Abstract
Objectives:
Patients with intermediate or advanced hepatocellular carcinoma (HCC) require repeated disease monitoring, prognosis assessment and treatment planning. In 2018, a novel machine learning methodology “survival path” (SP) was developed to facilitate dynamic prognosis prediction and treatment planning. One year after, a deep learning approach called Dynamic Deephit was developed. The performance of the two state-of-art models in dynamic prognostication have not been compared.
Methods:
We trained and tested the SP and Dynamic DeepHit models in a large cohort of 2511 HCC patients using time-series data. The time-series data were converted into data of time slices, with an interval of three months. The time-dependent c-index for OS at given prediction time (t = 1, 6, 12, 18 months) and evaluation time (∆t = 3, 6, 9, 12, 18, 24, 36, 48 months) were compared.
Results:
The comparison between SP model and Dynamic DeepHit-HCC model showed the latter had significant better performance at the time of initial admission. The time-dependent c-index of Dynamic DeepHit-HCC model gradually decreased with the extension of time (from 0.756 to 0.639 in the training set; from 0.787 to 0.661 in internal testing set; from 0.725 to 0.668 in multicenter testing set); while the time-dependent c-index of SP model displayed an increased trend (from 0.665 to 0.748 in the training set; from 0.608 to 0.743 in internal testing set; from 0.643 to 0.720 in multicenter testing set). When the prediction time comes to 6 months or later since initial treatment, the survival path model outperformed the dynamic DeepHit model at late evaluation times (∆t > 12 months).
Conclusions:
This research highlighted the unique strengths of both models. The SP model had advantage in long term prediction while the Dynamic DeepHit-HCC model had advantages in prediction at near time points. Fine selection of models is needed in dealing with different scenarios.
Introduction
Hepatocellular carcinoma (HCC) is the sixth most common cancer and the third leading cause of cancer-related death worldwide.1,2 Patients with intermediate stage HCC have poor prognosis and always require dynamic disease surveillance during comprehensive treatments, which consists of interventional therapies, targeted therapies and immunotherapy.3,4 The reported median survival time of patients with intermediate stage HCC ranges from 16.0 to 23.9.5,6
The 2022 update of BCLC strategy added two new concepts, the treatment stage migration (TSM) and untreatable progression, to facilitate dynamic adjustments of treatment plan when recommended treatment is not optimal. 7 In 2023, Alessandro Vitale proposed an evidence-based framework for treatment of HCC based on the novel concept of “multiparametric therapeutic hierarchy,” which allows dynamic adaption of the staging based algorithms. 8 Although considerable efforts had been made for optimizing and standardizing dynamic management of HCC patients, high-quality evidence is scarce to support refined adjustment of treatment plan. 9 Harnessing clinical big data to facilitate dynamic prognosis prediction and treatment planning for HCC patients represents a substantial challenge currently. 10
During the dynamic surveillance of patients with HCC, the time-series clinical data rapidly accumulate. 11 Modeling of these data may delineate the biological behaviors of HCC and help guide dynamic management. The classical autoregressive integrated moving average model (ARIMA) does not fit for survival data. 12 Temporal abstraction, 13 hidden Markov models, 14 and dynamic bayesian networks could be utilized for time-series survival data. 15 In recent years, a deep learning model called Dynamic Deephit based on RNN and attention mechanism was proposed, which theoretically can utilize a large fraction of repeated measurements and provide predictions with high consistency.16,17 Although these model have high accuracy in prediction, the nature of black box models make the results difficult to interpret.
In clinical practice, an ideal model should predict prognosis precisely while also being user-friendly and can provide guidance on treatment. 18 Our previous study in 2018 proposed a novel analytical approach called survival path, which converts the timeseries data into a cascading survival map, in which each survival path bifurcates at fixed time interval depending on selected prognostic features. 11 The model demonstrated to have higher superior or equal value than conventional staging systems in dynamic prognosis prediction for HCC patients at specific time interval. On the other hand, with the emergence of Dynamic DeepHit model in 2020, the value in dynamic prognostication between survival path model and Dynamic Deephit model for HCC patients haven’t been compared.
Therefore, in this study, we set out to compare the Survival Path model and Dynamic-Deephit model for dynamically prognosticating patients with HCC. Our study yielded significant findings on multiple aspects. Firstly, feature engineering was found playing a central role in enhancing SP model’s performance as well as its ability in generalization. Secondly, either the SP model or the Dynamic DeepHit model have its unique strengths in dynamic prognostication of patients with HCC. These advancements lay the groundwork for future research on developing novel machine learning tools for dynamic prognostication and management of HCC.
