Abstract
C-X-C chemokine receptor 4 and Notch1 have been shown to play oncogenic role individually. This study aimed to determine the combinatorial role of C-X-C chemokine receptor 4 and Notch1 in lung adenocarcinoma. Expression of C-X-C chemokine receptor 4 and Notch1 was detected in resected tumor samples from 185 patients with lung adenocarcinoma at stage I–IIIa by immunohistochemistry. Correlations of their immunoscores with clinicopathological characteristics and disease-specific survival were retrospectively investigated. A three-dimensional capillary-like sprouting model was established to assess the effects of C-X-C chemokine receptor 4 and Notch1 on angiogenesis in vitro. The results revealed that expression of C-X-C chemokine receptor 4 and Notch1 was elevated in lung adenocarcinoma tissues. The high co-expression of C-X-C chemokine receptor 4 and Notch1 was significantly correlated with tumor size, tumor status, nodal status, tumor stage, and lymphovascular invasion, as well as decreased disease-specific survival. Multivariate analysis showed that lymphovascular invasion (hazard ratio: 0.205, 95% confidence interval: 0.086–0.491, p < 0.0001) and co-expression of C-X-C chemokine receptor 4 and Notch1 (hazard ratio: 0.293, 95% confidence interval: 0.168–0.510, p < 0.0001) were independent indicators of poor prognosis in lung adenocarcinoma. Furthermore, Notch1 enhanced the effects of C-X-C chemokine receptor 4 to promote angiogenesis by regulating Flt1 and Flt4 in vitro. In conclusion, co-expression of C-X-C chemokine receptor 4 and Notch1 is associated with tumor progression and lymphovascular invasion and is an independent indicator of poor survival in lung adenocarcinoma. In lung adenocarcinoma patients with high C-X-C chemokine receptor 4 and Notch1 expression, simultaneous inhibition of both factors might be an effective treatment strategy.
Introduction
Lung cancer remains the most common incident cancer and the leading cause of cancer death in China. 1 Non–small cell lung cancer (NSCLC) makes up 85% of all lung carcinomas, and lung adenocarcinoma (LUAD) is the most common histological subtype. Although recent advances in diagnosis and therapy have improved outcomes for some LUAD patients, risks of distant metastasis via lymphovascular vessels still highlight systemic threats of this cancer. 2 Therefore, it is important to identify predictive markers of poor prognosis or lymphovascular involvement to tailor the treatment for high-risk patients.
Transmembrane C-X-C chemokine receptor 4 (CXCR4) and its ligand C-X-C chemokine 12 (CXCL12) play key roles in tumor-stroma crosstalk, including the regulation of tumor growth, metastasis, angiogenesis, and lymphangiogenesis.3–5 CXCR4 overexpression has been reported in more than 20 human solid tumors, and elevated CXCR4 expression is correlated with aggressive metastasis, advanced tumor stages, and poor survival in NSCLC patients.6–8
Notch signaling pathway is evolutionarily conserved and regulates cell differentiation, proliferation, and apoptosis in both physiological and pathological processes, including tumor angiogenesis and lymphangiogenesis.9–11 Binding of a Notch ligand (Dll1, Dll3, Dll4, Jagged1, and Jagged2) to a receptor (Notch1-4) triggers receptor cleavage and enables the intracellular domain of Notch (NICD) to translocate to the nucleus, affecting transcription of target genes such as Hes (hairy/enhancer of split) and Hey (Hes-related proteins) families. 9 Notably, ligands Jagged1 and Dll4 have opposing effects on angiogenesis. Jagged1 acts as a positive regulator of tip cell formation and sprouting to antagonize negative Dll4/Notch signaling. 12 Thus, Notch signaling is linked to be both oncogenic and tumor suppressive in tumorigenesis, depending on the tissue and cellular context.13–15 Notch signaling in NSCLC has also been investigated. Conflicting results was initially reported, but more recent studies support an oncogenic role for Notch1 in NSCLC.16–19
Additionally, interactions have been investigated between CXCR4 and Notch signaling. Wang et al. 20 first showed that Notch signaling regulated mobilization and homing of endothelial progenitor cells (EPC) by modulation of CXCR4 expression. Notch was also reported to promote tumor cell proliferation and migration in multiple myeloma and ovarian cancer through the upregulation of CXCR4/SDF-1 expression.21,22 However, the combinatorial expression of CXCR4 and Notch in human LUAD and the impact on prognosis have not been investigated.
