Abstract
Certain viruses use microRNAs to regulate the expression of their own genes, host genes, or both. A number of microRNAs expressed by herpes simplex virus type 2 have been confirmed by previous studies. However, whether these microRNAs play a role in the metastasis of lung cancers and how these viral microRNAs precisely regulated the tumor biological process in lung cancer bone metastasis remain obscure. We recently identified the high expression of an acutely and latently expressed viral microRNA, Hsv2-miR-H9-5p, encoded by herpes simplex virus type 2 latency-associated transcript through microarray and quantitative polymerase chain reaction analyses which compared the expression of microRNAs in bone metastasis from lung cancer with primary lung cancers. We now reported that Hsv2-miR-H9-5p was highly expressed in bone metastasis and closely associated with pathological and metastatic processes of lung cancers. The functions of Hsv2-miR-H9-5p were determined by overexpression which results in an increase in survival, migration, and invasion of lung cancer cells in vitro. We determined that Hsv2-miR-H9-5p directly targeted SOCS2 mechanistically by dual-luciferase reporter assay. Then, we investigated the functions of SOCS2 in the progress of lung cancers. Reduction of SOCS2 dosage by hsv2-miR-H9-5p induced increased migration and invasion of lung cancer cells. Overexpression of SOCS2 inverted these phenotypes generated by hsv2-miR-H9-5p, indicating the potential roles of SOCS2 in Hsv2-miR-H9-5p-driven metastasis in lung cancers. The results highlighted that Hsv2-miR-H9-5p regulated and contributed to bone metastasis of lung cancers. We proposed that Hsv2-miR-H9-5p could be used as a potential target in lung cancer therapy.
Introduction
Lung cancer is the leading cause of cancer-related deaths worldwide, mainly attributing to the high frequency of metastasis. 1 Despite advancements and improvements in surgical and medical treatments, the survival rate of lung cancer patients remains frustratingly poor. Local control for early-stage non-small-cell lung cancer (NSCLC) has dramatically improved over the last decades for both operable and inoperable patients,2–4 but metastasis is still a major factor contributing to the high mortality of lung cancer. According to the World Health Organization (WHO), lung cancer will cause about 2.5 million deaths per year by the year 2030. 5 These facts highlight a need for better understanding of the cellular and molecular events underlying the genesis of this disease for the development of more effective therapeutics. 6 MicroRNAs (miRNAs) are a type of small regulatory RNAs, and dysregulation of miRNA expression is a feature of malignancies; 7 recent studies have shown that miRNAs are also involved in the pathological process of lung cancers, including proliferation, 8 apoptosis,9,10 cell cycle, migration, and invasion,2,11 such as miR-543 promotes proliferation and invasion by inhibiting PTEN 12 and microRNA-140-3p inhibits proliferation, migration, and invasion of lung cancer cells by targeting ATP6AP2 in lung cancer. 13 So far, some studies suggested an association between virus-encoded miRNA and cancer progression and metastasis, such as Epstein–Barr virus–encoded miRNA BART1 induces tumor metastasis by regulating PTEN-dependent pathways in nasopharyngeal carcinoma. 14 However, direct links between hsv2-miR-H9-5p and lung cancer metastasis remain undefined. Identifying such mediators and deciphering the underlying mechanism are important for better understanding the lung cancer progression and metastasis process, as well as finding potential therapeutic targets for effective lung cancer treatment in clinic.
Through microarray data analysis, we have previously identified several potential candidates associated with metastasis of human lung cancer. In this study, we demonstrated the expression pattern of miRNAs in patients with lung cancer and associated bone metastasis. We found that hsv2-miR-H9-5p, a human herpes simplex virus type 2 (HSV-2)-encoded miRNA, was highly expressed in patients with bone metastasis from lung cancer compared with control subjects. Herpes simplex virus type 1 (HSV-1) and HSV-2 are closely related human herpes viruses. 15 It is now known that all metazoan organisms encode miRNAs, and recently, miRNAs were identified in several DNA viruses, many of them in human herpesviruses.16,17 Of all the viruses studied to date, herpesviruses are the hands down winners in terms of the number of miRNAs encoded. 18
Therefore, the aims of this study were to determine the miRNA expression pattern of patients with lung cancer and to determine the association and potential relationships among hsv2-miR-H9-5p, lung cancer, and bone metastasis.