Methods
Study design and patient cohorts
Between January 2007 and Jan 2015, 10621 consecutive patients with newly diagnosed HCC at Sun Yat-sen University Cancer Center (SYSUCC) were retrospectively reviewed to develop the derivation (training) cohort. Between February 2015 and January 2016, an independent consecutive series of 2105 HCC patients treated at SYSUCC were reviewed to develop the internal validation cohort. Besides, between February 2016 and August 2018, 6055 patients from SYSUCC were reviewed to develop the internal testing cohort. The inclusion criteria were as follows: (1) clinically diagnosed with intermediate stage (BCLC stage B) HCC; (2) complete data of any of the following at initial diagnosis: computed tomography (CT) or magnetic resonance imaging (MRI) of the abdominal region, radiography or CT of the chest, routine bloodwork test, biochemical routine test, serum AFP level, and coagulation indices. The exclusion criteria were: (1) with history of other malignancies; (2) ECOG PS score >1 at initial diagnosis. The Hospital Ethics Committee of SYSUCC approved this study (B2023-639-01), which waived the need for written informed consent based on the retrospective nature of the study. A total of 1000, 200, and 879 patients were included in the derivation cohort, internal validation cohort, and testing cohort, respectively. Public multi-center database of 414 patients with intermediate stage HCC from three medical centers in southern China, which contains times-series clinical data on imaging and blood tests, was utilized as multicenter testing cohort (Figure 1).

Flowchart of study design.
The majority of HCC patients received transarterial chemoembolization (TACE) based integrated therapies as first-line treatment, which is decided based on the decision of the multidisciplinary teams, including hepatologists, radiologists, and interventional radiologists. The subsequent therapies after failure of first-line treatment included ablation, targeted therapies and palliative chemotherapy. Patients were advised to receive followed-up monthly during the period of initial treatment, subsequently at every 2-3 months for the first 2 years if complete remission was achieved. The frequency gradually decreased to every 3-6 months after 2 years’ remission.
The building of Dynamic DeepHit model was based on the derivation cohort and internal validation cohort. Survival path models were built using the derivation cohort as it does not need internal validation when setting up model.
Workflow of survival path mapping
The survival path models were built based on published R package, SurvivalPath. 19 The interval for time slices was set at 3 months and survival path model with nine time slices were computed. The minimum splitting sample size was 15 and the alpha value of significance is set at 0.05. Two survival path models were built based on the trainning dataset: the model of raw variables and the model of curated variables. The included variables in each survival path model were list in Supplemental Table S1. The key difference between the two models is that, in the model of curated variables, the empirical binary variables with known clinical and prognostic significance were added, including variables defined by both size and number of intrahepatic lesions, variables summarize both vascular invasion and extrahepatic metastatsis, and variable that describe the change of lesions. All the empirical variables can be computed based on variables that included in the model of raw variables.
Workflow of dynamic deephit model for HCC
The Dynamic Deephit model for HCC adopted the same architecture of the model described in previous work by Changhee Lee et al. 16 and is developed under the Python 3.7 platform. The deep learning model contained a shared subnetwork that process longitudinal measurements and predicts the next measurements of time-varying covariates, and a set of causespecific subnetworks which estimates the joint distribution of the first hitting time of death. Learning curves were utilized to optimize parameters of the model, including number of iteration and size of internal validation during model training. The parameters of the final model were described in Supplemental Table S2. All variables utilized in building the survival path model were included in the construction of Dynamic Deephit-HCC model. The predicted results for patients of given prediction time, denoted as t, and evaluation time, denoted as ∆t, were further utilized to calculated time-dependent c-index C(t, ∆t) to assess the model’s ability in dynamic prognostication.
Evaluation for prognostic significance
A total of five models were included and compared in dynamic prognostication for HCC patients, including survival path model using raw variables, survival path model with curated variables, Dynamic DeepHit-HCC model, CNLC staging system 20 and BCLC staging system. 7 The comparison of prognostic significance between survival path model and other models were conducted in training dataset and testing datasets, respectively.
