Abstract
Prostate cancer is the most common male malignancies in the United States. The specific characteristics of different disease stages have been deeply investigated. We present our data on ALDH1A3 as a potential therapeutic target for the prostate cancer based on several functional investigations. Also, we used The Cancer Genome Atlas datasets for primary prostate cancer to detect the relevance of ALDH1A3 and prostate cancer luminal phenotype. We found that ALDH1A3 correlated with androgen receptor signaling pathway in primary prostate cancer, which is consistent with its luminal layer localization. Then, from the genetic manipulation assay, we knocked out the ALDH1A3 in PC-3 cells and found significantly reduced proliferation rate as well as the invasion ability. Furthermore, we looked up our single center primary prostate cancer post-operative follow-up data and suggested that the high level ALDH1A3 expression could predict the poor progression-free survival in a 158-patient cohort. We concluded that ALDH1A3, localized in luminal layer in prostate epithelium, is highly expressed in prostate cancer. It played important role in maintaining the proliferation, invasion, and cell cycle. It can also become the potential biomarker in the future to guide the therapeutic manipulations for primary prostate cancer.
Introduction
In 2016, the American Cancer Society estimates the numbers of new cancer cases and deaths of urologic cancers in the United States to be 334,830 and 58,380, respectively. Prostate cancer alone is the most common malignancy and the second leading cause of cancer causing deaths in men. 1 Since 2005, the next-generation sequencing (NGS) technologies 2 have allowed us to better understand the molecular classification of cancer, which may provide benefits for clinical practice in oncology, such as diagnosis, prognosis, and treatment decisions. Going along with proteomics in the clinical and biological applications, 3 The Cancer Genome Atlas (TCGA) has performed a lot of sequencing analysis, including whole exome sequencing, transcriptome profiling, and DNA methylation analysis. In prostate cancer, the TCGA was presented with a comprehensive molecular analysis of 333 primary prostate cancer, 4 which revealed a molecular taxonomy with seven different subtypes, such as specific gene fusions (erythroblast transformation specific (ETS) family of transcription factors), or mutations (speckle-type POZ protein (SPOP), forkhead box protein A1 (FOXA1)). The transcriptome analysis also included the androgen receptor (AR) output or variants.
The aldehyde dehydrogenase family 1 member A3 (ALDH1A3) catalyzes the oxidation of retinal to the pleiotropic factor retinoic acid using nicotinamide adenine dinucleotide (NAD+). The level of ALDHs enzymatic activity has been regarded as a cancer stem cell (CSC) marker and seems to correlate with tumor aggressiveness. 5 ALDH1A3 is highly expressed in different cancers, such as pancreatic cancer,6,7 ovarian cancer, 8 and high-grade glioma. 9 In breast cancer, ALDH1A3 has been shown to be negatively correlated with estrogen receptor (ER) and progesterone receptor (PR) expression and positively correlated with the expression of human epidermal growth factor receptor 2 (HER2). 10 And ALDH1A3 is an independent prognostic biomarker in variety of cancers. High level of ALDH1A3 is associated with poor prognosis and lymph node metastasis in pancreatic cancer. 7 In breast cancer, ALDH1A3 only indicated with the prognosis with a subgroup of patients. 10 But little evidence has been shown the association of ALDH1A3 with prostate cancer. In this study, we correlated with the clinical and AR pathway relevance using TCGA datasets, and investigated the mechanisms of prostate cancer progression, and showed that the expression of ALDH1A3 is associated with prostate cancer progression-free survival in our single center prostate cancer follow-up data.
Methods and materials
Data analysis from public datasets
The human multi-organ RNA sequencing (RNA-seq) data were analyzed from GTEx Portal, version V6p (http://www.gtexportal.org/home/). TCGA data for ALDH1A3 expression in benign versus tumor were investigated from FireBrowse portal (http://firebrowse.org/). Immunohistochemical staining for ALDH1A3 in benign human prostate was downloaded from Protein Atlas Database (http://www.proteinatlas.org/). The AR pathway correlation analysis was analyzed by cBioPortal (http://www.cbioportal.org/) based on the TCGA primary prostate cancer study. The expression of ALDH1A3 in different cell lines was investigated from Broad-Novartis Cancer Cell Line Encyclopedia (CCLE) database (https://portals.broadinstitute.org/ccle/home/477/).
