Abstract
IQ-domain GTPase-activating protein 1 is a scaffolding protein with multidomain which plays a role in modulating dishevelled (Dvl) nuclear translocation in canonical Wnt pathway. However, the biological function and mechanism of IQ-domain GTPase-activating protein 1 in invasive ductal carcinoma (IDC) remain unknown. In this study, we found that IQ-domain GTPase-activating protein 1 expression was elevated in invasive ductal carcinoma, which was positively correlated with tumor grade, lymphatic metastasis, and poor prognosis. Coexpression of IQ-domain GTPase-activating protein 1 and Dvl in the nucleus and cytoplasm of invasive ductal carcinoma was significantly correlated but not in the membrane. Postoperative survival in the patients with their coexpression in the nucleus and cytoplasm was obviously lower than that without coexpression. The positive expression rates of c-myc and cyclin D1 were significantly higher in the patients with nuclear coexpression of Dvl and IQ-domain GTPase-activating protein 1 than that with cytoplasmic coexpression, correlating with poor prognosis. IQ-domain GTPase-activating protein 1 significantly enhanced cell proliferation and invasion in invasive ductal carcinoma cell lines by interacting with Dvl in cytoplasm to promote Dvl nuclear translocation so as to upregulate the expression of c-myc and cyclin D1. Collectively, our data suggest that IQ-domain GTPase-activating protein 1 may promote the malignant phenotype of invasive ductal carcinoma via canonical Wnt signaling, and it could be used as a potential prognostic biomarker for breast cancer patients.
Introduction
Breast cancer is the most frequently developing cancer in case of women in many countries of the world.1–3 Invasive ductal carcinoma (IDC) accounts for 80% of all breast tumor, and it shows a worse survival rate than invasive lobular carcinoma. 4 This is a complex disease which is developed by genetic accumulation and epigenetic alterations. Preoperative chemoradiotherapy or hormonal therapy has increased the treatment efficiency of breast cancer patients. However, the anti-cancer therapy is not very effective especially in tumor metastasis patients. Tumor markers indicate the presence of disease and play an important role in patients’ treatment.5,6 Therefore, potential prognostic indicator of IDC requires further investigation.
IQ-domain GTPase-activating protein (IQGAP) family proteins are multidomain scaffolding proteins, containing three IQGAP isoforms: IQGAP1, IQGAP2, and IQGAP3. IQGAP1 is the most widely studied member of the IQGAP family. It is a key regulator of the cytoskeleton and participates in a number of biological processes, including cell adhesion, migration, and polarization.7,8 Previous reports indicate that IQGAP1 interacts with filamentous actin and stabilizes actin filaments through the calponin homology (CH) domain of its N-terminus. 9 Some protein factors interacting with IQGAP1, including extracellular signal–regulated kinase (ERK), Cdc42, Rac, and N-cadherin, modulate cognitive processes and spine density, showing obvious anomalous change in psychiatric disorders. 10 IQGAP1 is overexpressed in many tumors, including pancreatic cancer 11 and gastric cancer. 12 IQGAP1 overexpression was correlated with trastuzumab-induced resistance in breast cancer cell lines. 13 However, the expression of IQGAP1 in IDC has not been reported.
Dvl is recognized as a critical factor in Wnt pathway, which is an important component of biological response in tumor progression.14,15 There are many reports demonstrating the role of Dvl isoforms (Dvl-1, Dvl-2, and Dvl-3) in different tumors. Wnt-5a promotes the migration of breast cancer cell line via Dvl-2. 16 IQGAP1 regulates the nuclear translocation of Dvl in canonical Wnt signaling, 17 and the relationship between IQGAP1 and Dvl has been demonstrated in non–small cell lung cancer (NSCLC) tissues, 18 but their expression and mechanism had not been confirmed in breast cancer tissues and cell lines.
In this work, we tested the expression of IQGAP1 and Dvl in IDC and analyzed the correlation with clinicopathological parameters. In addition, we also sought to identify the involved mechanisms in breast cancer cell lines.
