temperature following the instructions for 1 h. Finally, the sections were visualized with diaminobenzidine (DAB Kit; ZSGB-BIO, Beijing, China), and hematoxylin was used to counterstain the nuclei.
Analysis and quantification of staining
Immunohistochemical staining was examined by two pathologists using a double-blind method. An immunoreactivity score (IRS) was applied wherein the staining intensity (SI) was graded as follows: negative = 0, weak = 1, moderate = 2, and strong = 3. The percentage of positively (PP) stained cells was scored in a range from 0–3 as follows: 0 (≤10%), 1 (11%–50%), 2 (51%–75%), and 3 (≥5%). The following calculation was applied: IRS = SI × PP, where IRS >3 was defined as positive and IRS ≤3 was defined as negative.
Cell lines and culture
Human HCC cell line MHCC97-H and SMMC-7721 were obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). Cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM; HyClone, Logan, UT, USA) with 10% fetal bovine serum (FBS; Tianhang Biotechnology, Zhejiang, China) in a humidified incubator containing 5% CO2 at 37°C.
Lentiviral infection
Lentiviral plasmids negative control (sh-NC) and interference for human MPRS23 (sh-MRPS23) were designed and produced from GeneChem (Shanghai, China). MHCC97-H and SMMC-7721 cells were cultured in six-well plates at a certain density. Transfection process was carried out with 20 multiplicity of infection (MOI) according to the instructions. After 72 h, the infection rate was detected by a fluorescence microscope. Then RT-qPCR and western blotting were carried out to detect the expression of MRPS23.
RNA extraction and RT-qPCR
Total RNA was isolated by TRIzol reagent (Thermo Fisher Scientific, Shanghai, China) according to the manufacture’s protocol. MRPS23 mRNA expression was analyzed by RT-qPCR. β-Actin mRNA was used as the reference. mRNA level was determined by SYBR Premix EX Taq™ II (TaKaRa, Dalian, China) on Bio-Rad IQ5 System (Bio-Rad, Hercules, CA, USA). The primers used for RT-qPCR were as follows—MPRS23: 5′-GGTTTGACGTATATGACGCCTT-3′ (forward) and 5′-CTCTAATCCGATCCTCGTGGTA-3′ (reverse); β-actin: 5′-CATGTACGTTGCTATCCAGGC-3′ (forward) and 5′-CTCCTTAATGTCACGCACGAT-3′ (reverse). Relative gene expression values were quantified by 2−ΔΔCt method. All measurements were performed three times.
Western blotting
The protein samples were prepared according to the normal procedure with lysis buffer (Beyotime, Shanghai, China) containing protease inhibitors (CoWin Bioscience, Beijing, China). Then, same amount of protein sample was separated by 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis and transferred to the polyvinylidene difluoride membranes (Millipore, Billerica, MA, USA). Then, the membranes were incubated overnight at 4°C with primary antibodies MRPS23 (1:1000; Sigma-Aldrich) and β-actin (1:2000; Proteintech, Wuhan, China) and were washed with Tris-buffered saline 0.1% Tween for three times. The membranes were incubated with corresponding horseradish peroxidase (HRP)-conjugated secondary antibody (1:3000; Proteintech) at room temperature for 2 h. The blots were visualized and captured by Bio-Rad ChemiDoc™ XRS+ and Image Lab™ Software (Bio-Rad).
Immunofluorescence
Immunofluorescence staining of cells was performed as previously described. Briefly, the samples were fixed in 4% paraformaldehyde solution at room temperature for 20 min and permeabilized in 0.5% Triton X-100 in phosphate-buffered saline (PBS) for 10 min at 4°C. The cells were then blocked with 3% bovine serum albumin (BSA) and incubated with primary antibody against MRPS23 (1:200; Proteintech) followed by a fluorescein isothiocyanate (FITC)-conjugated secondary antibody (ZSGB-BIO), counterstained with 4′,6-diamidino-2-phenylindole (DAPI; Beyotime), and visualized using a fluorescence microscope (Olympus, Tokyo, Japan).
Cell proliferation and colony formation assay
The Cell Counting Kit-8 (CCK-8) was used to measure cell proliferation in the different groups on days 1, 2, 3, 4, and 5 according to the manufacturer’s instructions. For all experiments, the cells were plated at a density of 2000 cells per well in 96-well plates. The absorbance of each well was measured 2 h after incubation with the CCK-8 reagents. Each experiment included five replicates and was repeated three times. The colony formation assay was used to estimate cell proliferation. Briefly, 400 cells were seeded onto a six-well plate and cultured for 14 days in DMEM supplemented with 10% FBS. The colonies were fixed with 95% ethanol and stained with a 4 g/L crystal violet solution. Each experiment was repeated three times.