The Harrell's c-indexes at different time slices were utilized to assess the value of prognostication between two survival path models. The time-dependent C-index can capture dynamic performance of the regression model over time and provide more precise assessment of models' discriminative capability than Harrell’s c-index. Given the unique feature of the Dynamic Deephit-HCC model that requires specific prediction and evaluation times for making predictions, time-dependent c-index C(t, ∆t) of different models/staging systems were further compared to assess the their ability in dynamic prognostication, where t indicates the prediction time which is the time when the prediction is made to incorporate dynamic predictions and ∆t denotes the evaluation time which is the time elapsed since the prediction is made.
Statistical analysis
Pearson χ 2 test was used to compare categorical variables between groups, respectively. To compare the efficacy in dynamic prognosis prediction different survival path models, the measurement of c-index in each time slice was computed. To further compare the efficacy in dynamic prognosis prediction between the survival path models, dynamic deephit model and staging systems, the measurements of time-dependent c-index of specific prediction time (t) and evaluation time (∆t) were computed using pec R package; means and standard deviations were obtained via ten random sampling of two thirds of cases. For category prediction models, subgroups less than three cases were omitted to reduce inference from extreme cases when computing c-index or time-dependent c-index. A random seed was set using the base R package. The comparison of c-index and time-dependent c-index between different models was conducted using Z test method. It is estimated that 288 times of tests were conducted during the comparison of time-dependent c-index between models. To avoid false positives caused by multiple tests, a meaningful alpha value is set at 0.0001 (0.05/288). All analyses were done using R 3.6.3 21
Results
The baseline characteristic of the patients
The training set consisted of 1000 HCC patients with a median age of 55 years (range, 14-85), the internal validation set consists of 200 HCC patients with a median age of 55 years (range, 20-82) and the testing set consisted of 897 HCC patients with a median age of 56 (range, 18-88) years (Table 1). The median follow-up time were 25.5, 22.3, and 24.6 months for the training set, internal validation set and internal testing set, respectively. Compared to the training set, the internal validation set and testing set had a lower proportion of patients with HBV infection. Besides, the proportion of patients with Child Pugh A class and those with multiple intrahepatic lesions (⩾4) in testing set were higher compared to the training set. Compared to the training set, the multicentric testing set had a higher proportion of patients with young age and female gender.
Baseline characteristics of training set, internal validation set, and testing sets at initial diagnosis.

Note. The distributions of baseline characteristics between training set and validation/testing sets are compared based on Chi-square test.
The survival paths built by training set with two strategies of feature selection
Although it’s widely accepted feature engineering is of vital importance for prognosis modeling, it’s impact in survival path mapping dealing with time-series data remains unknown. Two sets of variables, one consists of only raw variables and one consists of both raw and curated variables, were design to compute the survival path models. In the survival path model with curated variables, binary variables in terms of tumor diameter, 22 number of intrahepatic tumors, liver function, 23 AFP level,24,25 change of lesions after treatment and composite variable were designed, and added. 26
Compared to the eight variables in the SP model of raw variables, a total of twenty variables were put into the training of the SP model of curated variables (Table 2). The times-series data of HCC patients were firstly divided into data of time slices, with an interval of 3 months, and the two SP models were displayed in Figure 2. The included variables and their cutoff of each models were listed in Supplemental Table S1. In both training set and testing set, the c-index of the two models underwent rapidly rise from time slice no. 1 to time slice no. 3 and the trend become gentle in the following time slices. Except for time slice No. 2, SP model of curated variables have superior or non-inferior performance compared to the SP model of raw variables at all other time slices (Figure 3A). The peak c-index for both models were 0.841 and 0.786 in the training set, 0.761 and 0.722 in the internal testing set and 0.784 and 0.718 in the multicentric testing set, respectively.
The variables included in building the survival path models and Dynamic DeepHit-HCC model.