In this study, we performed a systematic immunohistochemical (IHC) analysis of CXCR4 and Notch1 expression in human LUAD samples to evaluate the correlation with clinicopathological parameters and to determine the impact on prognosis. Furthermore, a three-dimensional (3D) capillary-like sprouting model was established to assess potential roles of CXCR4 and Notch1 in angiogenesis in vitro.
Patients and methods
Patients
Written informed consent was obtained from all patients, and this study was approved by the Ethics Committee of Jinling Hospital, Medical School of Nanjing University. Case records of 185 patients diagnosed with LUAD at pathologic stage I–IIIa were retrospectively reviewed, who underwent radical resection between January 2011 and December 2012 at Jinling Hospital based on the following criteria: (1) no preoperative adjuvant radiotherapy or chemotherapy; (2) no previous malignancy; (3) adequate organ function; and (4) available for follow-up. Histopathologic features of tumors were defined according to the 8th edition of American Joint Committee on Cancer (AJCC) staging system. Patients’ characteristics and clinical features were obtained from hospital records. Follow-up was carried out on an outpatient basis at 3-month intervals for the first 1 year and thereafter, at 6-month intervals. Disease-specific survival (DSS) was considered as the time from pulmonary resection to tumor-related death.
Immunohistochemistry and scoring
Fresh tumor specimens and their corresponding normal tissues were obtained during surgery. Formalin-fixed paraffin-embedded tissue blocks were used for IHC staining using streptavidin–biotin labeling method as previously described. 23 The following primary antibodies were used: anti-CXCR4 (1:500, Abcam, ab124824), anti-Notch1 (1:200, Abcam, ab27526), anti-CD31 (1:50, Abcam, ab28364), or anti-LYVE-1 (1:250, Abcam, ab14917). Phosphate-buffered saline (PBS) without the primary antibody served as negative control.
Two experienced pathologists blinded to this study scored the stained sections independently. Briefly, the intensity of staining was graded as 0 (no staining), 1 (weak), 2 (moderate), and 3 (strong). The percentage of positive immunoreactive cells was graded as 0 (<5%), 1 (5%–25%), 2 (26%–50%), 3 (51%–75%), and 4 (>75%). Multiplication of the two scores resulted in a final immunoreactivity score from 0 to 12 for each case. The immunoscores were further grouped into two categories: negative/low expression (scoring of 0–4) and positive/high expression (scoring of 6–12).
Cell culture
Primary human umbilical vein endothelial cells (HUVEC) isolated from human umbilical veins were cultured for 2–5 passages on plates coated with collagen I (Corning, USA) in EGM-2 medium (Lonza, Switzerland) as previously described. 24 D551 fibroblasts (ATCC, USA) and 293T cells (ATCC, USA) were maintained in Eagle’s Minimum Essential Medium (EMEM; Gibco, USA) and Iscove’s Modified Dulbecco’s Media (IMDM; Gibco), respectively, supplemented with 10% fetal bovine serum (Gibco) and 0.01% Pen-Strep (Invitrogen, USA) in 5% CO2 at 37°C.
Plasmid construction and lentiviral infection
Human CXCR4 short hairpin RNA (shRNA) and Notch1 shRNA plasmids in pLKO.1 lentiviral vector, as well as a control scrambled shRNA plasmid, were purchased from Sigma-Aldrich (USA). The shRNA sequences were as follows: CXCR4 shRNA (NM_003467, TRCN0000256866, 5′-CCGGTCCTGTCCTGCTATTGCATTACTCGAGTAATGCAATAGCAGGACAGGATTTTTG-3′) and Notch1 shRNA (NM_017617, TRCN0000003358, 5′-CCGGGATGCCAAATGCCTGCCAGAACTCGAGTTCTGGCAGGCATTTGGCATCTTTTT-3′).