Materials and methods
Patients and tissue samples
Human lung cancer tissues were isolated from patients (n = 10) undergoing radical operation of lung cancer (5 men and 5 women; age range = 57–73 years with a median age of 67 years), and metastasized lung cancers consisted of cases with bone only as the control subjects (5 men and 5 women; age range = 54–74 years with a median age of 61 years). We obtained written informed consent from patients before sample analyses. Tissue samples were flash-frozen using liquid nitrogen immediately of surgical resection and stored at −80°C until analyses. The study was approved by the Human Ethics Committee of Changzheng Hospital of Second Military Medical University.
Microarray and data analysis
MiRNA expression profiling was analyzed using the miRCURY LNA Array (version 11.0) system. All the cancer tissues were shock-frozen in liquid nitrogen immediately after surgical resection in the operation room. Tissue specimens were homogenized in TRIzol reagent (Invitrogen), and total RNA was isolated according to the manufacturer’s protocol. The quantity and quality of RNA were measured by NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies), and RNA integrity was assessed to meet the requirement for microarray and quantitative reverse transcription polymerase chain reaction (qRT-PCR) analysis using standard denaturing agarose gel electrophoresis. Briefly, messenger RNA (mRNA) was purified from total RNA after removal of ribosomal RNA with an mRNA-Only Eukaryotic mRNA Isolation Kit (Epicentre Technologies). RNA samples were labeled with the Exiqon miRCURY Hy3/Hy5 Power Labeling Kit and hybridized on the miRCURY LNA Array (version 11.0) station. Scanning was performed with the Axon GenePix 4000B Microarray Scanner. GenePix pro version 6.0 was used to read image raw intensity. The intensity of the green signal was calculated after background subtraction, and replicated spots on the same slide were averaged to obtain median intensity. The median normalization method was used to acquire normalized data (foreground minus background divided by median). The median was the 50th percentile of miRNA intensity and was >50 in all samples after background correction. The threshold value for significance used to define upregulation or downregulation of miRNAs was a fold change >2, with a value of p < 0.05 calculated by the t-test. The miRNAs selected for investigation in our study were further filtered on the basis of expression levels and previously published data.
Real-time qRT-PCR
Total RNA was isolated from samples using TRIzol reagent. Isolated RNA was reverse-transcribed using miRcute miRNA First-Strand cDNA Synthesis Kit (TIANGEN), and quantitative PCR was performed with a LightCycler 480II (Roche) real-time PCR system. The Quantitative PCR conditions were as follows: 95°C for 30 s, followed by 40 cycles of 95°C for 5 s and 60°C for 30 s. A dissociation stage was added at the end of the amplification procedure. Melting curve analysis was performed to confirm the specificity of the PCR products. The primer sets specific for the miRNA amplification are as follows: for U6, 5′-ATTGGAACGATACAGAGAAGATT-3′ (forward) and for Hsv2-miR-H9-5p, 5′-CTCGGAGGTGGAGTCGCGGT-3′(forward). The relative expression of target gene SOCS2 (suppressor of cytokine signaling 2) was detected by qRT-PCR; the primer sets are as follows: 5′-GGAACGGCACTGTTCACCTTTA-3′ (forward) and 5′-AGCCTACAGAGATGCTGCAGAGA-3′ (reverse).
Cell culture
LTEP-α-2 and SPC-α-1 cell lines were obtained from Cell Bank of Type Culture Collection of Chinese Academy of Sciences, Shanghai Institute of Cell Biology, Chinese Academy of Sciences. Both cell lines obtained from the cell bank were passaged for fewer than 6 months. They were maintained in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum (FBS; Gibco) at 37°C and 5% CO2.
Reagents and vectors
Mature Hsv2-miR-H9-5P mimic and control RNA duplexes were obtained from GenePharma. The sequences of SOCS2-3′-UTR and its mutation were synthesized by Sangon Biotech and then cloned into the pmiR-GLO vector (Promega). Target gene SOCS2 was amplified by PCR, and products were electrophoresed on a 1% agarose gel and were visualized under ultraviolet (UV) light. The gel-purified PCR segment was cut with both EcoR I and BamH I restriction enzymes, and then, the segment was introduced into the pIRES2-EGFP vector (stored in our laboratory) to form the pIRES2-EGFP-SOCS2 plasmid. The accuracy of the fragment was identified by digestion with both EcoR I and BamH I and was verified as the correct clone through sequence analysis.