Cell migration and invasion assays
The cell migration and invasion assays were performed using Transwell chambers (8-µm pore size; Millipore) without (migration assay) or with (invasion assay) Matrigel (BD Biosciences, San Jose, CA, USA). Briefly, 500 µL of complete medium was added to the bottom of the chamber, and 200 µL of serum-free medium containing 5 × 104 cells was added to the upper chamber. After 24 h, the top surface of the insert membrane was removed using a cotton swab, and the cells on the bottom surface of the insert were fixed with 95% ethanol and stained with crystal violet solution. Finally, the number of cells in five randomly selected areas was counted under a microscope. Each experiment was repeated three times.
In vivo tumor growth
For the tumorigenesis assay, transfected MHCC97-H cells (2 × 106) were suspended in 200 µL of PBS and subcutaneously injected into nude mice (n = 5 mice). The tumors were harvested and weighed after 25 days. The tumor tissues were formalin-fixed and paraffin-embedded for hematoxylin and eosin (H&E) staining and IHC. Male nude mice (BALB/C nu/nu; 5 weeks old) were obtained from the Animal Center of the Chinese Academy of Sciences (Shanghai, China) and maintained under specific pathogen-free conditions in the Laboratory Animal Center of The Fourth Military Medical University (Xi’an, China). All experimental procedures were performed in accordance with the Guide for the Care and Use of Laboratory Animals and were approved by The Research Animal Care and Use Committee of The Fourth Military Medical University.
Statistical analysis
All data were analyzed using the statistical software SPSS 22.0 for Windows (IBM, Chicago, IL, USA) and GraphPad Prism 6 for Windows (GraphPad Software, La Jolla, CA, USA). The data are generally expressed as the mean ± standard deviation (SD). The differences between groups were analyzed using Student’s t test when comparing two groups with homogeneous variance. The χ2 analysis was used to analyze correlations between MRPS23 expression and the clinicopathological features. The p values for the Kaplan–Meier survival analysis were generated by the log-rank test. Cox proportional hazards regression analysis was used to identify risk factors. Univariate and multivariate analyses were performed to examine the influence of each clinical variable on survival. Significance was two-tailed, and significant differences were defined as p < 0.05.