The establishment of ALDH1A3 knockout cell line
We designed the guide RNA for ALDH1A3 from (http://crispr.mit.edu/), targeting the first exon. The sequence of the guide is as follows—ALDH1A3: TAGTCTGCGGCGCACCGGCT; green fluorescent protein (GFP): GGCGAGGAGCTGTTCACCG. Then, we ligated the guide to the LentiCrispr-V2 system followed by Sanger sequencing validation. Finally, we produced the lentivirus according to the protocol previously described. 11 After 2 days of the infection to the PC-3 cell line, the puromycin selection was performed. We performed Western Blot assay to validate the knockout efficiency after 14 days of infection.
Patients and tissue microarrays
The protocol to generate the tissue microarrays (TMAs) in this study has been described previously in detail. 11 Briefly, a total of 158 patients with prostate cancer were included in this study. Those who have lost follow-up or benign tissue on the TMA were excluded. All patients in this study were treated by radical prostatectomy between 2008 and 2012 at the urology department of The First Affiliated Hospital of Nanjing Medical University (Nanjing, China). All the patients were recruited following informed consent, and the protocol was approved by ethical committee of The First Affiliated Hospital of Nanjing Medical University. In this study, the clinical and pathological features of all patients were summarized in Table 1. Progression was defined as biochemical recurrence (BCR) or metastasis. BCR was defined as two consecutive increases post-operative prostate-specific antigen (PSA) 0.2 ng/mL or greater in serum. For the staining score system, 12 the percentage of positive tumor cells was determined by at least five areas at 400× magnification and assigned to one of the following five categories—0: <5%, 1: 5%-25%, 2: 25%–50%, 3: 50%–75%, and 4: >75%. The intensity of immunostaining was scored as follows—1: low, 2: moderate, and 3: strong. The immunohistochemistry (IHC) score for ALDH1A3 on prostate cancer slides was: low expression: <8 and high expression: ≥8.
Characteristics of the 158 patients treated by radical prostatectomy.
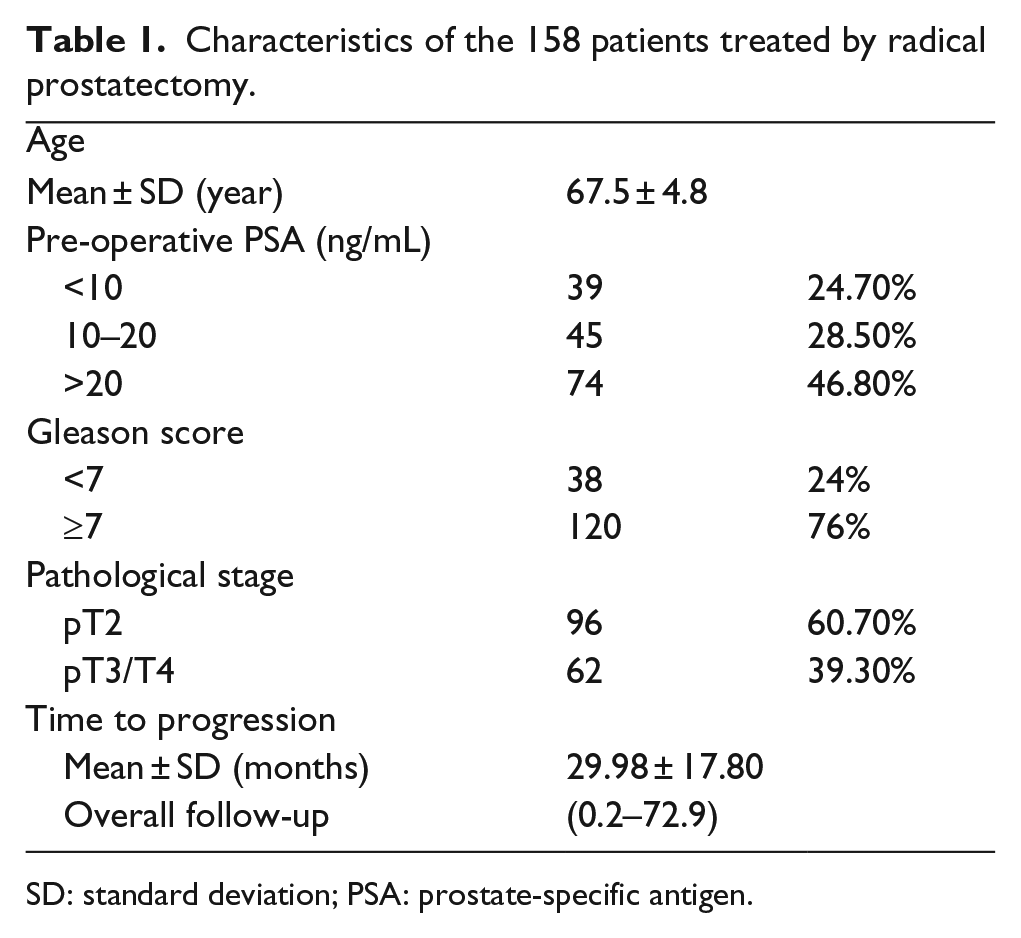
SD: standard deviation; PSA: prostate-specific antigen.