Dual-luciferase reporter assays
Dual-luciferase reporter assays were conducted to confirm that SOCS2 gene containing the predicted Hsv2-miR-H9-5p binding site was indeed target of Hsv2-miR-H9-5p. The human SOCS2 3′-UTR containing the predicted binding sites for Hsv2-miR-H9-5p was cloned into the pmiR-GLO dual-luciferase reporter plasmid (Promega) downstream of the firefly luciferase coding region as pmiR-GLO-SOCS2-WT (wild type, WT). Corresponding reporters containing mutations in the seed region of Hsv2-miR-H9-5P binding sites were generated through synthesizing DNA sequences containing described mutations (Sangon Biotech), annealing, and then cloning into the pmiR-GLO vector as pmiR-GLO-SOCS2-MUT (mutated type, MUT). HEK293T cells were co-transfected with the appropriate reporter plasmids and Hsv2-miR-H9-5P mimic in a 24-well plate. Lipofectamine 2000 (Invitrogen) was used as the transfection reagent according to the manufacturer’s instructions. Cells were lysed 24 h after transfection and analyzed for firefly and Renilla luciferase activities using the Dual-Luciferase Assays (Promega) on a Varioskan Flash Multimode Reader (Thermo Scientific).
Transwell cell migration assay and matrigel invasion assay
The ability of cell migration was evaluated using the Costar Transwell polycarbonate permeable supports. For the Transwell migration assay, a total of 105 cells resuspended in 100 µL of serum-free DMEM media were applied to the upper chamber of the device, and 600 µL of medium containing 10% FBS was added to the lower chamber. A polycarbonate membrane with a pore size of 8 µm was placed between the two chambers. Cells were allowed to incubate for 12–18 h at 37°C. The cells that had not yet migrated were removed from the upper surface of the membranes with cotton swabs. Adhered cells, attached to the lower surface of the membranes, were stained with 4′,6-diamidino-2-phenylindole (DAPI)–phosphate-buffered saline (PBS) to visualize the nuclei. Cell numbers in five predetermined fields in each replicate were counted under the microscope (OLYMPUS DP80). All assays were independently repeated at least for three times. Cell invasiveness was examined using a Matrigel basement membrane matrix invasion assay (Millipore) according to the manufacturer’s instructions. Cell invasion assays were performed as the migration assays except the Transwell membrane was precoated with Matrigel (Millipore) and the cells were incubated for 18–24 h.
Cell proliferation assay and apoptosis
For Cell Counting Kit-8 (CCK-8; Dojindo Molecular Technologies) assay, lung cancer cells (2 × 103 cells/well) were seeded into 96-well plates; 48 h after transfection with Hsv2-miR-H9-5P mimic and negative control using lipofectamine 2000 (Invitrogen), cells were incubated in CCK-8 solution for 2 h at 37°C. Cell proliferation was detected by CCK-8 assay at various time points according to the guidance of the manufacturer. The amount of formazan dye was measured by absorbance at 450 nm with a Varioskan Flash Multimode Reader (Thermo Scientific). Apoptosis was also assessed using an Annexin V–Fluorescein Isothiocyanate (FITC) Apoptosis Detection Kit (BD Biosciences) according to the manufacturer’s protocol, but for the cells that transiently transfected with pIRES2-EGFP plasmid, apoptosis was examined by flow cytometry after the cells were stained with 100 µL cell suspension containing 5 µL Annexin V-allophycocyanin (APC) at room temperature in the dark for 10–15 min.
Western blot analysis
For protein quantification, total cell extracts of LTEP-α-2 and SPC-α-1 which both transfected with Hsv2-miR-H9-5p mimic and negative control after 24 and 48 h were subjected to sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and electrotransferred to polyvinylidene difluoride membranes (Bio-Rad). After blocking in 20 mmol/L Tris-HCl, pH 7.6, containing 150 mmol/L NaCl, 0.1% Tween 20, and 5% (wt/vol) nonfat dry milk, the blots were incubated with the specific mouse anti-human SOCS2 (MAB4979; R&D) and anti–glyceraldehyde 3-phosphate dehydrogenase (GAPDH; sc-365062; Santa Cruz Biotechnology, Inc.) overnight at 4°C. The blots were then incubated with horseradish peroxidase–conjugated secondary antibody (1:2000; Jackson ImmunoResearch Laboratories) for 2 h while shaking at room temperature. The immunoreactivity was visualized by an enhanced chemiluminescence detection agents (agent A:agent B = 1:1) according to the manufacturer’s instructions. The densitometry plots were analyzed using ImageJ software.
Statistical analysis
For statistical analyses, mean values with standard deviation (SD) are shown in most graphs that were generated from several repeats of biological experiments. The p values were obtained from t-tests with paired or unpaired samples, with significance set at p < 0.05.