Results
MRPS23 mRNA and protein levels are highly expressed in HCC tissues
Data from the TCGA analysis showed that MRPS23 expression was higher in the HCC tissues than in the ANLTs (Figure 1(a)). To confirm this result, we quantified the MRPS23 expression levels in 50 pairs of HCC and ANLTs using RT-qPCR. As shown in Figure 1(b), MRPS23 mRNA was significantly overexpressed in the HCC tissues compared with the levels seen in the ANLTs (p < 0.05). Next, we investigated MRPS23 protein expression by western blotting. In total, 8 pairs of HCC samples were assayed by western blotting, as shown in Figure 1(c). MRPS23 was overexpressed in HCC tissues from 5/8 patients (62.5%), which was consistent with the MRPS23 mRNA expression results.
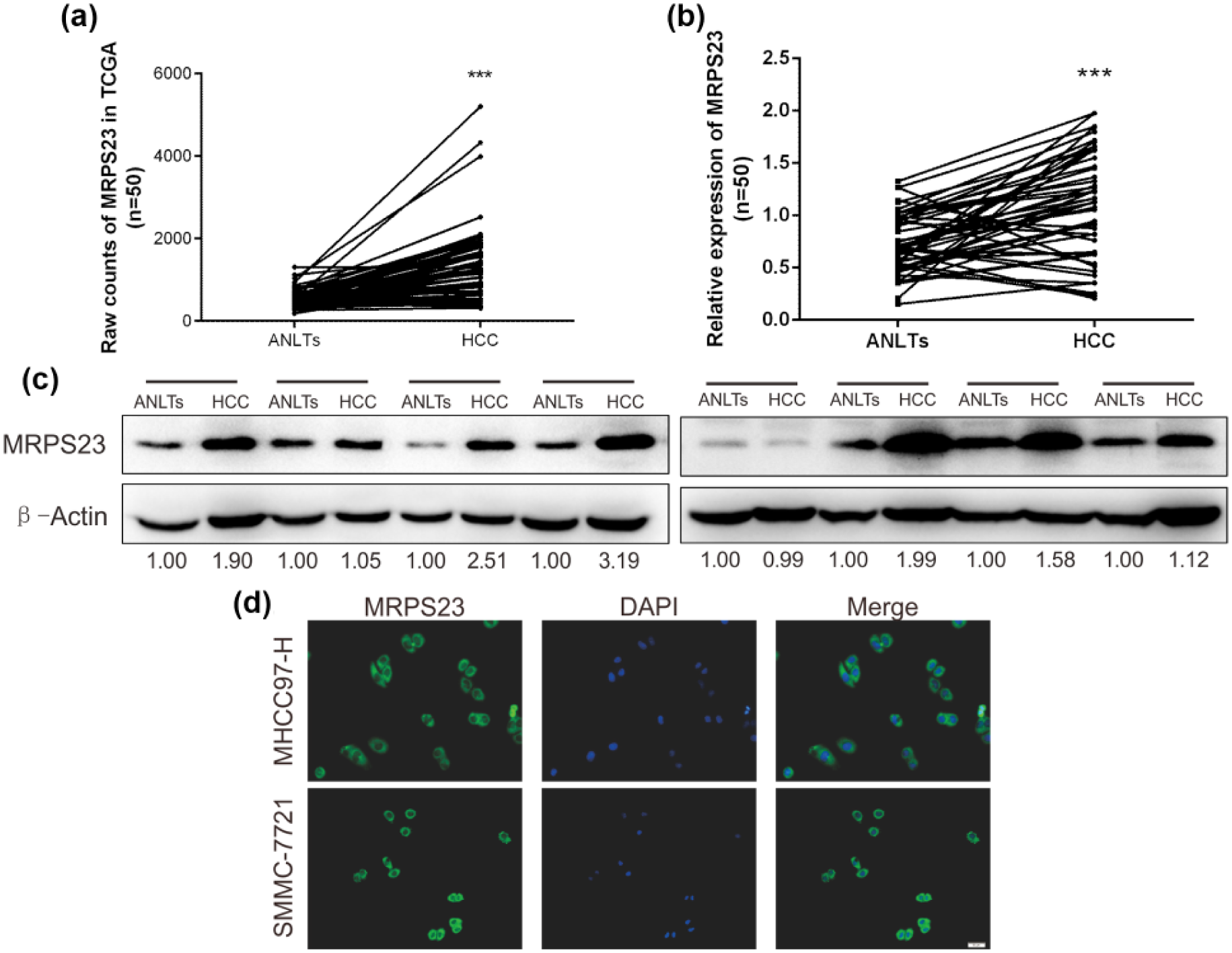
Expression analyses of MRPS23 in HCC and ANLTs. (a) Expression of MRPS23 in TCGA database (n = 50). (b) The mRNA levels of MRPS23 in 50 pairs of HCC and ANLTs by RT-qPCR (n = 50). (c) The protein levels of MRPS23 in 8 pairs of HCC and ANLTs were detected by Western blot analysis. (d) Immunofluorescence was carried out to identify the distribution of MRPS23 in HCC cells (***p < 0.001).
Next, we used IHC to confirm MRPS23 expression in the HCC and ANLT tissues. We found that MRPS23 expression was significantly upregulated in the HCC tissues. In the TMA, 65/95 (68.42%) of the HCC tissues showed high MRPS23 expression, whereas only 24/85 (28.24%) of the ANLTs showed high MRPS23 expression. Thus, the MRPS23 levels were significantly increased in the HCC tissues compared with those in the ANLTs (p < 0.05, Table 1). Moreover, the immunofluorescence analysis performed to confirm this result showed that MRPS23 was primarily expressed in the cytoplasm. Indeed, Figure 1(d) showed that MRPS23 was mainly expressed in the cytoplasm in both the MHCC97-H and SMMC-7721 cells.
Expression of MRPS23 in HCC by IHC.
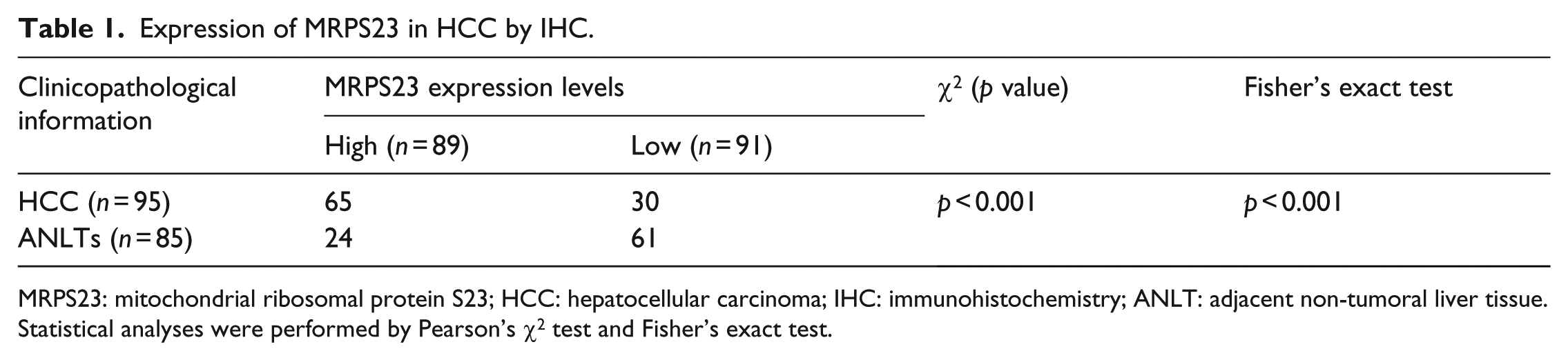

MRPS23: mitochondrial ribosomal protein S23; HCC: hepatocellular carcinoma; IHC: immunohistochemistry; ANLT: adjacent non-tumoral liver tissue.
Statistical analyses were performed by Pearson’s χ2 test and Fisher’s exact test.