Western Blotting
The protein expression of ALDH1A3 by Western Blot assay was performed according to the protocol previously described. The antibodies against ALDH1A3 (Abcam, USA) and glyceraldehyde 3-phosphate dehydrogenase (GAPDH; Bioworld Technology, Inc., USA) were used in Western Blot assay in accordance with the manufacturer’s instructions.
Cell cycle analysis
The cell cycle distribution was performed by flow cytometry machine according to the cell cycle analysis protocol described previously. 11 After 14 days of transduction, cells from three different groups (parental, sg-GFP, and sg-ALDH1A3) were harvested followed with 70% ethanol fixation for 12 h in -20. The fixed cells were incubated in 50 mg/mL of propidium iodide and 1 mg/mL of RNase for 30 min at room temperature. The experiments were performed in triplicates.
Cell invasion assay
After 14 days of infection, we used invasion assay to examine whether ALDH1A3 null could decrease the metastasis on PC-3 cells. Each lower compartment of the transwell (BD Biosciences, USA) was filled with 500 µL of 0.5% fetal bovine serum (FBS) in medium to act as chemoattractant. A total of 5 × 104 cells in 200 µL of serum-free medium were placed in the upper chamber that was coated with Matrigel (BD Biosciences) in accordance with the manufacturer’s protocol.
Cell proliferation assay
After 14 days of infection, the cells were seeded onto 96-well plates at a density of 2000 cells/well and cultured for 24, 48, 72, and 96 h. Cell proliferation was determined using Cell Counting Kit-8 (CCK-8) assay (Beyotime Biotechnology, China) according to the manufacturer’s protocol. Absorbance was detected at the wavelength of 450 nm. Three wells were measured for cell viability in each treatment group.
Xenograft studies
The study was approved by Medical Laboratory Animal Welfare and Ethics Committee of Nanjing Medical University. BALB/c nude mice were randomly divided into two groups and each contained three mice. Cells (5 × 106 cells in 200 µL) were mixed with the same volume of Matrigel Matrix and injected subcutaneously into the left armpit. The volume of the xenografts was measured every week with calipers. The volume was calculated with the formula of length × width 2 × 0.52. The mice were humanely sacrificed 7 weeks after injection, and the xenografts were dissected. The methods were performed according to the approved guidelines.
Statistical analyses
Differences in vitro experiment between groups were subjected to Student’s t test. p < 0.05 was considered to be statistically significant. All the statistical calculations were performed using GraphPad Prism v 6.0 software (GraphPad Prism version 6.00 for Windows, GraphPad Software, La Jolla California USA, www.graphpad.com). The progression-free survival was assessed using the Kaplan–Meier curve and log-rank tests.
Results
ALDH1A3 is specifically expressed in human prostate
In order to determine the association of ALDH1A3 and prostate cancer, we first investigated the expression level in human normal prostate among the whole body organs. Surprisingly, ALDH1A3 is highly expressed in prostate and salivary glands according to the multi-organ RNA-seq data with GTEx Portal 13 (Figure 1(a)). Next, we aimed to investigate the ALDH1A3 expression level in the human prostate cancer. We analyzed TCGA data through FireBrowse portal (http://firebrowse.org/). Consistently, it expressed highest in human prostate cancer among pancreatic-cancer samples. The cancer samples are higher than the benign part (Figure 1(b)), which strongly indicated the relationship with tumor characteristics.