Results
MiRNAs are differentially expressed in bone metastasis of lung cancer versus primary lung cancer
The microarray cohort of subjects included 10 patients with bone metastasis of lung cancer and 10 patients of primary lung cancer without known metastasis. To determine the differential specific candidate miRNA involved in the pathological process of lung cancer metastatic to bone, we screened miRNA expression of cohorts mentioned above using the miRCURY LNA Array (version 11.0) system. The levels of miRNAs differed profoundly between the two cohorts, as illustrated in the heat map shown in Figure 1(a). A series of miRNAs was aberrantly expressed in bone metastasis of lung cancer compared to primary lung cancer. In consideration of the fold changes and p value of Hsv2-miR-H9-5p, we selected it as the aim of our research. We further confirmed the differential expression of Hsv2-miR-H9-5p by real-time PCR (Figure 1(b)). In addition to the microarray cohorts, we studied a second group composed of 10 patients with bone metastasis of lung cancer and 24 control subjects for independent validation. The real-time PCR results matched the microarray information (Figure 1(c)). To confirm the accuracy of the data, we selected hsa-miR-211-3p (Figure 1(d) and (e)) and hsa-miR-9-3p (Figure 1(f) and (g)) as control. These data indicated that the significantly altered expression of Hsv2-miR-H9-5p might be involved in the pathogenesis or phenotypic behavior of lung cancer metastasis.

Heat map of microRNA (miRNA) microarray expression and verification of the differential expression of Hsv2-miR-H9-5p. (a) Heat map of microRNA (miRNA) microarray expression data from tissue samples of patients with lung cancer (n = 10) and patients with bone metastasis of lung cancer as control subjects (n = 10). The expression of miRNA is hierarchically clustered on the y-axis, and primary lung cancer samples or control tissue samples are hierarchically clustered on the x-axis. The legend on the right indicates the miRNA represented in the corresponding row. The relative miRNA expression is depicted according to the color scale shown on the top. Red indicates upregulation and blue indicates downregulation. Numbers with C present bone metastasis of lung cancer samples; numbers with B, primary lung cancer samples. (b) Validation of miRNA microarray data by quantitative reverse transcription polymerase chain reaction. The microarray cohort included 10 lung cancer patients and 10 lung cancer control subjects. (c) Independent validation of differential expression of Hsv2-miR-H9-5p in patients. Quantitative reverse transcription polymerase chain in an independent validation set of 24 lung cancer patients and 10 bone metastasis control subjects. (d) Validation of relative hsa-miR-211-3p expression by quantitative reverse transcription polymerase chain reaction. The microarray cohort included 10 lung cancer patients and 10 lung cancer control subjects. (e) Independent validation of differential expression of hsa-miR-211-3p in patients. Quantitative reverse transcription polymerase chain in an independent validation set of 24 lung cancer patients and 10 bone metastasis control subjects. (f) Validation of relative hsa-miR-9-3p expression by quantitative reverse transcription polymerase chain reaction. The microarray cohort included 10 lung cancer patients and 10 lung cancer control subjects (**p < 0.01). (g) Independent validation of differential expression of hsa-miR-9-3p in patients. Quantitative reverse transcription polymerase chain in an independent validation set of 24 lung cancer patients and 10 bone metastasis control subjects (***p < 0.005, the relative expression of hsv2-miR-H9-5p was normalized to expression of U6). Gene expression was determined by the ΔCT method; data in (b)–(g) are shown as box plots. Each box represents the 25th–75th percentiles. Lines inside the boxes represent the median. Lines outside the boxes represent the minimum and the maximum.
Hsv2-miR-H9-5P promotes lung cancer cell invasion and metastasis in vitro
In view of bone metastasis of lung cancer with high expression of Hsv2-miR-H9-5P which implied that it may promote the metastasis of lung cancer, to test and verify the speculation, we upregulated the expression of Hsv2-miR-H9-5P in two Hsv2-negative lung cancer cell lines (LTEP-α-2 and SPC-α-1) using Hsv2-miR-H9-5P mimic and its negative control. Transwell migration assays (Figure 2(a)) confirmed that upregulation of Hsv2-miR-H9-5P dramatically increased the cell migration of either LTEP-α-2 or SPC-α-1 compared to relative negative control. Matrigel invasion assay (Figure 2(b)) revealed that the upregulation of Hsv2-miR-H9-5p significantly increased the cell invasion of both LTEP-α-2 and SPC-α-1 compared with the negative control. Together, these results suggested that Hsv2-miR-H9-5p propelled the migration and invasion of lung cancer cells in vitro.