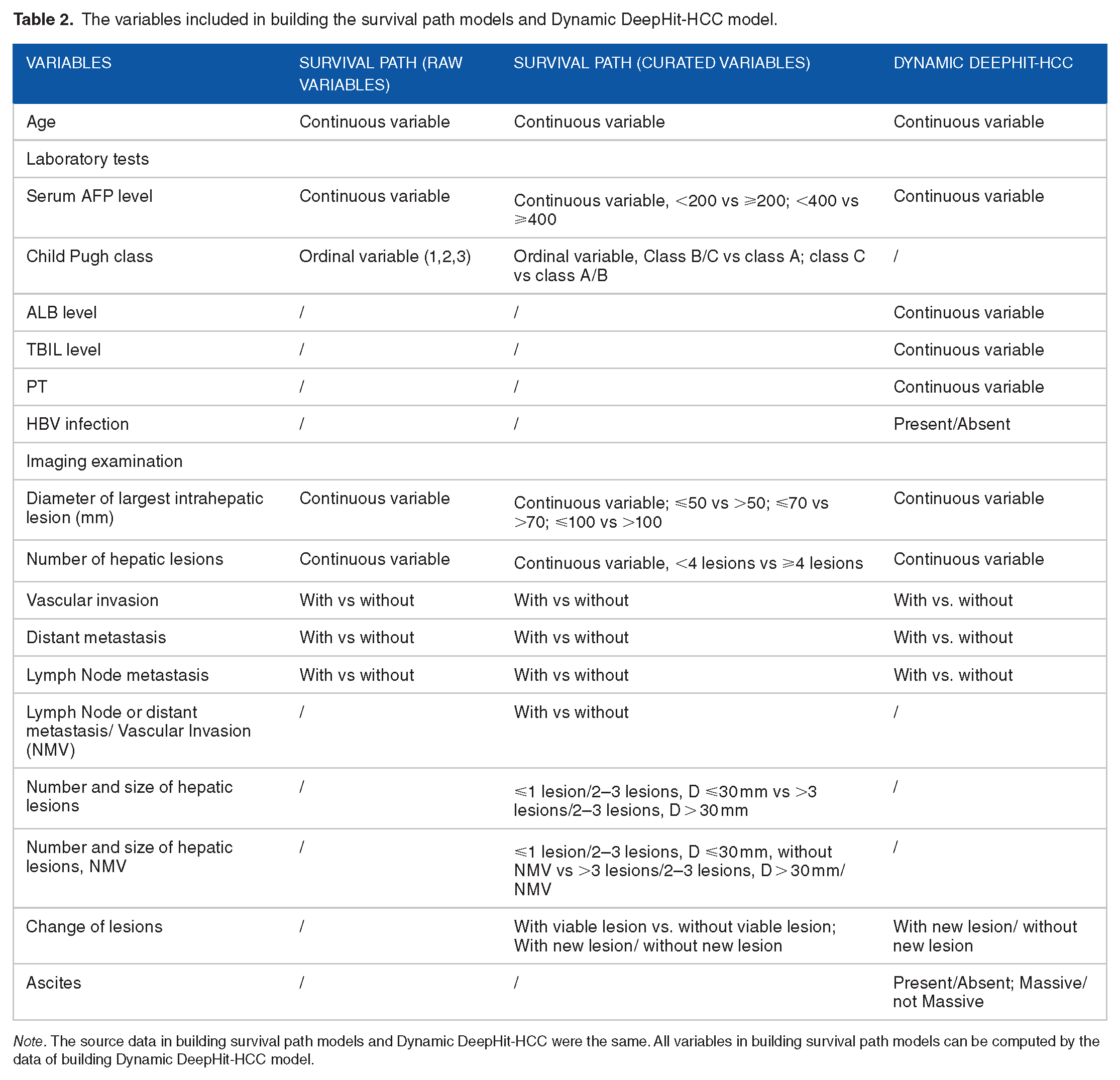
Note. The source data in building survival path models and Dynamic DeepHit-HCC were the same. All variables in building survival path models can be computed by the data of building Dynamic DeepHit-HCC model.