High MRPS23 expression is associated with a poor prognosis in HCC patients
After confirming that MRPS23 was abnormally expressed in HCC, we investigated the relationship between MRPS23 and clinical prognostic features. The survival information for the 342 patients was divided into two groups according to the median MRPS23 expression level (high MRPS23: n = 172 and low MRPS23: n = 170). Kaplan–Meier analysis with a log-rank test was performed to evaluate the influence of MRPS23 on the overall survival (OS) of HCC patients. The patients in the high MPRS23 expression group had significantly lower OS than the patients in the low-expression group (Figure 2(a)). Then, the HCC patients were divided into three groups (Stage 1, Stage 2, and Stage 3/4) according to their clinical stages (American Joint Committee on Cancer standard). The results showed that high MRPS23 expression was mainly associated with a poor prognosis in Stage 1 patients (Figure 2(b)). In contrast, the p values obtained for the Stage 2 (Figure 2(c)) and Stage 3/4 (Figure 2(d)) patients were insignificant possibly due to limitations in the sample size; these limitations might be overcome by studying a larger sample. Our data indicate that the value of MPRS23 may be more significant and sensitive in Stage 1 patients. Thus, MRPS23 expression may be a prognostic indicator for early-stage HCC.
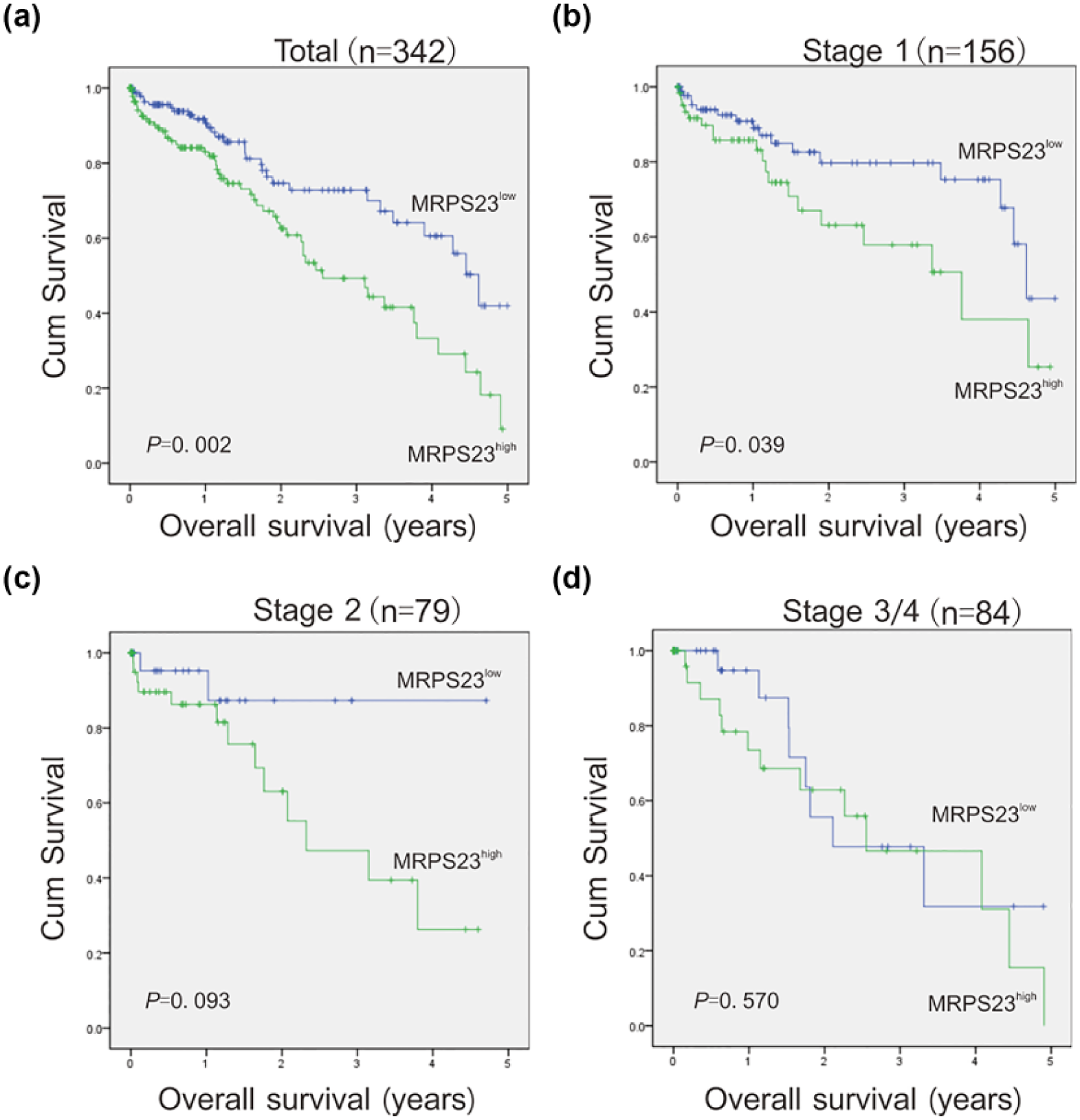
Survival analysis of HCC patients based on MRPS23 expression by Kaplan–Meier method. (a) Overall survival rate in total HCC patients with high/low MRPS23 expression. (b–d) Overall survival rate in the different clinical stages of HCC patients with high/low MRPS23 expression. The clinical stages were based on the AJCC criteria. Totally, 342 samples were taken into two groups: total (MRPS23high: n = 172 and MRPS23low: n = 170), Stage 1 (MRPS23high: n = 64 and MRPS23low: n = 92), Stage 2 (MRPS23high: n = 48 and MRPS23low: n = 31), and Stage 3/4 (MRPS23high: n = 48 and MRPS23low: n = 36). Clinical stage of 23 patients were unclear.