Materials and methods
Tissue samples of the patients
We collected 147 specimens from IDC patients who underwent complete cutting operation in the First Affiliated Hospital of China Medical University between 2005 and 2009. All resected tumor tissues of these patients were obtained in accordance with Human Subject Research Protocols approved by the ethics committees of the First Affiliated Hospital of China Medical University. Resected IDC tissues were obtained with written informed consent from adult patients. No patients had received preoperative chemotherapy or radiotherapy before operation. All the patients had complete follow-up data. The survival of the patient was defined as the time from the surgery day to the death day attributed to migration or recurrence or to the end of the follow-up period. Formalin-fixed paraffin-embedded sections of tumor tissues were stained with hematoxylin and eosin, and reviewed independently by three well-experienced pathologists, using the World Health Organization criteria (WHO Classification of Tumours of the Breast. 4th ed.). 24 Therefore, the grade of IDC was categorized into three groups: I (low grade), II (moderate grade), and III (high grade). All patients were women, and the age ranged from 32 to 81 years old (mean: 51 years). The clinicopathological factors including age, tumor size, lymph nodal status, grade, estrogen receptor (ER), and progesterone receptor (PR) status of the patients are listed in Table 1. We also collected 30 cases of adjacent normal tissue samples from the same patients as negative control.
IQGAP1 expression in relation to clinicopathological variables.

IQGAP1: IQ-domain GTPase-activating protein 1; ER: estrogen receptor; PR: progesterone receptor.
Immunohistochemistry
The steps of immunohistochemistry are the same as the previous. 18 Primary antibodies are listed as follows: IQGAP1 (1:50; sc-376021; Santa Cruz Biotechnology, Dallas, TX), Dvl (1:50; sc-7400; Santa Cruz Biotechnology), cyclin D1 (1:100; sc-20044; Santa Cruz Biotechnology), and c-myc (1:100; sc-40; Santa Cruz Biotechnology).
Evaluation of immunohistochemical staining
The immunohistochemical staining was independently evaluated by two pathologists, who were blinded to the clinicopathological information of the patient. The evaluation standard of immunohistochemical staining is the same as the previous. 18
Cell culture and transfection
The human normal breast cell line MCF-10A and human breast cancer cell lines MCF-7, BT-474, and MDA-MB-231 were purchased from American Type Culture Collection (Manassas, VA). These cell lines were cultured in RPMI-1640 or Dulbecco’s Modified Eagle’s Medium (DMEM) medium containing 10% fetal bovine serum (FBS) at 37°C with 5% CO2.
The expression of IQGAP1 in these cell lines was screened (Supplementary Figure 1). MCF-7 (with low level of IQGAP1) and MDA-MB-231 (with high level of IQGAP1) were selected for subsequent experiments. Lipofectamine 2000 (Invitrogen, Carlsbad, CA) was used to transfect the plasmid and siRNA into cells.
The pEGFP-IQGAP1 and pEGFP were provided by Prof. Kozo Kaibuchi. IQGAP1 small interfering RNA (siRNA; 5′-AACGCCGATCTAGTGTAACTGGCTGACTC-3′, 5′-ATACTGTATCACTAGTTCGGCCCCTGTCTC-3′) and the scrambled siRNA (5′-AAGTAGCAAGGACGCCATTGTCCAGTCTC-3′, 5′-AAACATTGCCGTCCTTGGTTCGCTCTCTC-3′) were designed and produced by the Shanghai GenePharma Co., Ltd (Shanghai, China).
Protein extraction
We prepared whole cell lysates with radioimmunoprecipitation assay (RIPA) lysis buffer. Nuclear and cytoplasmic extracts were prepared using Nuclear and Cytoplasmic Protein Extraction Kit (Beyotime Biotech, Beijing, China), according to the manufacturer’s instructions. We quantified protein concentrations using a bicinchoninic acid assay (BCA; Sigma, St. Louis, MO) and western blot analysis.
Western blot analysis
The protein was separated by 12% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE). After transferring to polyvinylidene fluoride (PVDF) membranes (Millipore, Billerica, MA), the membranes were blocked with 5% bovine serum albumin (BSA) in Tris-buffered saline (TBS) containing 0.05% Tween 20 before being incubated with primary antibodies overnight at 4°C. Primary antibodies are listed as follows: IQGAP1 (1:500; sc-376021; Santa Cruz Biotechnology), Dvl (1:500; sc-7400; Santa Cruz Biotechnology), cyclin D1 (1:400; sc-20044; Santa Cruz Biotechnology), c-myc (1:300; sc-40; Santa Cruz Biotechnology), LaminB1 (1:500; sc-30264; Santa Cruz Biotechnology), β-actin (1:3000; Sigma), and α-tubulin (1:500; sc-32293; Santa Cruz Biotechnology). After being incubated with corresponding secondary antibody (Santa Cruz Biotechnology), the signal was developed with enhanced chemiluminescence (ECL; Thermo Fisher Scientific, Waltham, MA). We used β-actin, α-tubulin, and LaminB1 as reference control.