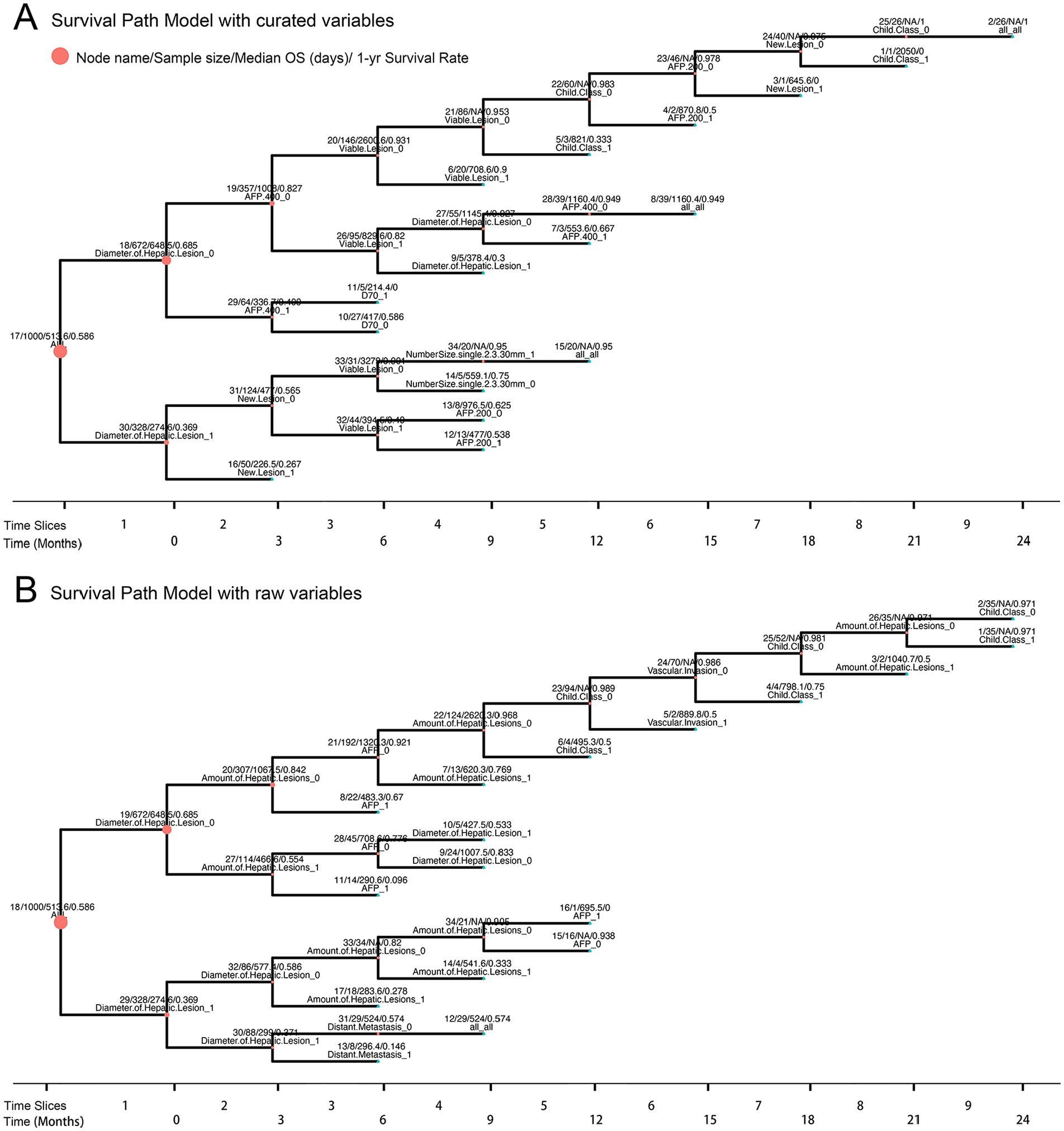
The survival path models constructed based on training set.

Comparison of C-index and time dependent c-index between different models: (A) The line chart represent change of c-index of survival path models along with time slices. The c-index of the two models underwent rapidly rise from time slice no. 1 to time slice no. 3 and the trend become gentle in the following time slices. Except for time slice No. 2, Survival path model built on curated variables have superior or non-inferior performance compared to the model built on raw variables at different time slices. The change of time dependent c-index along with different evaluation time for the two machine learning models in the training set (B), internal testing set (C) and multicentric testing set (D). ***P-value < .0001.
Comparison of the survival path model with Dynamic DeepHit-HCC model and conventional staging systems
The value of prognostic prediction between the survival path model, Dynamic Deephit-HCC model and conventional staging system were compared at different prediction times in both training and testing datasets (Table 3). It’s interesting to note that Dynamic DeepHit-HCC had significantly better performance in prognosis prediction compared to survival path model for patients initially admitted (prediction time at 1 month). However, the time-dependent c-index of Dynamic DeepHit-HCC model gradually decreased with the extension of evaluation time. By contrast, the fluctuation of time-dependent c-index of survival path model was relatively stable. When the prediction time is more than 6 months since onset of the treatment, the survival path model significantly outperforms the dynamic DeepHit model in long-term prognosis prediction (evaluation time > 12 months) (Figure 3B-D). The predictive ability of survival path models at different time points is superior or not inferior to traditional CNLC and BCLC staging systems (Supplemental Table S3). The feature importance of key variables at critical time points in Dynamic Deephit-HCC model were displayed in Table 4. The influential variables of Survival Path model at corresponding time slice were displayed in Table 5. It is notable that the variable “New lesion” appeared two times in survival path model by time slice no. 7 while in Dynamic Deephit-HCC model the importance of the variable rank 9th to 10th among the 15 variables included. Besides, the increased risk of “D. of Hep Lesions” become negative at distant prediction time (t = 18; ∆t = 18).