A third-generation lentiviral gene delivery system was used for transfection, as previously reported. 25 Briefly, 293T packing cells were co-transfected via CaPO4 with the core lentiviral plasmids and either pLKO-CXCR4 shRNA, pLKO-Notch1 shRNA, pLKO-scrambled shRNA, or pCCL-RFP. Medium was replaced 20 h later with EGM-2 medium. About 48 h after the transfection, lentivirus-containing supernatants were collected, filtered, and added onto low-passage HUVECs.
Western blot analysis
Total cell lysates were isolated from HUVECs. Protein samples (30 µg) were separated by 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto polyvinylidene fluoride (PVDF) membranes. The membranes were blocked with 5% bovine serum albumin (BSA) and then incubated overnight at 4°C with primary antibodies: anti-CXCR4 (1:100, Abcam, ab124824), anti-Notch1 (1:1000, Abcam, ab52627), or anti-Tubulin (1:500, Abcam, ab6046). Bands were developed following incubation with horseradish peroxidase (HRP)-conjugated secondary antibodies.
Quantitative reverse transcription polymerase chain reaction
Total RNA was extracted from HUVECs using RNeasy Mini Kit (Qiagen, USA) and reverse transcribed into complementary DNA (cDNA) using a PrimeScript RT reagent kit (Takara, Japan). The cDNA was then used as the template for polymerase chain reaction (PCR) detection of the expression of target genes with PowerUp SYBR Green Master Mix (ThermoFisher, USA). Primer sequences are listed in Supplementary Table 1. Data were analyzed according to comparative threshold cycle value (2−ΔΔCt) method, normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH).
The 3D bead sprouting assay
The fibrin bead sprouting assay was carried out according to previous protocols. 26 HUVECs were first transfected with plasmid pCCL-RFP to stably express red fluorescent protein for better visibility. HUVECs were mixed with Cytodex3 beads (Sigma-Aldrich) at a ratio of 400 cells per bead in EGM-2 and shaken gently every 20 min for 4 h in 37°C incubator. The following day, HUVECs-coated beads were washed and resuspended in PBS containing 3 mg/mL fibrinogen (Sigma-Aldrich) and 0.15 U/mL aprotinin (Sigma-Aldrich), at a concentration of about 300 beads/mL. A total of 500 µL fibrinogen/beads solution was then mixed gently with 0.625 U/mL thrombin (Sigma-Aldrich) in 24-well plate. Afterward, the solution was incubated for 25 min to solidify and then covered with 50,000 D551 cells in 1 mL EGM-2, which settled to form a feeder layer on top of the fibrin clot. Medium was changed every other day, and images were taken with an Olympus IX70 microscope on day 5.
Quantification of vessels in vitro
All fibrin bead sprouting assays were replicated at least three times with similar results. Only sprouts greater than half the bead diameter in length were included and 50 beads were assessed for each group. Sprout number per bead, sprout length, and the proportion of lumenized sprouts were measured and analyzed with NIH ImageJ software.
Statistical analysis
All statistical analyses were performed using IBM SPSS Statistics (Version 21; IBM Corp., USA). Continuous data were expressed as the mean ± standard deviation and categorical data as percentages. Pearson chi-square and Fisher’s exact tests were used to analyze the associations between CXCR4 or Notch1 expression and clinicopathological variables. Spearman correlation analysis was used for correlation between groups. Survival analysis was performed with the Kaplan–Meier method, and the log-rank test was used for comparisons. In addition, Cox proportional hazards model was used for univariate and multivariate analyses. The value of p < 0.05 was considered statistically significant.