Hsv2-miR-h9-5p propels cell migration and invasion of lung cancer cells in vitro. (a) Transwell migration assays. Hsv2-miR-h9-5p increased the cell migration of both LTEP-α-2 and SPC-α-1 compared with negative control (NC; *p < 0.05). (b) Matrigel invasion assays. Hsv2-miR-h9-5p increased the cell invasion of both LTEP-α-2 and SPC-α-1 compared with NC (*p < 0.05).
Hsv2-miR-H9-5p promoted cell proliferation and induced apoptosis resistance in lung cancer cells
Hsv2-miR-H9-5P was encoded by latency-associated transcript (LAT) sequences, which is a single viral gene—the LAT gene is abundantly transcribed during latency. In view of anti-apoptosis of LAT and the evolutionary conservatism, we speculated that Hsv2-miR-H9-5p may perform similar effect. 19 To investigate the biological effect of Hsv2-miR-H9-5p upregulation on lung cancer progression, the LTEP-α-2 and SPC-α-1 cell lines were used to transfect with Hsv2-miR-H9-5P mimic or negative control. The qRT-PCR was performed to confirm Hsv2-miR-H9-5P expression of LTEP-α-2 and SPC-α-1 cells after 48 h transfection. CCK-8 assay showed that Hsv2-miR-H9-5P upregulation significantly increased the rate of cell proliferation (Figure 3(a)). Flow cytometry was used to assess apoptosis in lung cancer cells transfected with Hsv2-miR-H9-5p mimic or negative control. Significant differences in Annexin-V-positive apoptotic cells were observed in the Hsv2-miR-H9-5p mimic-treated group in comparison to cells transfected with negative control. Hsv2-miR-H9-5p mimic and negative control induced apoptosis in 7.89% + 0.24% and 14.78% + 0.35% of LTEP-α-2 cells, respectively (p < 0.05, Figure 2(b)). In SPC-α-1 cells, they were 7.95% + 0.1% and 13.8% + 0.28%, respectively (p < 0.05, Figure 3(b)); in terms of cell morphology, cells transfected with mimic exerted strong resistance to cisplatin-induced apoptosis (Figure 3(c)), which further verified the results of flow cytometry. For cell cycle analysis, there were no significant differences. Such experiments demonstrated that Hsv2-miR-H9-5p can promote proliferation, and its overexpression augmented the resistance of lung cancer cells to apoptosis induced by the cisplatin treatment. 18 These results suggest that Hsv2-miR-H9-5p plays an oncogenic role in lung cancer cells in vitro.

Hsv2-miR-H9-5p plays an oncogenic role in lung cancer cells in vitro. (a) CCK-8 assay reveals that hsv2-miR-h9-5p upregulation induced growth in LTEP-α-2 and SPC-α-1 lung cancer cells (**p < 0.01). (b) Annexin V–FITC/PI staining of cells treated with 20 µM cisplatin for 24 h (*p < 0.05). (c) Representative morphology of lung cancer cells transduced with Hsv2-miR-h9-5p mimic and negative control treated with 20 µM cisplatin for 24 h.
Hsv2-miR-H9-5p regulates cellular SOCS2 expression through directly binding to the SOCS2 3′-UTR in lung cancer cells
We investigated Hsv2-miR-H9-5p-targeted genes that are associated with lung cancer metastasis by TargetScan6.0. In fact, from an evolutionary standpoint, although many viruses have evolved with their own functional miRNA genes, they also utilize the mechanism of modulating cellular miRNAs to influence a repertoire of mRNA targets. In this situation, some kind of virus appears to have evolved to encode miRNAs which can target an already existing miRNA/targetome pathway. 18 Salient examples of this is the Kaposi’s sarcoma herpesvirus (KSHV) encoded miR-K12-11 and Marek’s disease virus (MDV-1) encoded miR-M4 which have the identical seed sequence as that of the cellular miRNA, miR-155. Having identical seed sequences, these virus-encoded miRNAs target a highly overlapping set of genes with that of miR-155.20–22 Considering this possibility, we first searched the identical or similar seed sequences of Hsv2-miR-H9-5P and no significant similarity found. We next examined the mechanisms by which Hsv2-miR-H9-5P promotes tumor progression by combining a computational method (TargetScan 6.0, November 2011; http://www.targetscan.org/vert_60/) with the mRNA microarray expression data (Figure 4(a)). In previous experiments of apoptosis, we observed the strong anti-apoptosis effect of Hsv2-miR-H9-5p, and combined with the function of SOCS2, we presumed among the candidate genes SOCS2 as a potential target of Hsv2-miR-H9-5P; SOCS2 is a protein of interest given that SOCS2 negatively regulates cancer progression and has correlation with patients’ prognosis and survival.