Immunoprecipitation
We added sufficient amount of antibodies to total protein and gently rotated overnight at 4°C. Immunocomplexes were captured using protein A/G agarose beads (Beyotime Biotech, Beijing, China) and gently rotating for 3 h at 4°C. Then, we centrifuged the mixtures at 1500g for 5 min (4°C) and discarded the supernatant. Precipitates were washed three times with ice-cold RIPA buffer and boiled for 5 min to dissociate the immunocomplex from the beads. We collected the supernatant by centrifugation, and it was used for the western blot analysis.
Transwell assay
We used transwell membranes (8 µm pore polycarbonate membrane) in 24-well plates and applied Matrigel (BD Biosciences, New Jersey, USA) to the upper surface of the membrane in each well. Cells (treated with different factor, 3 × 105 cells in 100 µL serum-free medium) were added in the upper chamber, and the lower chamber was covered with the medium containing 10% FBS. After incubation for 20 h, the cells were fixed and stained with hematoxylin (Sigma). For each filter, we randomly selected 10 fields (400× magnification) under the microscope to count the number of invaded cells.
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide assay
Cells (treated with different factor) were plated on 96-well plates (3000 cells/well) and cultured in medium containing 10% FBS for 4 days. For evaluating the survival status of different cells, we added 20 µL of 5 mg/mL MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) solution to each well and incubated it (4 h at 37°C). Then, the media was discarded from each well, and the resultant MTT formazan was solubilized in dimethyl sulfoxide (DMSO; 150 µL). The results were evaluated with a spectrophotometer (490 nm) by a microplate reader (Bio-Rad Laboratories, Hercules, CA).
Colony formation assay
Cells (treated with different factor) were seeded on 6-cm dishes (1000 cells/dish) and cultured for 12 days. The plates were washed with phosphate-buffered saline (PBS) and stained with Giemsa. We counted the number of colonies (each with more than 50 cells) by QZD microscope (Olympus, Japan).
Statistical analysis
SPSS version 17.0 was used for all above analyses. The chi-square test was performed to calculate different correlations. Kaplan–Meier analysis was performed to estimate the IDC patients’ survival and the differences of survival time among the groups of patients. A two-tailed p < 0.05 was considered to be statistically significant in difference.
Results
IQGAP1 is highly expressed in IDC and is associated with poor prognosis in patients
We used immunohistochemistry to detect 147 IDC specimens and 30 normal breast tissues. IQGAP1 expression was positive in 121 cases of IDC (positive expression rate: 82.3%; 121/147), while duct epithelial cells from normal breast tissues showed weak or no staining (Figure 1(a)). In the positive group of IDC, there were cytoplasmic expression (49.0%; Table 1, Figure 1(b)), membranous expression (24.5%; Table 1, Figure 1(c)), and nuclear expression (8.8%; Table 1, Figure 1(d)). The analysis of the correlation between IQGAP1 expression and clinicopathological factors (Table 1) showed that IQGAP1 expression was positively correlated with tumor grade (p = 0.015) and lymphatic metastasis (p = 0.000), but it was not significantly associated with age, tumor size, or ER/PR status (p > 0.05).

Expression and localization of IQGAP1 in (a) normal duct epithelium and (b–d) IDC. (a) Negative IQGAP1 staining in the duct epithelial cells. Positive IQGAP1 staining in IDC was seen in (b) cytoplasm, (c) membrane, and (d) nucleus (original magnification: 400×; scale bar: 20 µm).
Postoperative survival time in IDC patients with IQGAP1 positive expression was significantly shorter than in those with negative expression (61.146 ± 2.748 vs 95.500 ± 2.451 months, p < 0.05; Figure 2(a)), and the survival time in patients with nuclear expression was significantly shorter than those with cytoplasmic or membranous expression (25.077 ± 3.477 vs 51.672 ± 2.726 vs 75.614 ± 3.576 months, p < 0.05; Figure 2(b)). Based on the above, IQGAP1 positive expression was associated with poor prognosis of IDC patients.