To validate this result, IHC was performed on HCC tissues and ANLT specimens to evaluate MRPS23 expression. The result showed significant difference between HCC tissues and ANLTs (Figure 3(a)). TMA was used to detect the expression of MRPS23. The HCC tissues displayed much stronger staining of MRPS23 than the ANLTs (Figure 3(b) and (c)). Data presented in Tables 1 and 2 demonstrates that MRPS23 had significantly upregulated expression in HCC. The patients with high MRPS23 indeed had a shorter OS than the low-expression group (Figure 3(d)). The relationship between MRPS23 and various clinicopathological characteristics of patients was showed in Table 2. High MPRS23 expression may be correlated with tumor size (p < 0.05) and tumor–metastasis–node (TNM) stage (p < 0.05). Univariate Cox regression analysis indicated that MRPS23 expression, metastasis, and TNM stage were significantly associated with the OS of patients with HCC. Furthermore, multivariate Cox regression analysis demonstrated that MRPS23 expression, metastasis, and TNM stage were the most valuable independent prognostic factors for the survival of HCC patients (Table 3). In conclusion, these results indicate that high expression of MRPS23 is related to HCC progression and can be a prognostic indicator of early stage HCC.

The protein expression of MRPS23 in HCC by immunohistochemistry. (a) The representative HCC tissues with ANLT images of HCC frozen tissue show the staining of MRPS23. (b) The representative TMA images show the different staining intensities of MRPS23 in HCC and ANLTs. (c) The IRS scores of MRPS23 in HCC tissues and ANLTs by the immunohistochemical staining of TMA. (d) Kaplan–Meier analysis of the overall survival of patients with MRPS23 high/low expression based on the TMA immunohistochemical staining results (***p < 0.001).
Correlations between MRPS23 expression and clinicopathological variables in tissues of HCC patients.

MRPS23: mitochondrial ribosomal protein S23; HCC: hepatocellular carcinoma; TNM: tumor–node–metastasis.
Statistical analysis was performed by Pearson’s χ2 test. Gender, metastasis, and tumor number were analyzed by Fisher’s exact test.
Significant values.
Univariate and multivariate analyses of factors associated with survival.
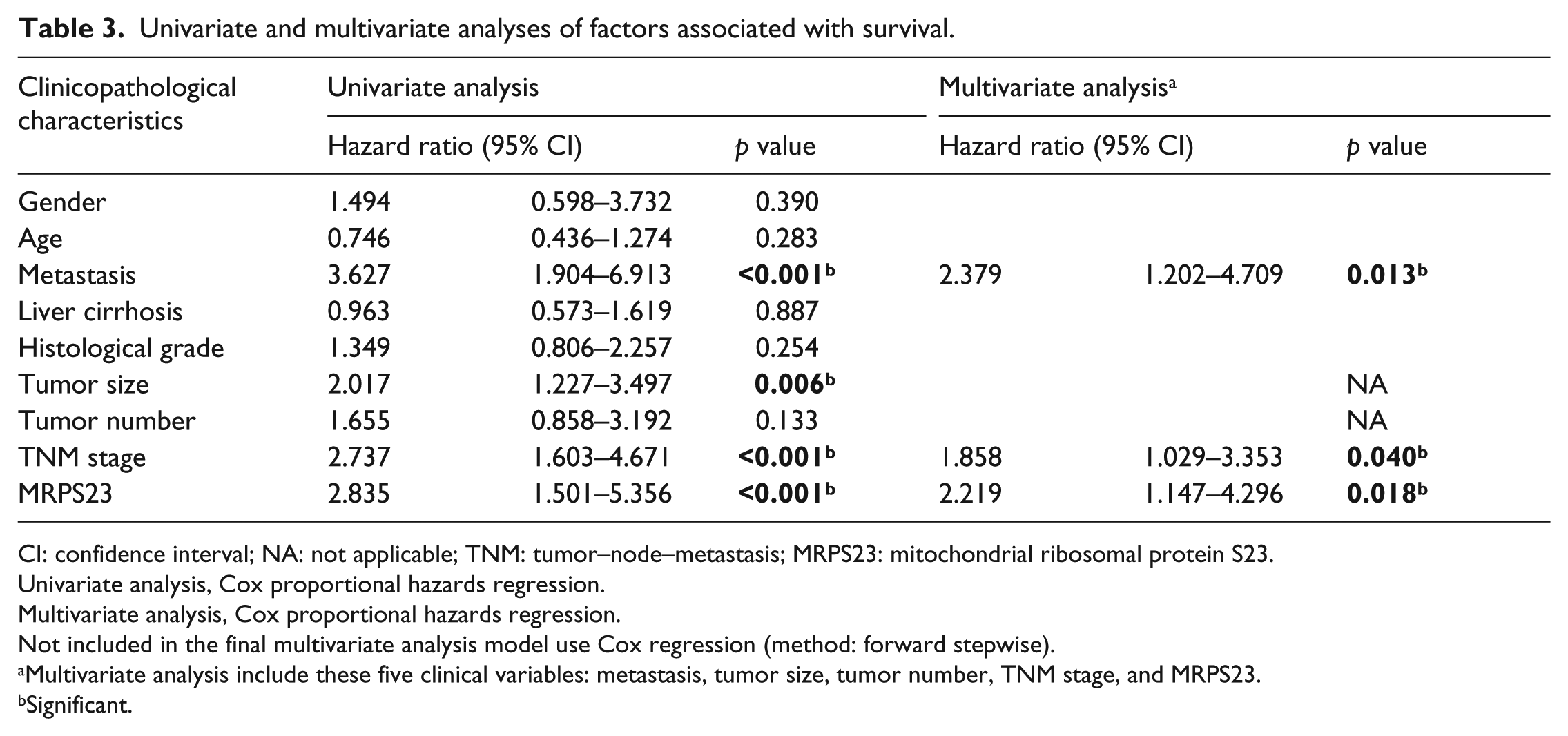
CI: confidence interval; NA: not applicable; TNM: tumor–node–metastasis; MRPS23: mitochondrial ribosomal protein S23.