The top 10 most influential covariates of dynamic Deephit-HCC model at specific prediction time.
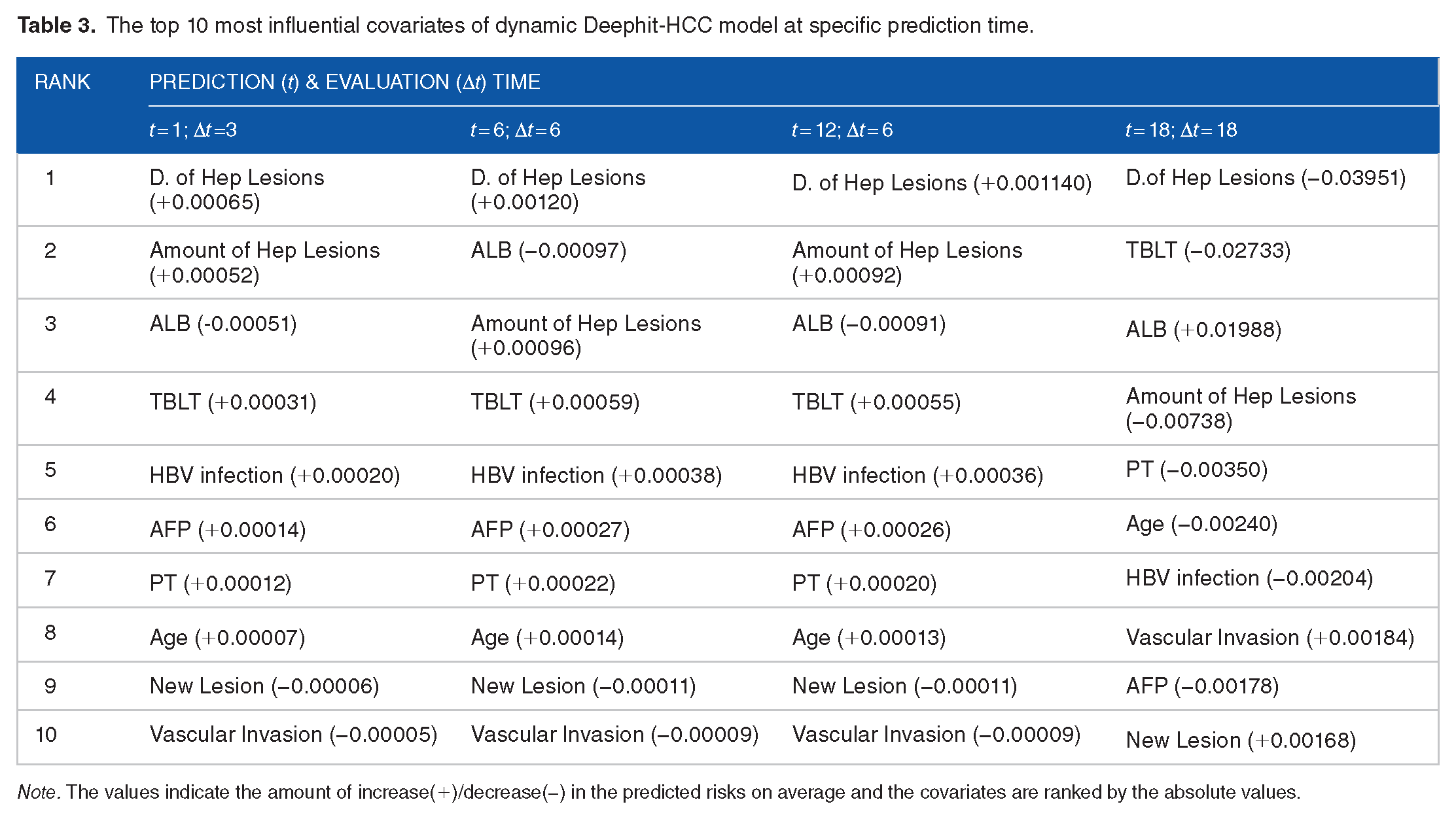
Note. The values indicate the amount of increase(+)/decrease(−) in the predicted risks on average and the covariates are ranked by the absolute values.