Results
CXCR4 and Notch1 expression in human LUAD tissues
Representative IHC staining patterns of CXCR4 and Notch1 are shown in Figure 1(a). In general, CXCR4 and Notch1 expression was absent or very weak in normal lung tissues. In contrast, CXCR4 staining was observed in the membrane and cytoplasm in LUAD tissues, and 99 (53.5%) samples displayed a high expression. Similarly, Notch1 staining was observed mainly in the membrane, and 84 (45.4%) samples showed a high expression in cancer tissues. Moreover, dual CXCR4/Notch1 positive expression was observed in 41 (22.2%) cancer tissues. No significant difference was found between the expression levels of CXCR4 and Notch1 (p = 0.0756) (Figure 1(b)). However, positive association between the expression of CXCR4 and Notch1 was noted statistically significant in the same cancer specimens (r = 0.457, p < 0.0001) (Figure 1(c)).
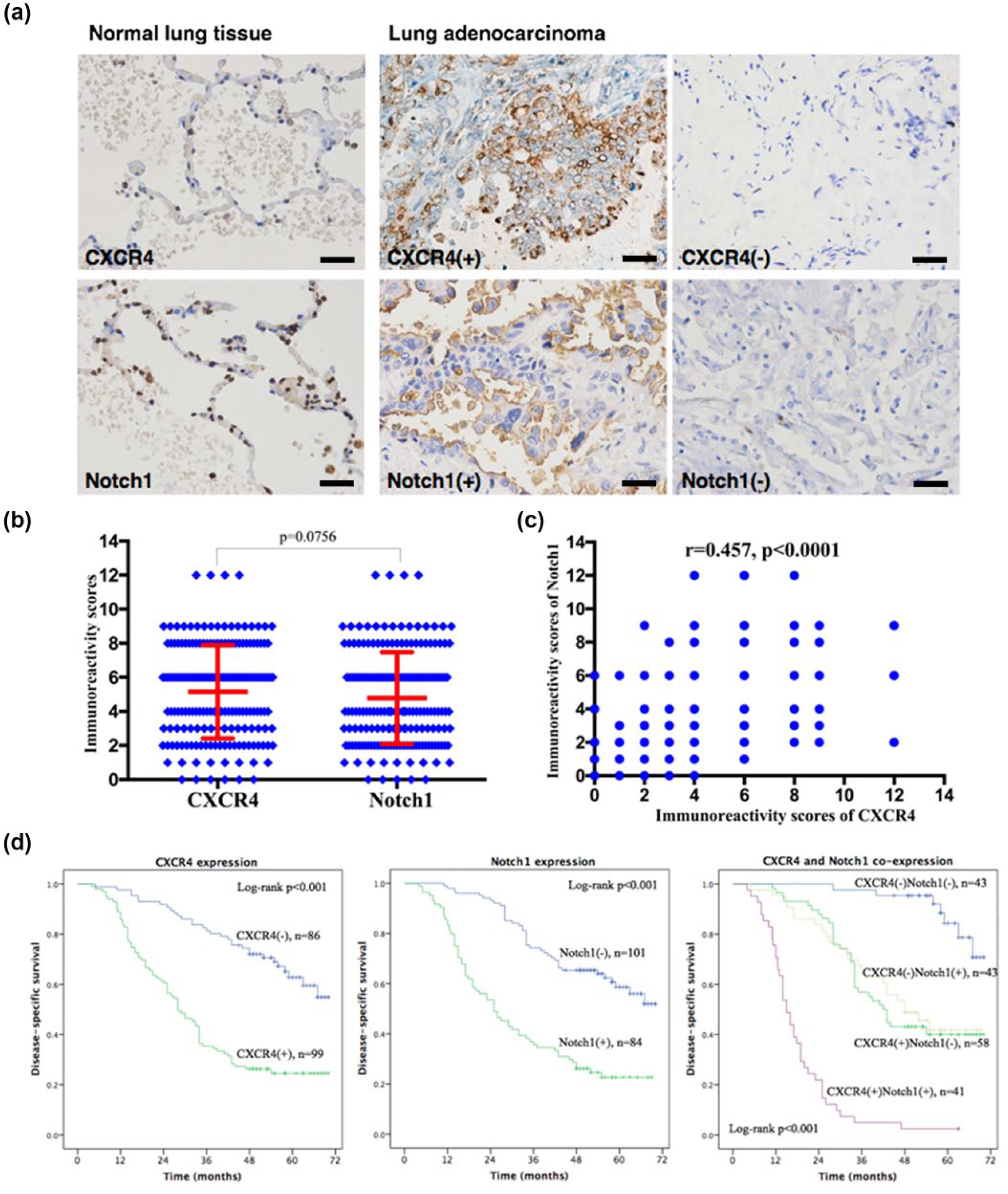
CXCR4 and Notch1 expression in normal and adenocarcinoma lung tissues. (a) Representative images of CXCR4 (upper panels) and Notch1 (lower panels) staining (scale bars represent 50 µm); (b) immunoreactivity scores of CXCR4 and Notch1 in all lung adenocarcinoma samples; (c) Spearman correlation analysis of CXCR4 and Notch1 immunoreactivity scores in lung adenocarcinoma samples; (d) Kaplan–Meier survival curves for disease-specific survival of lung adenocarcinoma patients according to the expression status of CXCR4 and Notch1.