Kaplan–Meier curves showing overall survival among IDC patients with different expression and location of IQGAP1 and Dvl. The Kaplan–Meier curve shows that (a) IQGAP1 overexpression is associated with poor prognosis and (b) the group with nuclear positive expression was significantly lower than those with cytoplasmic and membranous positive expression. (c) The group without cytoplasmic and nuclear coexpression of IQGAP1 and Dvl (82 cases); the group with cytoplasmic and nuclear coexpression of IQGAP1 and Dvl (65 cases). (d) The group without cytoplasmic and nuclear coexpression of IQGAP1 and Dvl (82 cases); the group with cytoplasmic coexpression of IQGAP1 and Dvl (56 cases); the group with nuclear coexpression of IQGAP1 and Dvl (9 cases). (e) The group without cytoplasmic coexpression of IQGAP1 and Dvl (91 cases); the group with cytoplasmic coexpression of IQGAP1 and Dvl (56 cases). (f) The group without nuclear coexpression of IQGAP1 and Dvl (138 cases); the group with nuclear coexpression of IQGAP1 and Dvl (9 cases).
Coexpression of IQGAP1 and Dvl mainly in the nucleus and cytoplasm of IDC
IQGAP1 expression was significantly associated with Dvl in IDC (p = 0.000; Table 1). The positive expression rate of Dvl was 83.7% (123/147): 63.3% (93/147) for cytoplasmic expression, 12.2% (18/147) for membranous expression, and 8.2% (12/147) for nuclear expression. Cytoplasmic coexpression rate of IQGAP1 and Dvl was 77.8% (56/72; Table 2, Figure 3(a) and (b)) and 69.2% (9/13) for nuclear coexpression (Table 2, Figure 3(e) and (f)). Cytoplasmic coexpression was significantly correlated (p = 0.000, r = 0.295), also the nuclear coexpression (p = 0.000, r = 0.695); coexpression rate of IQGAP1 and Dvl in the membrane was only 16.7% (6/36; Table 2, Figure 3(c) and (d)), and membranous coexpression was not correlated (p > 0.05).
The relationship between IQGAP1 and Dvl expression in IDC.

IQGAP1: IQ-domain GTPase-activating protein 1; IDC: invasive ductal carcinoma.
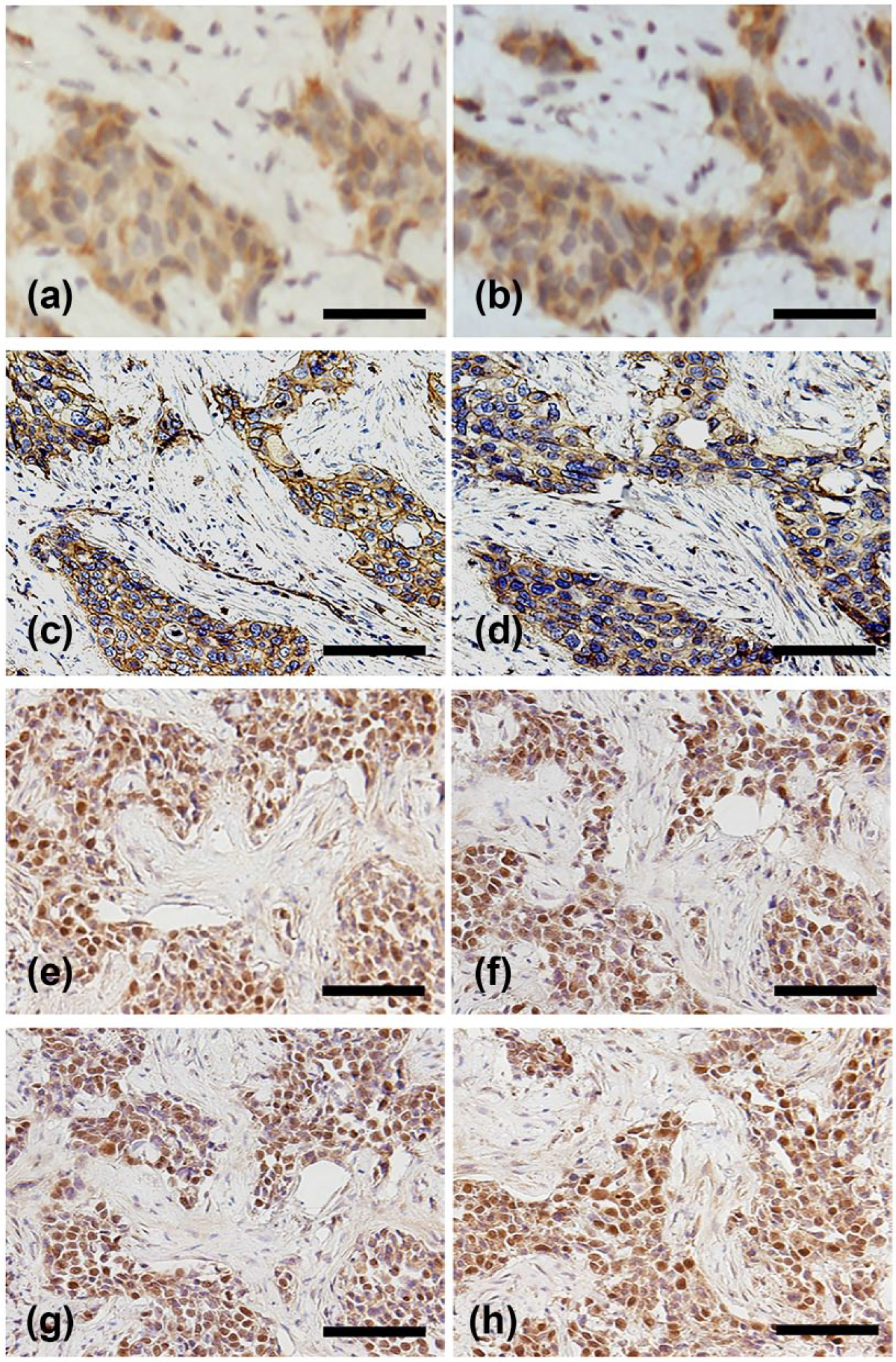
The expression and location of IQGAP1 and Dvl in IDC. Cytoplasmic coexpression of (a) IQGAP1 and (b) Dvl. Membranous coexpression of (c) IQGAP1 and (d) Dvl in IDC. Coexpression of (e) IQGAP1 and (f) Dvl in the nucleus of IDC. Positive expression of (g) c-myc and (h) cyclin D1 in IDC (original magnification: 400×; scale bar: 20 µm).