The influential variables of Survival Path model using curated variables at specific prediction.

Note. The key bifurcation variables were same across different evaluation time given specific prediction time.
Discussion
To our knowledge, this is the first study to compare the performance of two state-of-the-art machine learning approaches (Survival Path and Dynamic-Deephit) for dynamic prognosis prediction in HCC patients. Our study found that the Dynamic DeepHit-HCC model has higher predictive ability for patient at initial admission. However, when the prediction time is 6 months or even later since onset of treatment, the survival path model significantly outperforms the dynamic DeepHit model in long-term prognosis prediction. These results suggested that although deep learning models have significant advantages in high-throughput data and accuracy in prediction, the tree based models such as survival path models also have unique advantages in dynamic prediction based on time series data when key features identified. In addition, by contrast to the nature of black box model of deep learning, the survival path model has better visualization ability and is therefore may easier be accepted by oncologists.
In recent years, the application of traditional machine learning models in predicting prognosis, including support vector machines, Bayesian models, and linear regression, is gradually decreasing with the development of deep learning technologies. 27 Tree models still have certain advantages over deep learning models because of their nature of white box model and have no strict restriction on sample size. 28 Numerous studies have shown that the selection of key features in tree model is crucial for its effectiveness. 29 Although the original data used in the SP model with curated variables and SP with raw variables are the same, we have added variables with classical cutoff values in SP with curated variables, including the widely accepted serum AFP cutoff values and tumor diameter cutoff values. In addition, we have also added composite variables used in BCLC staging 30 and CNLC staging, 31 and variables for describing tumor changes based on time series. 32 In SP with curated variables, a total of 12 new variables were added, and we found that these variables can significantly enhance the model’s ability to dynamically predict prognosis and its generalization ability in the testing sets. Our result suggests that a well trained survival path model suitable for dynamic prognosis prediction of cancer requires identification and inclusion of key classification features.
To optimize the the Dynamic DeepHit-HCC model, learning curves were utilized for parameter setting the key model parameters. The Dynamic DeepHit-HCC model achieved good performance in both the training and testing sets. Compared to the Survival Path model, a peerless advantage of the Dynamic Deephit-HCC model is its ability to predict the survival probability of any evaluation time from any prediction time. In addition, at the initial admission of HCC patients, the Dynamic DeepHit-HCC model has significantly better predictive accuracy for prognosis than the Survival Path model and conventional staging systems. The reason for this phenomenon may be that Dynamic DeepHit, as a deep learning model, can directly integrate effective information of all variables for prediction since initial admission, 33 while survival paths bifurcated using only one key variable (in this case, the diameter of intrahepatic lesion) at time slice no. 1, with information in other variables unexploited. The disadvantage of Dynamic Deephit-HCC is that the model's long terms predictive ability (evaluation time > 12 months) gradually decreases. The reason for this trend may be because the majority of data in training the model are of early time slices, and hence the model has abundant raw data for short-term prediction, while in terms of long-term prediction, the fact of limited data may impede the performance of deep learning models. 34 In contrast, the bifurcation of the Survival Path model depends on the selection of key variables at at each time slices, and its long term prognostic ability is highly correlated with key factor identified by the data of distant time slice. Besides, the variables used in the survival path model are key prognostic factors that are fundamentally recognized in clinical practice, thereby minimizing the risk of overfitting. These two key differences in building models may explain the phenomenon that Survival Path model has better long term predictive ability for HCC patients. The differences in prognostic ability between the SP model and the Dynamic Deephit model-HCC suggested they can be applied in different scenarios to facilitate personalized prognosis prediction.
Compared to the Dynamic DeepHit model, one major drawback of the Survival Path model is that the requirement of sample size increase exponentially with the increase of time slices. If the sample size in specific node is not large enough to bifurcate when there do exist key variable that predict prognosis, the associated information will be lost. In this study, due to the restriction of sample size, we did not conduct further analysis after the ninth time slice. Developing node fusion technique to combine similar nodes may overcome this dilemma as the sample sizes of these node could be added up to support further bifurcation. In addition, the predictive ability of survival pathway model for patients at initial admission is relatively week, partly due to insufficient utilization of initial data and limited information carried by single categorical variables. Multi-classification of the nodes at early time slices may improve the performance of the survival path model in predicting prognosis. There still remains a lot of exploratory work to be done in improving the Survival Path model.