Clinicopathological significance of CXCR4 and Notch1 expression
CXCR4 and Notch1 expression was correlated with clinicopathological characteristics of LUAD patients (Table 1). Neither CXCR4 nor Notch1 expression was associated with patients’ gender, age, or smoking history. Positive CXCR4 expression was significantly associated with tumor size (p = 0.012), tumor status (p = 0.001), nodal status (p < 0.001), AJCC tumor stage (p < 0.001), and lymphovascular invasion (p < 0.001), while high Notch1 expression was significantly associated with nodal status (p < 0.001), AJCC tumor stage (p < 0.001), and lymphovascular invasion (p < 0.001). In addition, high co-expression of CXCR4 and Notch1 was significantly associated with tumor size (p = 0.021), tumor status (p = 0.041), nodal status (p < 0.001), AJCC tumor stage (p < 0.001), and lymphovascular invasion (p < 0.0001).
Correlations between CXCR4, Notch1 expressions, and clinicopathological characteristics (n = 185).

CXCR4: C-X-C chemokine receptor 4; AJCC: American Joint Committee on Cancer.
Bold values indicate significant p values (<0.05).
CXCR4/Notch1 expression and patient survival
The medium follow-up was 46.5 months, and there were 105 tumor-related deaths. Kaplan–Meier survival curves showed that positive expression of either CXCR4 or Notch1 was significantly associated with decreased DSS in all LUAD patients (both p < 0.001) (Figure 1(d)). Furthermore, patients with co-expression of CXCR4 and Notch1 exhibited the worst prognosis (p < 0.001), while patients with negative expression for both proteins had the highest survival rate (Figure 1(d)).
Cox regression survival analysis
To determine independent risk factors of survival, Cox regression univariate and multivariate analyses were performed (Table 2). The univariate analysis revealed that tumor size, tumor status, nodal status, AJCC tumor stage, lymphovascular invasion, and co-expression of CXCR4 and Notch1 had significant prognostic value for DSS (p < 0.001). By multivariate analyses, tumor size (hazard ratio (HR): 0.383, 95% confidence interval (CI): 0.209–0.704, p = 0.002), lymphovascular invasion (HR: 0.205, 95% CI: 0.086–0.491, p < 0.0001), and co-expression of CXCR4 and Notch1 (HR: 0.293, 95% CI: 0.168–0.510, p < 0.0001) were identified as independent prognostic factors of DSS in LUAD patients.
Cox regression analyses of disease-specific survival in lung adenocarcinoma patients.

CXCR4: C-X-C chemokine receptor 4; AJCC: American Joint Committee on Cancer; HR: hazard ratio; CI: confidence interval; NS: not significant.
Bold values indicate significant p values (<0.05).