The expression of c-myc and cyclin D1 was obviously positive in the nuclear coexpression group of Dvl and IQGAP1 (Figure 3(g) and (h)), and their positive expression rates were 100% (9/9) and 88.9% (8/9), respectively. In the cytoplasmic coexpression group of Dvl and IQGAP1, the positive rates of c-myc and cyclin D1 were 53.6% (30/56) and 33.9% (19/56), respectively. The positive expression rates of c-myc and cyclin D1 in nuclear coexpression group were remarkably higher than cytoplasmic coexpression group (p < 0.05).
Coexpression of IQGAP1 and Dvl in the nucleus or cytoplasm was associated with poor prognosis of IDC
Kaplan–Meier analysis suggested a obviously lower survival time in IDC patients with cytoplasmic and nuclear coexpression of IQGAP1 and Dvl than those without coexpression (45.818 ± 2.979 vs 84.839 ± 2.981 months, p < 0.05; Figure 2(c)), and the survival time of nuclear coexpression group was significantly lower than that of cytoplasmic coexpression group (21.889 ± 3.498 vs 49.181 ± 3.099 months, p < 0.05; Figure 2(d)).
Postoperative survival time in the group with cytoplasmic coexpression of IQGAP1 and Dvl was obviously lower than that without coexpression (49.181 ± 3.099 vs 79.198 ± 3.335 months, p < 0.05; Figure 2(e)), as well as in the nuclear coexpression (21.889 ± 3.498 vs 75.154 ± 2.802 months, p < 0.05; Figure 2(f)). Therefore, cytoplasmic or nuclear coexpression of IQGAP1 and Dvl was significantly associated with poor prognosis of IDC.
IQGAP1 increases cell proliferation and invasion
To further confirm the effect of IQGAP1 on the biological behavior, breast cancer cell lines MCF-7 cells (low level of IQGAP1 expression) were transiently transfected with IQGAP1 and MDA-MB-231 cells (high level of IQGAP1 expression) were transfected with siRNA-IQGAP1 (Figure 4(a)); normal breast cell line MCF-10A cells were transiently transfected with IQGAP1 (Supplementary Figure 2(a)). The result showed that IQGAP1 had a significant effect on increasing cell proliferation, colony formation, and invasiveness (Figure 4(b)–(d), Supplementary Figure 2(b)–(d)).