The interpretability of machine learning models is particularly important in the clinical medicine. 35 Although the survival path model could be visualized using tree diagram, its interpretation is still complex. In recent years, increasing studies have shown that large language models (LLM) can facilitate interpretation of machine learning models. 36 Establishing an online LLM platform to enable more convenient invocation and interpretation of survival path model might have a positive impact on the promotion and application of this methodology.
Our study has several limitations. First, it is a retrospective study using dataset of single center. Second, the time-series data after patients lost surveillance/follow-up could not be obtained and hence were unable to be included in the analysis, which may be a source of bias. Therefore, to demonstrate the value of this methodology in HCC, a large-scale multi-institutional prospective study for both modeling and validation is needed.
Conclusions
Dynamic DeepHit-HCC model had advantages over survival path model in dynamic prognostication of HCC patients at early prediction times. The survival path model outperforms the dynamic DeepHit model in long-term prognosis prediction (evaluation time > 12 months) at late prediction times (⩾6 months). The SP models, given its features of easy to visualize, use and understand, have the potential to enter the clinic in the near future.
Supplemental Material
sj-docx-1-cix-10.1177_11769351241289719 – Supplemental material for Machine Learning for Dynamic Prognostication of Patients With Hepatocellular Carcinoma Using Time-Series Data: Survival Path Versus Dynamic-DeepHit HCC Model
Supplemental material, sj-docx-1-cix-10.1177_11769351241289719 for Machine Learning for Dynamic Prognostication of Patients With Hepatocellular Carcinoma Using Time-Series Data: Survival Path Versus Dynamic-DeepHit HCC Model by Lujun Shen, Yiquan Jiang, Tao Zhang, Fei Cao, Liangru Ke, Chen Li, Gulijiayina Nuerhashi, Wang Li, Peihong Wu, Chaofeng Li, Qi Zeng and Weijun Fan in Cancer Informatics
Supplemental Material
sj-docx-2-cix-10.1177_11769351241289719 – Supplemental material for Machine Learning for Dynamic Prognostication of Patients With Hepatocellular Carcinoma Using Time-Series Data: Survival Path Versus Dynamic-DeepHit HCC Model
Supplemental material, sj-docx-2-cix-10.1177_11769351241289719 for Machine Learning for Dynamic Prognostication of Patients With Hepatocellular Carcinoma Using Time-Series Data: Survival Path Versus Dynamic-DeepHit HCC Model by Lujun Shen, Yiquan Jiang, Tao Zhang, Fei Cao, Liangru Ke, Chen Li, Gulijiayina Nuerhashi, Wang Li, Peihong Wu, Chaofeng Li, Qi Zeng and Weijun Fan in Cancer Informatics
Supplemental Material
sj-xlsx-3-cix-10.1177_11769351241289719 – Supplemental material for Machine Learning for Dynamic Prognostication of Patients With Hepatocellular Carcinoma Using Time-Series Data: Survival Path Versus Dynamic-DeepHit HCC Model
Supplemental material, sj-xlsx-3-cix-10.1177_11769351241289719 for Machine Learning for Dynamic Prognostication of Patients With Hepatocellular Carcinoma Using Time-Series Data: Survival Path Versus Dynamic-DeepHit HCC Model by Lujun Shen, Yiquan Jiang, Tao Zhang, Fei Cao, Liangru Ke, Chen Li, Gulijiayina Nuerhashi, Wang Li, Peihong Wu, Chaofeng Li, Qi Zeng and Weijun Fan in Cancer Informatics
Footnotes
Acknowledgements
We would like to thank Ms. Juan Nie for providing continuous encouragement to Dr. Lujun Shen in the past seven years in the pursuit of excellence in medicine. We would also like to thank Mr. Tongzhou Wang, Dr. Mengxuan Cui and Dr. Yao Wang for technical support in programming.
Authors’ Contributions
Lujun Shen and Weijun Fan and conceptually designed the research; YJ, TZ, FC and LS collected data and screened data; LS and YJ analyzed the data; LS, YJ, TZ, FC, LK, CL, GN, WL, PW, CL, QZ and WF validate the interpretation and drafted and revised the manuscript. All authors have read and approved the final manuscript.
Availability of data and materials
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Science and Technology Project of Guangzhou City (202201011375).
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