Altered CXCR4 and Notch1 expression affects capillary-like sprouting and lumen formation in vitro
Microvessel density (MVD) and lymphaticvessel density (LVD) were evaluated in human LUAD specimens by IHC staining using CD31 and LYVE-1, respectively (Figure 2). The MVD and LVD were observed to be significantly associated with CXCR4 and Notch1 expression. Thus, the 3D fibrin bead sprouting assay was performed to evaluate the impact of CXCR4 and Notch1 on angiogenesis in vitro (Figure 3(a)). Sprout number, sprout length, and proportion of visible lumenized sprouts were quantified (Figure 3(b)). CXCR4 knockdown HUVECs formed fewer sprouts (p = 0.003) and shorter sprout length (p = 0.008), with the lumen proportion marginally decreased (p = 0.035). In contrast, Notch1 knockdown HUVECs exhibited fewer sprouts (p = 0.006), shorter length (p = 0.016), and more lumen-forming defect (p < 0.001). Moreover, after both CXCR4 and Notch1 knockdown, HUVECs almost completely abrogated sprouting, with only few scattered vessels. Taken together, these data indicate that CXCR4 and Notch1 are essential to capillary-like sprouting and lumen formation in vitro.

Microvessel density (MVD) and lymphaticvessel density (LVD) in human lung adenocarcinoma specimens. (a) MVD and LVD were evaluated in normal and adenocarcinoma lung tissues by immunohistochemistry using CD31 (upper panels) and LYVE-1 (lower panels), respectively (scale bars represent 50 µm); (b) immunoreactivity scores of CD31 (MVD) in all lung adenocarcinoma samples according to the expression status of CXCR4 and Notch1; (c) immunoreactivity scores of LYVE-1 (LVD) in all lung adenocarcinoma samples according to the expression status of CXCR4 and Notch1.
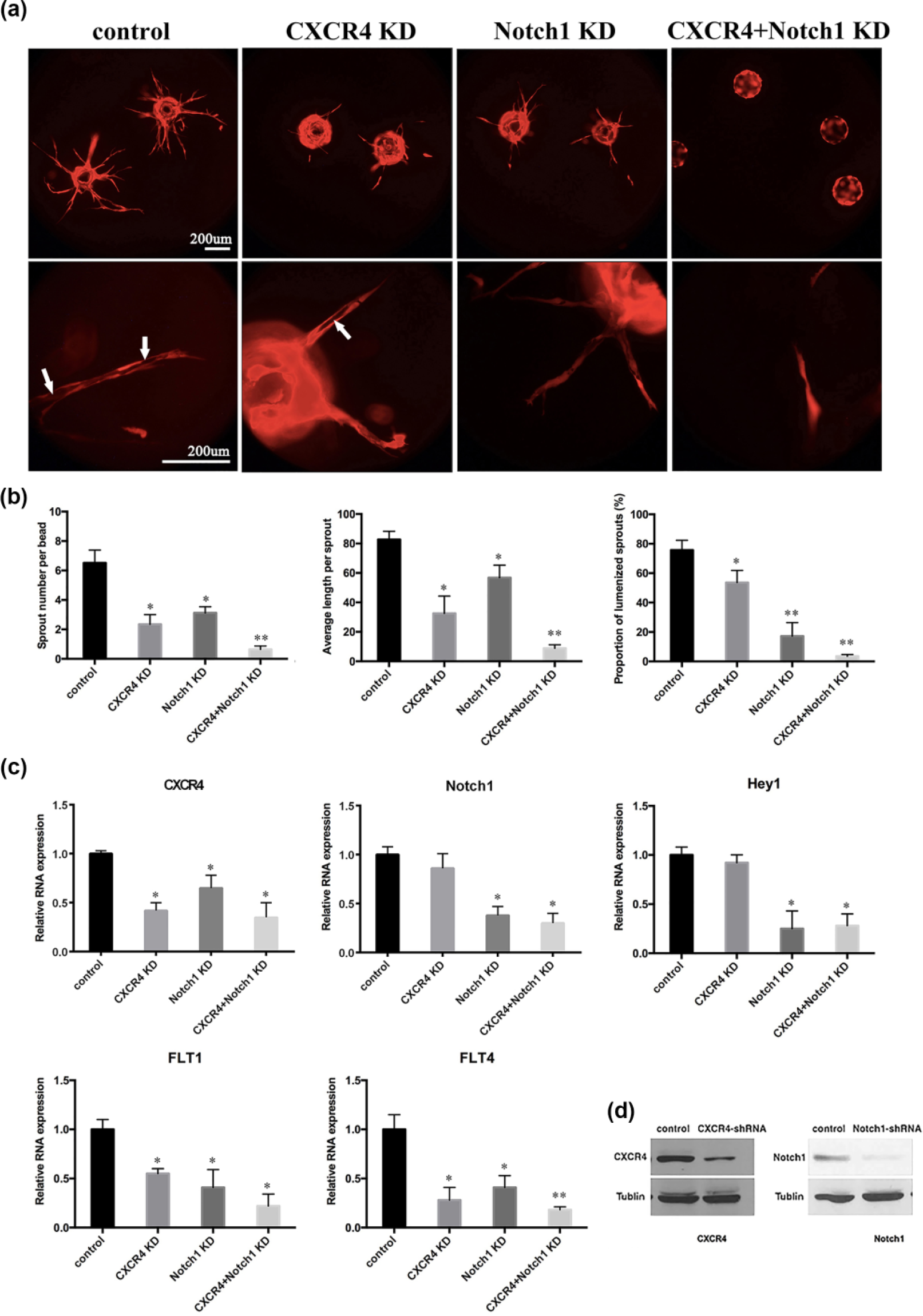