RNA-seq analysis for ALDH1A3 in multi-organ and pancreatic-cancer tissue expression. (a) From GTEx database, the ALDH1A3 is highly expressed in human prostate tissue and minor salivary gland, while the other organs had low expression. (b) From the TCGA data (accessed from FireBrowse portal), the expression of ALDH1A3 in prostate cancer samples is highest among all.
ALDH1A3 is expressed in luminal compartment and correlated with AR signaling pathway
First, with the help of Protein Atlas Database (http://www.proteinatlas.org/), we found that the ALDH1A3 is specially located in human prostate luminal epithelial cells not in the basal layer (Figure 2(a)). We speculate that the ALDH1A3 may have a relationship with AR signaling pathway. Thus, we searched the TCGA primary prostate cancer data 4 and correlated the ALDH1A3 messenger RNA (mRNA) level with AR pathway downstream genes. Thanks to the cBioPortal, 14 we found that several AR target genes, including KLK3, KLK2, TMPRESS2, FKBP5, ABCC4, PMEPA1, GNMT, ELL2, and HERC3, showed positive correlation with ALDH1A3 mRNA expression, conversely, the basal cell markers, TP63, KRT5, and NGFR, showed negative correlation (Figure 2(b)). Based on these data, we speculate that the ALDH1A3 may play an important role in the prostate luminal layer biological event and probably interacts with AR signaling pathway.

Correlation with luminal phenotype in prostate. (a) The Immunohistochemical staining for ALDH1A3 indicates the luminal layer localization (data from Protein Atlas Database; http://www.proteinatlas.org/). (b) The mRNA level of ALDH1A3 is positively correlated with luminal markers but negatively correlated with basal markers in prostate cancer according to the primary prostate cancer TCGA data (analyses from cBioPortal Pearson’s correlation tool).
ALDH1A3 knockout can deteriorate the prostate cancer proliferation and progression
In order to study the mechanisms of the ALDH1A3 in the prostate cancer proliferation and progression, we knocked out the ALDH1A3 in prostate cancer cells with Crispr systems. First, we checked the CCLE database and found that the ALDH1A3 is highly expressed in human prostate cancer cell line PC-3 (Figure S1). We targeted the first exon to knockout ALDH1A3 expression in PC-3 cells and validated with Western Blotting (Figure 3(a)). Next, we found that the proliferation rate of ALDH1A3 knockout cells decreased compared with sg-GFP control group (p < 0.01; Figure 3(b)). The invasion assay showed the similar results (Figure 3(c)). As a stem cell marker, ALDH1A3 may affect the cell cycle. We did the cycle assay to analyze the proportion of cells in G1 phase and found the knockout group significantly decreased compared with the control groups (Figure 3(d)). We did xenograft by subcutaneously injecting these groups of cells into nude mice. After 5 weeks, we found a significant difference between sg-GFP group and ALDH1A3 knockout group in tumor volume. All the animals were sacrificed by the end of week 7, the xenografts were removed from the body and were shown in Figure 3(e).

Functional analysis of ALDH1A3 knockout in prostate cancer cells. (a) Western Blotting validation for Crispr–ALDH1A3. The efficiency for Crispr is 70%–80% after puromycin selection. (b) CCK-8 analysis to determine the growth curve or proliferation rate showed that sg-ALDH1A3 group significantly decreased the proliferation (p < 0.01) after 14 days of infection. (c) Invasion assay after 14 days of infection showed that the knockout group had less invasive phenotype compared with the control group. (d) Cell cycle analysis indicated that the percentage of PC-3 knockout group cells in G1 stage was higher than that of the control group (p < 0.05). (e) In vivo xenograft showed the volume growth curve indicated the deteriorated proliferation when we target the ALDH1A3 by the end of the seventh week (p = 0.023).