Univariate analysis, Cox proportional hazards regression.
Multivariate analysis, Cox proportional hazards regression.
Not included in the final multivariate analysis model use Cox regression (method: forward stepwise).
Multivariate analysis include these five clinical variables: metastasis, tumor size, tumor number, TNM stage, and MRPS23.
Significant.
Downregulation of MRPS23 attenuates HCC cell proliferation in vitro
To investigate the role of MRPS23 in HCC progression, we downregulated MRPS23 expression by transfecting the HCC cell lines MHCC97-H and SMMC-7721 with lentiviral particles. A negative control was also included. The RT-qPCR and western blotting assays were used to verify MRPS23 downregulation. The results showed that MRPS23 was significantly downregulated in the MHCC97-H and SMMC-7721 cells (Figure 4(a) and (b)).
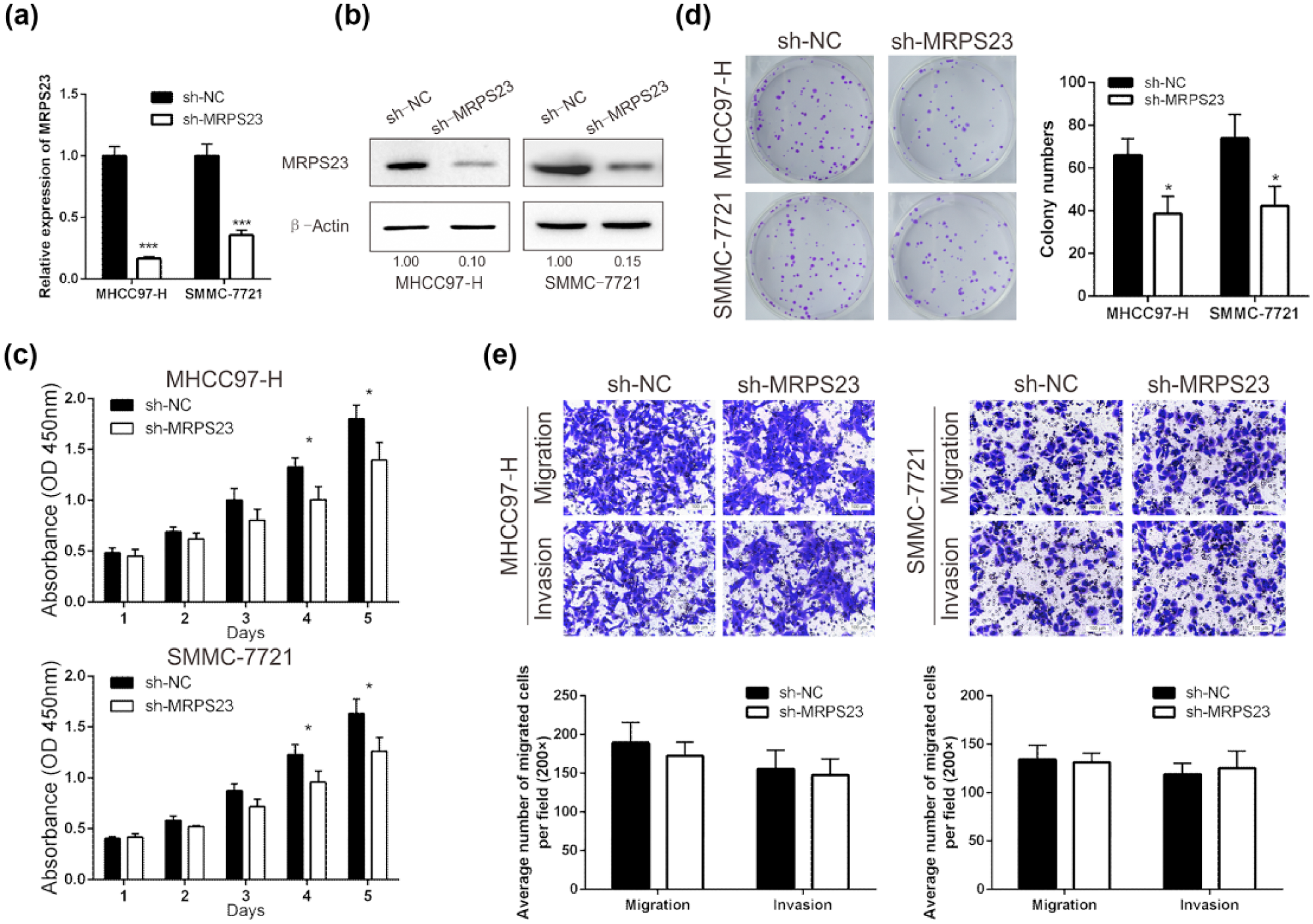
The effect of MPRS23 on HCC cell proliferation, colony formation, migration, and invasion in vitro. (a) RT-qPCR and (b) Western blotting analyses for the expression of MRPS23 after transfection with sh-MRPS23 or sh-NC in MHCC97-H and SMMC-7721. (c) Reduction of MPRS23 inhibited the growth rate of HCC cells detected by CCK-8 proliferation assay. (d) Colony-forming ability of HCC cells was attenuated after the decrease of MRPS23. (e) Migration and invasion assays of HCC cells were detected after transfection with sh-MRPS23 or sh-NC. Migrated and invaded cells were counted in five randomly selected areas under the microscopic field (200×; *p < 0.05; ***p < 0.001).
To determine the function of MRPS23 in the HCC cell lines, the CCK-8 assay was performed to assess the proliferation of the transfected HCC cells. Downregulation of MRPS23 inhibited HCC cell proliferation compared with that in the NC group (Figure 4(c)). Next, colony formation assays were performed to confirm the above result. Downregulation of MRPS23 led to a reduction in the colony counts compared with those in the control group (Figure 4(d)). However, the downregulation of MRPS23 had no significant changes on migration and invasion in vitro in either the MHCC97-H or SMMC-7721 cells (Figure 4(e)).
Downregulation of MRPS23 inhibits HCC cell tumor growth in vivo