CXCR4 and Notch1 knockdown disrupt capillary-like sprouting and lumen formation in vitro. (a) Representative images of HUVEC (red) sprouting in fibrin bead sprouting assay, with individual or combined knockdown of CXCR4 and Notch1 (scale bars represent 200 µm). White arrows indicate lumen-containing sprouts. (b) Quantification of sprout number, sprout length, and proportion of lumenized sprouts. (c) Relative mRNA levels of CXCR4, Notch1, Hey1, Flt1, and Flt4 in HUVEC with individual or combined knockdown of CXCR4 and Notch1. (d) Validation of shRNA-mediated knockdown of CXCR4 and Notch1 in HUVEC by Western blot analysis.
Furthermore, mRNA levels of related genes were tested following CXCR4 and Notch1 interruption in HUVECs (Figure 3(c) and (d)). Generally, a positive modulation on CXCR4 by Notch1 was observed, while silencing CXCR4 showed no significant effect on Notch1 or its downstream target Hey1. Flt1 and Flt4 were significantly reduced in HUVECs with CXCR4 or Notch1 knockdown and further reduced in HUVECs with both CXCR4 and Notch1 knockdown.
Discussion
Lung cancer remains the leading cause of death in malignancy. Candidate biomarkers are highly required to serve as possible therapeutic targets and to predict outcomes in LUAD patients. In this study, we assessed expression patterns of CXCR4 and Notch1 in stage I–IIIa LUAD samples and evaluated their clinical significance.
CXCR4 is widely expressed in malignant tumors, and its oncogenic role has been confirmed.3–5 Multiple studies showed that CXCR4 could be detected by IHC in NSCLC tumors, and that the nuclear expression was associated with survival benefits while cytomembranous expression suggested poor outcomes.7,8,23,27 Moreover, tumors with CXCR4 overexpression were more likely to be diagnosed with adenocarcinoma cancer. 7 Consistent with these findings, our results showed a cytomembranous expression pattern of CXCR4 in LUAD samples, and aberrant CXCR4 expression was significantly correlated with tumor progression, lymphovascular invasion, and poor survival (Figure 1 and Table 1). Franco et al. 28 demonstrated that high cytomembranous expression of CXCR4 in NSCLC tumor cells significantly increased microvascular density and microvessel invasion by tumor cells, consistent with our data that high expression of CXCR4 in LUAD samples indicated enhanced microvessel and LVD (Figure 2).