Hsv2-miR-H9-5p suppresses SOCS2 expression by directly targeting the SOCS2 3′-UTR. (a) Heat map showed an intersection of mRNA microarray expression data and the target genes that TargetScan6.0 predicted. (b) Dual-luciferase assay of pmiR-GLO-SOCS2-3′-UTR reporter in cells into which Hsv2-miR-H9-5p had been transduced and predicted Hsv2-miR-H9-5p target sequence in the 3′-UTR of SOCS2 (WT 3′-UTR) and mutant containing the mutated nucleotides in the SOCS2 3′-UTR (MUT 3′-UTR; *p < 0.05). (c) Real-time PCR analysis of SOCS2 after Hsv2-miR-h9-5p mimic and NC transfection at different times in LTEP-α-2 and SPC-α-1 cells. GAPDH was used as a loading control (*p < 0.05). (d) Western blot of SOCS2 after LTEP-α-2 cells were transfected with mimic or NC after 48 and 72 h. GAPDH served as the loading marker (*p < 0.05). (e) Western blot of SOCS2 after SPC-α-1 cells were transfected with mimic or NC after 48 and 72 h. GAPDH served as the loading marker (*p < 0.05). (f) Immunohistochemical staining data shows that SOCS2 is downregulated in the lung cancer tissues in comparison with normal adjacent tissues.
Bioinformatics analysis showed that SOCS2 3′-UTR region contains the site complementary with the seed sequences of Hsv2-miR-H9-5p (Figure 4(b)). To clarify whether SOCS2 is a direct cellular target for Hsv2-miR-H9-5p, we performed dual-luciferase reporter assays by co-transfection of a wild-type (WT) or mutant (MUT) SOCS2 3′-UTR-containing luciferase reporter vector with Hsv2-miR-H9-5p mimic. The luciferase activity of the WT SOCS2 3′-UTR but not the mutant 3′-UTR was significantly reduced by Hsv2-miR-H9-5p mimic alone but not by the negative control (Figure 4(b)).
The effects of Hsv2-miR-H9-5p on SOCS2 mRNA and protein expression were examined in lung cancer cell lines LTEP-α-2 and SPC-α-1. Upregulation of Hsv2-miR-H9-5p significantly reduced the expression of SOCS2 in both LTEP-α-2 and SPC-α-1 cell lines compared with the relative control as revealed by qRT-PCR (Figure 4(c)) and western blotting (Figure 4(d) and (e)). Immunohistochemical staining data showed that SOCS2 was downregulated in the lung cancer tissues in comparison with the normal adjacent tissues (Figure 4(f)). Taken together, these data indicated that SOCS2 was a direct cellular target of Hsv2-miR-H9-5p in lung cancer cell lines.
Restored SOCS2 inverts the phenotypes produced by Hsv2-miR-H9-5p
We transiently transfected SOCS2 expression vector pIRES2-EGFP-SOCS2 into LTEP-α-2-mimic and SPC-α-1-mimic cells. Restored SOCS2 expression significantly inhibited cell proliferation (Figure 5(a)), promoted apoptosis (Figure 5(b) and (c)), and reduced the migration (Figure 5(d)) and invasion (Figure 5(e)) of both LTEP-α-2-mimic and SPC-α-1-mimic cells compared with the relative pIRES2-EGFP blank plasmid control (Blank). Obviously, the reconstitution of SOCS2 inversed the results obtained from the upregulation of Hsv2-miR-H9-5p.