We explored the effects of MRPS23 on the growth of HCC cells using tumorigenicity assays in nude mice. The experimental details were described in the Methods section. After 25 days, the weights of the tumors generated from the sh-MRPS23 group were significantly smaller than the weights of the tumors generated from the control group (Figure 5(a) and (b), p < 0.05). Meanwhile, Figure 5(c) showed the representative H&E staining of neoplasms. Figure 5(d) showed the representative IHC staining of MRPS23 in tumors. This result confirmed that MPRS23 affected the growth of HCC cells in vivo.

MRPS23 downregulated in MHCC97-H cells inhibited tumor growth in vivo. Transfected MHCC97-H cells were injected subcutaneously into nude mice (n = 5). After 25 days, the mice were euthanized and the tumors were excised. (a) The tumors of each group are shown. (b) The growth of tumor was measured by tumor weight. (c) Representative H&E staining of neoplasms. (d) Representative IHC staining of MRPS23 in tumors (**p < 0.01).
Discussion
As with many other cancers, the development of HCC is a multistep process involving the accumulation of genetic and epigenetic alterations.21,22 Typically, the activation of oncogenes or the inactivation of cancer suppressor genes is the key step in this process. 23 Therefore, identifying a series of molecular biomarkers may play an important role in the process of overcoming cancer, 3 especially the identification of driver oncogenes, and may promote a better understanding of cancer. 24 Surveillance using cancer biomarkers can enable early detection and has improved the prognosis of HCC in Japan. 25
There are some molecular indices which are associated with HCC proliferation. For example, RecQL1 is a key molecule in DNA repair and cell cycle replication; studies found that RecQL1 can be a biological marker to predict the malignancy and progression of HCC; meanwhile, silencing the expression of RecQL1 could prevent the growth of HCC. 26 PCBP2 is a member of PCBP family; overexpression of PCBP2 contributes to poor prognosis and plays an important role in HCC proliferation and depletion of PCBP2 inhibits the proliferation of HCC cell by influencing the level of cyclin-dependent kinase inhibitor p27. 27 Receptor for activated C kinase 1 (RACK1) gene is associated with apoptosis and proliferation of HCC; inhibition of RACK1 can suppress cell proliferation of HCC cells. 28 In addition, there are many other molecules such as Lamp2a, 29 pyruvate kinase 2 (PKM2), 30 and Fanconi anemia complementation group D2 (FANCD2) 31 that are involved in the proliferation of tumor cells.