Notch signaling is either oncogenic or tumor suppressive, depending on cellular responses and crosstalk with other signal-transduction pathways.13–15 Zheng et al. 16 showed that overexpression of active Notch1 inhibited the growth of human LUAD A549 cells through cell cycle arrest and interfered with tumor growth in nude mice. A recent study indicated that higher Notch1 mRNA expression predicted better overall survival in LUAD. 29 However, more experimental and clinical studies support an oncogenic role for Notch1 in NSCLC.17–19 In this study, our data support oncogenic roles of Notch1 in LUAD. Patients with positive Notch1 expression were more likely to have advanced stage, increased MVD and LVD, and poor outcomes. Additionally, through mining of TCGA data (LUAD, TCGA Provisional, 522 samples) by cBioPortal for Cancer Genomics (http://cbioportal.org/), overall survival analyses in LUAD were performed regarding mRNA levels of CXCR4 and Notch1 (Supplementary Figure 1). Patients with upregulated CXCR4 mRNA expression tended to have reduced overall survival, although not significantly (p = 0.239), while patients with upregulated Notch1 mRNA expression showed significantly worse overall survival (p = 0.0312). All these evidences support the oncogenic role for Notch1 in LUAD.
In addition, we found that both CXCR4 and Notch1 were frequently expressed in LUAD samples, and there was a positive correlation between their expression levels. The co-expression of CXCR4 and Notch1 significantly suggested advanced cancer stage and lymphovascular invasion in LUAD and was identified as an independent prognostic predictor for DSS. It was widely accepted that a systematic lymph node dissection during lobectomy was crucial for exact lung cancer staging. 30 Lack of appropriate number of dissected lymph nodes might cause false-negative nodal status, generating incorrect staging and treatment strategies. In this situation, co-expression of CXCR4 and Notch1 in tumors could predict lymphovascular invasion and poor prognosis and help tailor the staging and treatment for high-risk patients. Further prospective studies in larger scales of patients are needed to evaluate the clinical application of CXCR4 and Notch1 as predictors and biomarkers in LUAD.
Other than in tumor cells, CXCR4 and Notch1 are expressed in stromal cells, including endothelial cells. The role of CXCR4 and Notch1 in angiogenesis and lymphangiogenesis has been investigated individually.4,5,12 However, the interactions between CXCR4 and Notch1 in regulating angiogenesis have not been reported. In this study, we found that silencing CXCR4 or Notch1 significantly suppressed capillary-like sprouting and lumen formation of HUVECs by downregulating Flt1 and Flt4, which are involved in the regulation of angiogenesis and lymphangiogenesis. 31 Furthermore, knockdown of both CXCR4 and Notch1 made additive effects on angiogenesis in vitro. Previous studies have indicated similar positive interactions between CXCR4 and Notch1 in EPC cells, multiple myeloma, and ovarian cancer.20–22 Williams et al. 32 showed that activated Notch signaling by Dll4 downregulated CXCR4 in HUVEC, which supported our results indirectly, considering the negative Dll4/Notch effects. However, the synergistic functions of CXCR4 and Notch1 in promoting angiogenesis in vivo and in endothelial cells in LUAD tissues should be carefully evaluated in future studies. In addition, precise molecular mechanisms still remain to be clarified for better understanding of the interaction between Notch and CXCR4 during tumorigenesis and angiogenesis.
In conclusion, this study profiled the expression patterns and clinical significance of CXCR4 and Notch1 in human LUAD. Co-expression of CXCR4 and Notch1 is associated with tumor progression and lymphovascular invasion and is an independent indicator of worse survival of LUAD patients. In vitro sprouting assay revealed positive interaction between Notch1 and CXCR4 by modulating Flt1/Flt4 on angiogenesis. These findings suggest therapeutic potential of CXCR4 and Notch1. Simultaneous inhibition of both factors might be an effective treatment strategy in LUAD patients with high CXCR4 and Notch1 expression.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (No. 81172032) and Jiangsu Planned Projects for Postdoctoral Research Funds (No. 1601104B).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