IQGAP1 increases IDC cell invasion, proliferation, and colony formation. (a) Western blot analysis of IQGAP1 protein level in MCF-7 cells transfected with IQGAP1 cDNA (IQ) and MDA-MB-231 cells transfected with siRNA-IQGAP1 (si-IQ). Transfection of IQGAP1 cDNA (IQ) into MCF-7 cells increases (b) cell invasion, (c) cell proliferation, and (d) cell colony formation. Transfection of siRNA-IQGAP1 (si-IQ) into MDA-MB-231 cells decreases (b) cell invasion, (c) cell proliferation, and (d) cell colony formation. (b) The graph in shows the number of invading cells under different treatments. (d) The graph shows the number of colony formation under different treatments (*p < 0.05 compared with negative control (NC); error bars: SD).
IQGAP1 regulates the nuclear translocation of Dvl in canonical Wnt signaling
To further explain the mechanisms by which IQGAP1 modulated biological behavior of breast cancer cells, we detected the interaction between IQGAP1 and Dvl. The result confirmed that IQGAP1 interacted with Dvl in MDA-MB-231 and MCF-7 cells (Figure 5(a) and (b)). IQGAP1 could increase nuclear translocation of Dvl and upregulate the expression of c-myc and cyclin D1 (Figure 5(c)).

IQGAP1 interacts with Dvl so as to activate canonical Wnt signaling. Immunoprecipitation assay shows that IQGAP1 interacts with Dvl in (a) MDA-MB-231 and (b) MCF-7 cells. (c) The promotion efficiency of IQGAP1 transfection (IQ) to Dvl nuclear translocation and the expression of cyclin D1 and c-myc in MCF-7 cells. The inhibition efficiency of siRNA-IQGAP1 (si-IQ) to Dvl nuclear translocation and the expression of cyclin D1 and c-myc in MDA-MB-231 cells. α-tubulin was used as cytoplasm control. LaminB1 was used as nuclear control (NC: negative control; si-NC: scrambled siRNA for control).
Discussion
As a scaffolding protein, IQGAP1 could regulate the actin cytoskeleton and tumor progression.19,20 In a number of biological processes, scaffolding proteins influence cellular signaling by interacting with different partner proteins.21,22 Dvl is an important mediator in Wnt signaling, and its nuclear localization is necessary for canonical Wnt pathway. 23 IQGAP1 interacted with Dvl and modulated its nuclear translocation to promote the activation of canonical Wnt signaling in Xenopus embryos. 17 Our previous report discovered that cytoplasmic and nuclear coexpression of Dvl and IQGAP1 were associated with poor prognosis of NSCLC. 18
IQGAP1 had elevated expression in many kinds of human tumors.11,12 So far, the expression of IQGAP1 in IDC has not been reported. In this study, we found that IQGAP1 has high expression in IDC, mainly located in the cytoplasm, correlating with tumor grade and lymphatic metastasis. Cytoplasmic or nuclear coexpression rate of Dvl and IQGAP1 was significantly higher than membranous coexpression; cytoplasmic and nuclear coexpression of Dvl and IQGAP1 were significantly correlated but not the membranous coexpression. The coexpression of Dvl and IQGAP1 in the nucleus and cytoplasm was correlated with poor prognosis in IDC. The postoperative time in the patients with nuclear coexpression was remarkably worse than that with cytoplasmic coexpression, in concert with IDC cell study in vitro (immunoprecipitation and biological behavior assay). These results indicated that IQGAP1 might interact with Dvl in the cytoplasm of IDC cell, and their interaction might be an important step in regulating the nuclear translocation of Dvl but not in the membrane.
Above mentioned results provide evidence for a vital role of IQGAP1 in promoting malignant phenotype of IDC. We conclude that IQGAP1 may contribute to the progression of IDC, cytoplasmic and nuclear coexpression of Dvl and IQGAP1 are associated with poor prognosis of IDC. IQGAP1 may regulate Dvl nuclear translocation in the cytoplasm of IDC cell but not in the membrane. Nuclear coexpression of Dvl and IQGAP1 is correlated with the positive expression of c-myc and cyclin D1, which indicate that nuclear coexpression of Dvl and IQGAP1 is correlated with the activation of canonical Wnt signaling. To sum up, blocking the interaction between Dvl and IQGAP1 in the cytoplasm so that inhibiting Dvl nuclear translocation might be one of the effective indicators of breast cancer prognosis.
Footnotes
Acknowledgements
The authors thank Prof. Kozo Kaibuchi (Department of Cell Pharmacology, Graduate School of Medicine, Nagoya University, Japan) for providing plasmids, mentioned in this manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of China (No. 81602022 to H.-Y.Z. and No. 81301930 to L.-H.Y.) and the General Project of Education Department of Liaoning Province (Grant No. L2015595 to L.-H.Y.).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