Previous studies have found that the abnormal expression of MRPS23 has potential value in several cancer types,18–20 but its role in HCC has not been studied. The use of novel biomarkers to identify patients who are at higher risk of HCC and who have a poor prognosis is an important method to reduce mortality and medical costs, especially in the early stages of HCC. 32
In this study, we examined MRPS23 expression in HCC. First, we found that the MRPS23 mRNA and protein levels were significantly upregulated in the HCC samples compared with those in the ANLTs. Having determined that abnormal of MRPS23 expression was an important characteristic in HCC, we showed that high MRPS23 expression was correlated with the tumor size and TNM stage. We also found that patients with high MRPS23 expression levels had significantly poorer OS. The multivariate analysis found that the TNM stage, MRPS23 level, and metastasis were the main factors that correlated with the OS rates of HCC patients. These results confirmed that MRPS23 might serve as an independent prognostic factor for HCC.
Here, we evaluated the potential value of MRPS23 as a molecular marker for HCC that could reflect the prognosis of HCC patients. To validate our results, we combined the data available from the TCGA databases with the TMA results to generate a large number of varied samples for our analysis.
The first study of MRPS23 in cancer investigated its role in cervical carcinomas. The authors found several genes associated with lymph node involvement in cervical cancer; for instance, CKS2, MRPL11, MRPS23, LSM3, and PDK2 were upregulated in node-positive tumors. The authors also found that these genes were significantly related to survival. The overexpression of MRPS23 and other genes indicated high proliferation activity and oxygen consumption, which might suggest the underlying functions of MRPS23 in tumors. MRPS23 may play similar roles in HCC, and MRPS23 overexpression may be a powerful driver of abnormal tumor proliferation and metabolism. Therefore, MRPS23 is a promising target for tumor therapy. However, after downregulation of MRPS23, we did not find significant changes in tumor metastasis ability in vitro. In a study of MRPS23 in human breast cancer, the researchers found that MRPS23 was significantly upregulated in luminal subtype patients. 18 This conclusion was consistent with our findings in HCC, suggesting that MRPS23 might be a novel oncogene in several tumor subsets, especially in patients with highly proliferative cancer subtypes.
Importantly, we found that MRPS23 played a role in HCC cell proliferation. Tumor proliferation is the most important step in the tumorigenic cascade. 33 We investigated the effect of MRPS23 on HCC proliferation and found that MHCC97-H and SMMC-7721 cells treated with sh-MRPS23 displayed a decreased proliferation ability compared with the control group. Furthermore, we confirmed that MRPS23 affected the growth of HCC cells in vivo using nude mouse transplantation tumor experiments.
As for the potential mechanisms, MRPS23 expression was associated with proliferation, oxidative phosphorylation, and tumor size. 19 Tumor size is a strong prognostic factor for HCC. High proliferation rate may be the result of abnormal expression of MRPS23. High expression of MRPS23 may change the hypoxia state of tumor cell and affect the tumor proliferation and angiogenesis. MRPS23 as the structural components of mitochondria, upregulation of MRPS23 may increase mitochondrial activity, energy production through oxidative phosphorylation, and a high oxygen consumption rate. Meanwhile, high expression of MRPS23 may cause changes in the tumor microenvironment and promote tumor growth. The Warburg effect might be associated with mitochondrial damage in cancer cells, and abnormal expression of MRPS23 might regulate the metabolism of tumor cells and thus affect the proliferation of tumors
This study provides a better understanding of the malignant biological behaviors of HCC and identifies MRPS23 as a novel molecular marker for malignancy and prognosis in HCC. Knowledge of HCC can provide potential treatment strategies. However, there are many mechanisms by which MRPS23 may contribute to HCC that need to be fully elucidated, especially the association between abnormal mitochondrial function and the development of cancer,14,15,34 disorders in the expression of MRPS23 and mitochondrial function,16,35 and aberrant cancer cell metabolism, 36 tumor apoptosis,37,38 and autophagy. 39 Although we have performed basic studies investigating MRPS23 and HCC, there are many areas that need further exploration and validation. Additional studies are needed to determine the utility of MRPS23 and to optimize breakthroughs in HCC research.
Footnotes
Acknowledgements
The authors thank Kun Liu, Huiming Wang, and Siyang Liang for collecting the samples of HCC. The authors also thank Wei Liu, Yan Qiu, Xiaojuan Wang (The Xijing Hospital of The Fourth Military Medical University) for technical assistance. M.P., J.W., Q.H., and G.Z. contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work received support from the National Natural Science Foundation of China (No. 81272648, No. 81670593, and No. 81401940).