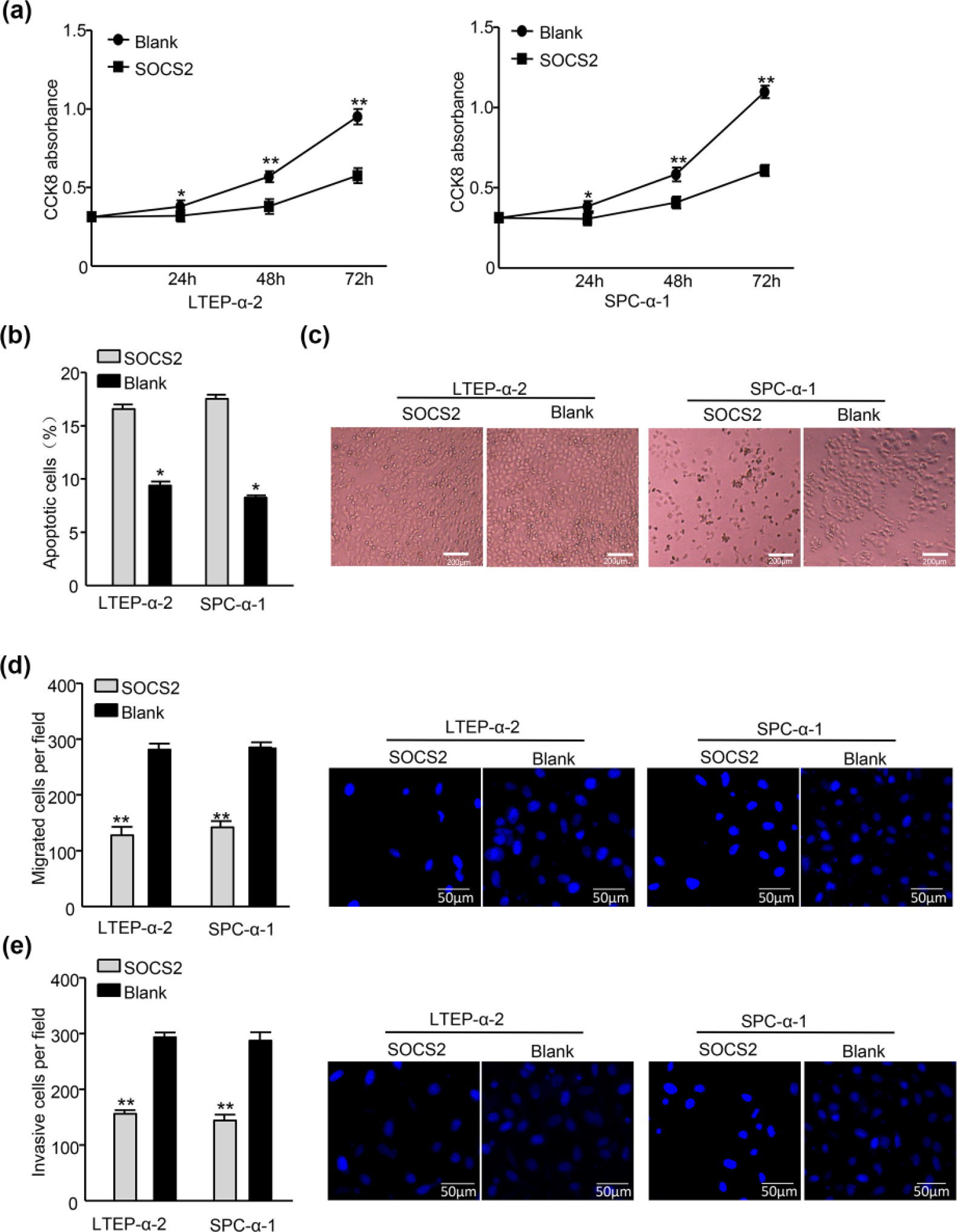
Reconstitution of SOCS2 inverts the phenotypes generated by Hsv2-miR-H9-5p. (a) CCK-8 assay reveals that SOCS2 upregulation decreased growth of LTEP-α-2-mimic and SPC-α-1-mimic lung cancer cells (**p < 0.01). (b) Flow cytometric analysis of lung cancer cells transduced with SOCS2 or blank plasmid and Annexin V–APC/PI staining of cells treated with 20 µM cisplatin for 24 h (*p < 0.05). (c) Representative morphology of lung cancer cells transduced with SOCS2 and blank plasmid treated with 20 µM cisplatin for 24 h. (d) Reconstitution of SOCS2 reduced the cell migration of both LTEP-α-2-mimic and SPC-α-1-mimic cells (*p < 0.05). (e) Reinstallation of SOCS2 reduced the cell invasion of both LTEP-α-2-mimic and SPC-α-1-mimic cells (*p < 0.05).
Discussion
The increasing data have provided compelling evidence that the aberrant expression of some miRNAs can enhance initiation and malignant progression of various types of cancers. MiRNAs have been shown to regulate multiple aspects of the metastatic process, 23 mainly through negatively regulating multiple target genes. We identified a number of miRNAs that were aberrantly expressed in lung cancer compared with lung cancer bone metastasis by microarray.