ALDH1A3 can predict the progression-free survival in primary prostate cancer patients
We did TMA for 158 primary prostate cancer tissues and stained with anti-ALDH1A3 antibody to calculate the IHC score (all the scores were reviewed by pathologist M.B.). It has been clearly shown that ALDH1A3 is expressed in the cytoplasm of primary prostate cancer cells (Figure 4(a)). Furthermore, we cut off the IHC score to distinguish high-expression and low-expression group. We found that ALDH1A3 high-expression group showed poorer progression-free survival compared with ALDH1A3 low-expression group (Figure 4(b)) but not the overall survival (data not shown).

Tissue microarray data suggested the prognosis value in ALDH1A3. (a) Immunohistochemical staining for ALDH1A3, the positive cells are luminal layer when is consistent with the Protein Atlas data. (b) From our single center prostate cancer follow-up database, we found that the ALDH1A3 high-expression group has lower progression survival time.
Discussion
In the recent several years, with the advance on the high throughput sequencing techniques, more and more datasets have revealed the clinical implications of specific genes. More and more studies focus on looking for or validating the biomarkers to predict the prognosis or treatment response. Recently, poly (ADP-ribose) polymerase (PARP) inhibitor olaparib has received the breakthrough therapy designation by the Food and Drug Administration (FDA) in treatment of metastatic castration resistance prostate cancer (mCRPC) patients with DNA-repair-gene mutations. This designation is mainly derived from a phase II TOPARP-A trial data that a comprehensive analysis of targeted NGS, exome and transcriptome analysis, and polymerase chain reaction testing has successfully divided a subgroup of patients based on biomarkers of DNA-repair pathway whose overall response rate can reach 90%. 15 Another study using circulating tumor cell (CTC) number measured with the CellSearch assay is prognostic for survival in metastatic castration-resistant prostate cancer before and after therapy. In that study, the conclusion shows a 30% CTC decline after treatment from an initial count more than 5 cells/7.5 mL is independently associated with mCRPC overall survival following abiraterone and chemotherapy. 16 On our hands, ALDH1A3 has a relationship with progression-free survival after prostatectomy, which has significant clinical implications for patients with primary prostate cancer.
The theory that CSCs could be one of the mechanisms of treatment resistance has been widely accepted. As a stem cell marker, ALDH1A3 has been investigated into the relationship with chemotherapy resistance. 17 Several drug-resistant cell lines have high levels of ALDH1A3, including A375 in melanoma and 1295Lu-resistant treated with temozolomide, taxol, or doxorubicin 18 and HT29-taxol resistant colon cancer cells. 19 The study shows downregulation of ALDH1A3 expression in pleural malignant mesotheliomas increased sensitivity to pemetrexed and cisplatin. 7 More details have been discussed elsewhere. 20 In order to address the therapeutic significance of ALDH1A3, we knocked out this gene and found the functional relevance with the cell cycle, which could lead to decreased proliferation, invasion both in vitro and in vivo. Combining all these evidence, it can be stated that ALDH1A3 could be potentially a target in the future to overcome the chemoresistance via intervening cell cycle mechanisms. Further studies are needed to validate this.
Even with more approved and advanced androgen synthesis inhibitors and AR antagonists, prostate cancer will still develop resistance, and the most important mechanism to address this has been derived from AR signaling axis. Fortunately, many preclinical studies or clinical trials have focused on AR-interacting proteins such as chaperones,21,22 pioneer transcription factors,23,24 and AR transcriptional coregulators.25–27 Given the resistance from directly targeting AR, these novel AR-interacting proteins probably provide alternative therapeutic methods to overcome the resistance. Recent studies show that ALDH1A3 interacts with AR signaling pathway, 28 which is consistent with our hypothesis. We demonstrate that ALDH1A3 has a correlation with AR target genes, especially located in the luminal layer. Although downregulation of ALDH1A3 could deteriorate the AR signaling pathway is unknown, the relationship between AR and ALDH1A3 is still needed to be uncovered.
Footnotes
Acknowledgements
S.W., C.L., M.B., and X.L. contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Project Funded by the Priority Academic Program Development (PAPD) of Jiangsu Higher Education Institutions, by the Jiangsu Provincial Special Program of Medical Science (BL2012027), by the Program for Development of Innovative Research Team in The First Affiliated Hospital of Nanjing Medical University, by the National Natural Science Foundation of China (grant number: 81171963, 81201998, and 81201571), and by the Jiangsu Graduate Students Research and Innovation Program (KYLX15_0965).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