Among them, we focused on HSV2-encoded miRNA, Hsv2-miR-H9-5p, which we found to regulate SOCS2 expression by directly targeting the SOCS2 3′-UTR. To date, several researches verified the carcinogenic effect of viruses and viruses-derived miRNAs, a hallmark of viruses is their incredible mastery of the art of cellular hijacking to achieve their own tenets of self-preservation and proliferation. With their limited genetic capacity, they rely exquisitely on a wide range of host functions to achieve these tenets. Nevertheless, they seem never content to simply utilize existing cellular functions in their natural state but instead modulate and fine-tune each of these pathways/functions in ways that provide a greater advantage for the virus. Given the now well-established role of miRNAs in regulating almost every aspect of the cellular regulatory machinery, it is not surprising to find viruses interacting with these pathways as a means of modulating the host environment to suit their needs. Although the study of how viruses utilize miRNA pathways in their life cycle is in its infancy, we know that this intersection with miRNA pathways occurs through virus-mediated modulation of existing cellular miRNA expression and/or through the evolution of a virus-encoded repertoire of miRNAs. 18
To the best of our knowledge, we are the first to report a link between Hsv2-miR-H9-5p and lung cancer bone metastasis. With experimental approaches, we found that Hsv2-miR-H9-5p is important in the progression and metastasis of lung cancer. We determined the correlation of Hsv2-miR-H9-5p with bone metastasis of lung cancer patients. The change in expression of Hsv2-miR-H9-5p detected by microarray analysis was further confirmed by qPCR.
However, the specific underlying pathophysiological mechanisms linking Hsv2-miR-H9-5p with lung cancer bone metastasis are yet to be defined. How, then, does Hsv2-miR-H9-5p exert its influence on lung cancer bone metastasis?
Herpesviruses encode various miRNAs to regulate viral replication and to counteract host defense such as apoptosis and immunity. 3 Cytomegalovirus (CMV) exploits miR-UL112-1 to evade immune surveillance by inhibiting MICB gene 24 and to suppress expression of the major immediate-early genes. 25 In consideration of the anti-apoptotic function of LAT that already reported 15 and the conserved mechanism in these closely related herpes simplex viruses combined with the bioinformatics analysis, we confirmed that Hsv2-miR-H9-5p directly binds the 3′-UTR of SOCS2 mRNA and inhibits its translation in lung cancer cells with dual-luciferase assay.
SOCS2 was involved in many types of cancer, including breast cancer, gastric cancer, and prostate cancer. Several mechanisms have been proposed to mediate SOCS2 dysregulation in cancers. In our studies, we demonstrated that Hsv2-miR-H9-5p inhibited SOCS2 expression and confirmed that SOCS2 was a direct target of Hsv2-miR-H9-5p in lung cancer cells. We found that knockdown of SOCS2 induced cell anti-apoptosis and promoted the migration and metastasis similar to the phenotypes induced by Hsv2-miR-H9-5p mimic, and further experiments revealed that SOCS2 overexpression inverted the effect of Hsv2-miR-H9-5p in lung cancer cells.
In conclusion, we have demonstrated that Hsv2-miR-H9-5p was highly expressed in bone metastasis of lung cancer patients and closely associated with invasive biological characteristics of the patients. Hsv2-miR-H9-5p increased the cell migration and invasion of lung cancer cells in vitro by directly targeting SOCS2.
Our findings provided new insights into the invasiveness and metastasis of lung cancer regulated by miRNAs derived from diverse viruses and rational for the development of clinical intervention strategies for lung cancer bone metastasis.
With the experiment proceeded, we noticed that there were some limitations to the present research. First, in our initial microarray cohort consisting of 10 lung cancer bone metastasis cases and 10 control subjects, the tissue samples were not matched one by one, which might potentially confound our results because of the individual difference. In order to eliminate these variants and to further validate the microarray data results, we expanded the sample size. However, there were no significant differences between the results. Second, our findings from Chinese lung cancer and bone metastasis patients may not be generalizable to other populations because of the variants across populations. Furthermore, due to the limitation of the virus packaging and the level of the experimental technique, we tried and failed to produce the viral vector which can generate Hsv2-miR-H9-5p especially, and we had to suspend the subsequent animal experiments, so whether the assessment of specificity of Hsv2-miR-H9-5p changes was specific for lung cancer bone metastasis will need to be the focus of future studies.
Footnotes
Acknowledgements
The authors would like to thank Aiying Du for critical reading and critical discussion during the experiments. X.W., S.L., and Z.Z. equally contributed to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
